Abstract
Oculocutaneous albinism (OCA) is characterized by hypopigmentation of the skin, hair and eye, and by ophthalmologic abnormalities caused by a deficiency in melanin biosynthesis. In this study we recruited 321 albino patients and screened them for the genes known to cause oculocutaneous albinism (OCA1–4 and OCA6) and ocular albinism (OA1). Our purpose was to detect mutations and genetic frequencies of the main causative genes, offering to albino patients an exhaustive diagnostic assessment within a multidisciplinary approach including ophthalmological, dermatological, audiological and genetic evaluations. We report 70 novel mutations and the frequencies of the major causative OCA genes that are as follows: TYR (44%), OCA2 (17%), TYRP1 (1%), SLC45A2 (7%) and SLC24A5 (<0.5%). An additional 5% of patients had GPR143 mutations. In 19% of cases, a second reliable mutation was not detected, whereas 7% of our patients remain still molecularly undiagnosed. This comprehensive study of a consecutive series of OCA/OA1 patients allowed us to perform a clinical evaluation of the different OCA forms.
Similar content being viewed by others
Introduction
Oculocutaneous albinism (OCA) is characterized by general hypopigmentation of skin, hair and eye, and by specific ophthalmologic abnormalities caused by a deficiency in melanin biosynthesis. Ocular abnormalities include congenital nystagmus, reduced pigmentation of the iris and retinal pigment epithelium, photophobia, foveal hypoplasia, reduced visual acuity and misrouting of the optic nerves that can result in strabismus and reduced stereoscopic vision. These features are common to all types of albinism and are probably related to melanin reduction during embryonic development and early postnatal life.1, 2 OCA can affect all ethnic groups with an overall prevalence of ∼1/17 000 people3 and is commonly subdivided into four main types (OCA1–4) because of mutations in the TYR, OCA2, TYRP1 and SLC45A2 genes, respectively.4, 5, 6, 7 To date, over 600 different recessive mutations have been described in these four genes.8 At least three additional loci involved in nonsyndromic OCA have been reported: an OCA locus mapping in 4q24 found in a consanguineous Pakistani family, whose gene (termed OCA5) has not yet been identified,9 and two genes, SLC24A510 and C10orf11,11 named OCA6 and OCA7.12 OCA6 has not yet a worldwide prevalence, but it is present in European population accounting for ∼1.25% of OCA cases.13
Furthermore, an X-linked form of ocular albinism (OA1), whose phenotype is mainly restricted to eyes and optic system, is associated with mutations in the GPR143 gene.14 OA1 patients are characterized by the presence of macromelanosomes in both skin and retinal pigment epithelium.15 Heterozygous females are asymptomatic, but could present iris transillumination and blotchy pattern of retinal pigmentation as a result of random X-chromosome inactivation.14
Albinism causes significant eye morbidity and amblyopia in children and ophthalmological evaluation allows an earlier diagnosis of OCA and OA in newborns. In OCA patients hypopigmentation of the skin causes severe photosensitivity and an increase risk of developing sun-induced skin cancer. This implies the need for albino patients of accurate dermatological controls during the lifetime.16, 17 In addition, hearing deficits have been reported in albino subjects of different species; in fact, although melanin seems not to be essential for normal auditory function, it is thought to have a protective role against age-related hearing impairment18 and reduced recovery of auditory thresholds after noise exposure.19 For these reasons, the majority of our patients underwent ophthalmological, dermatological, audiological and genetic evaluations within a 1-day multidisciplinary day hospital.
In this study we analyzed a consecutive series of 321 albino patients reporting genetic frequencies of the main causative genes and 70 novel mutations. This comprehensive work allowed us to perform clinical evaluation of our albino patients and to estimate the relative frequencies of the different molecular forms in our population.
Materials and methods
Patient recruitment
A total of 321 unrelated albino subjects were recruited from the Medical Genetics Unit and the Department of Pediatric Ophthalmology of the ASST Niguarda Hospital of Milan through a multidisciplinary diagnostic workup that included ophthalmologic exams, dermatological and ENT (ears, nose and throat) visits and genetic counseling. Furthermore, different Genetic Units and Ophthalmology Centers in Italy contributed to our cohort of albino patients.
Blood samples were also collected, when available, from proband parents. This study was conducted according to the Declaration of Helsinki and to the Italian laws. A signed informed consent for the genetic analysis was obtained from all involved subjects and from parents of probands <18 years old.
Ophthalmologic evaluation
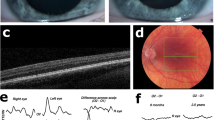
Albino patients of Niguarda Hospital underwent full ophthalmological and instrumental evaluations including: orthoptic evaluation, visual acuity, cicloplegic refraction, iris transillumination with slitlamp biomicroscope and macular translucency with indirect ophthalmoscope. Visual acuity assessment for distance and near was evaluated in monocular and binocular vision with appropriate tests (Pigassou figures, E test and Snellen charts).
Using spectral domain optical coherence tomography (Heidelberg Engineering, Heidelberg, Germany), we evaluated the macular morphological characteristics with attention to the presence or absence of foveal depression. We could not employ spectral domain optical coherence tomography in newborns and in noncompliant patients.
Visual evoked potentials were performed to evaluate the optic nerve fiber decussation (Retimax, CSO, Scandicci, Italy).
Dermatological evaluation
Albino patients were carefully examined at the ASST Niguarda Hospital, Department of Medicine, by well-trained dermatologists. Besides traditional dermatological examination by epiluminescence light microscopy, digital epiluminescence was employed in order to evaluate lesions at higher definition and magnification with a better diagnostic accuracy.
Audiological evaluation and ENT examination
Patients were visited at the Department of Otolaryngology by an ENT specialist with a standard examination, focusing on eardrum integrity and conformation, nystagmus existence, cleft shape, bite abnormality, nose airway clearance and tonsil dimension.
Adult patients received audiological evaluations including pure-tone audiometry and acoustic reflex study. Conventional audiometry was usually used in children aged ⩾5 years. For the younger infants (<5 months), audiologist employed behavioral observation audiometry; visual reinforcement audiology testing has been used to evaluate the hearing of infants from 6 months to 2–3 years, whereas condition play audiometry was useful for developmental age of 2–3 to 4–5 years. Young patients were examined with tympanometry and acoustic reflex study when it was possible to perform it.
DNA analysis
DNA was extracted from peripheral blood according to standard protocols.
The coding regions and adjacent flanking regions of TYR, OCA2, TYRP1, SLC45A2, SLC24A5 and GPR143 genes were PCR amplified and sequenced on an ABI Prism 3730 genetic analyzer and the SeqScape software (Life Technologies, Carlsbad, CA, USA) was used for mutation detection. For the amplification of the TYR locus we used specific primers able to discriminate between TYR gene and TYRL pseudogene exons 4 and 5 to allow direct sequencing and identification of TYR mutations and single-nucleotide polymorphisms.20 The lists of primers and protocols are available on request. Analysis of the 2.7 kb deletion of OCA2 gene was performed according to literature.21 To screen for TYR and OCA2 exons rearrangements (deletion/duplication), multiplex ligation-dependent probe amplification analysis was performed (SALSA MLPA Kit P325, MRC-Holland, Amsterdam, The Netherlands). Raw data were analyzed using the Coffalyser software v.9 (MRC-Holland).
In silico analysis
To assess variant nomenclature according to HGVS and the pathogenicity of the genetic alterations found, two web-bioinformatic tools were used, respectively: Mutalyzer 2.0.beta-2622 and MutationTaster;23 computer-assisted analysis for splice-site prediction was accomplished using the NNSPLICE 0.9 program (http://www.fruitfly.org/seq_tools/splice.html).
Results
In this study we recruited 321 albino patients and screened them for the main genes known to cause oculocutaneous albinism (OCA1–4 and OCA6) and ocular albinism (OA1). The age at diagnosis varied from 3 months to 81 years, although the majority of our patients are in pediatric age; 279/321 patients are Italian, whereas the remaining belong to different ethnic groups (Table 1). Clinical history and physical examination did not reveal any evidence of dysmorphic features in the analyzed subjects.
Genetic analysis identified 184 subjects with mutations in the TYR gene, associated with OCA1 phenotype, 138 of which harboring two TYR mutations, whereas 46/184 subjects were carriers of a single TYR mutation. Within the latter group, 39/46 patients were carriers of the p.R402Q thermo-sensitive variant in trans, as confirmed by familial segregation analysis, 30 of which with mild hypopigmentation and/or mainly ocular phenotype, whereas another one (patient 489 (pt489)) also had a SLC45A2 (OCA4) mutation. One patient (pt476) was carrier of a single TYR and TYRP1 (OCA3) mutation. In 6/46 patients we did not identify a second mutated allele. We found 24 novel TYR mutations: 15 missense, 2 splice-site mutations, 1 deletion in frame, 4 frameshifts and 2 exon deletions (of the single exon 3 and of the single exon 5).
We found OCA2 mutations in 65 albino subjects: 54/65 patients were homozygous or compound heterozygous and 3/54 also had a mutation in TYRP1 (pt145) and TYR (pt292 and pt537), respectively. In addition, 11/65 subjects presented only one OCA2 mutation and one patient (pt307) also had the p.Y438* mutation on a TYRP1 allele. We identified 23 OCA2 novel mutations including 10 missense, 1 nonsense, 8 splice site, 2 frameshift and 2 homozygous deletions: one from exons 12 to 18 and one from exons 7 to 8. The recurrent ‘African’ 2.7 kb deletion was identified in 4 patients: 3 from Cameroun and 1 from Nigeria.
Four patients were found to be mutated in the TYRP1 gene (OCA3), with 3 with biallelic mutations, the nonsense mutation p.Y519* in homozygous state, the novel missense mutation p.G267E in homozygous state and the missense mutations p.E140* and p.R164G in compound heterozygosity, whereas the last one had only the TYRP1 p.R93H mutation in heterozygous state.
SLC45A2 (OCA4) mutations were found in 26 affected subjects, 23/26 with biallelic SLC45A2 mutations and 3/26 with a single pathological mutation. We identified 11 SLC45A2 novel mutations: 5 missense, 1 nonsense, 4 frameshifts and 1 splice-site mutation.
SLC24A5 mutational screening identified one patient (pt162) with two OCA6 mutations: the p.Tyr72* and the c.385+2T>G (IVS3+2T>G) inherited from his heterozygous parents.
Finally, in 17 OA1 male patients we identified GPR143 mutations, 11 of which are here described for the first time. Among these new mutations, 4 are missense, 5 are frameshift and 2 involve splice sites.
However, 24/321 patients (7%) remain still unresolved as they have only OCA gene polymorphic variants.
As a result of this molecular study we estimated the frequencies of the different molecular forms in our cohort of 321 consecutive albino patients. We identified causative mutations accounting for: 44%, TYR; 17%, OCA2; 1%, TYRP1; 7%, SLC45A2; and <0.5%, SLC24A5. Male patients with a OA1 phenotype and GPR143 defects account for 5% of all cases. In 19% of OCA patients, a second reliable mutation (see Discussion below) was not detected, whereas 7% of our cases did not show any genetic alterations in the tested genes. These results are summarized in Figure 1. Frequencies calculated only on Italian cases (N=279) do not significantly differ (data not shown).
Clinical diagnosis of albinism was mainly established through ophthalmologic examinations. The distribution of eye disorder severity has been divided into six groups: nystagmus (presence/absence), visual acuity for distance, iris transillumination, macular transparency, foveal morphology and visual evoked potential decussation (presence/absence) (Table 1).
At dermatological examination, actinic keratoses were detected in sun-exposed areas of three adult patients (pt340 age 37, pt264 age 47 and pt344 age 29 years). They were topically treated by specific drugs. A case of malignant cutaneous melanoma was identified on the right leg of a 40-year-old male (pt369) patient histologically classified as superficial spreading, Clark level 4, Breslow 0.8. He was promptly sent to the surgical department where he underwent specific treatment protocol. The same patient had multiple actinic keratoses on the face and also on the back of the hands. In spite of the lack of melanin, skin pigmented lesions were present in most of the adult patients, the majority of which were benign (common nevi and angiomas), without submicroscopic and vascular atypias.
After audiological evaluation, the majority of the patients showed normal hearing thresholds, tympanometric and acoustic reflex results. In four subjects (pt187 age 40, pt205 age 49, pt266 age 39 and pt345 age 38 years) the pure tone audiogram showed a bilateral slight sensorineural hearing loss in the high frequencies. Four patients (pt197 age 50, pt376 age 26, pt383 age 44 and pt493 age 59 years) showed bilateral whereas two other patients (pt316 age 44 and pt340 age 37 years) showed unilateral mild sensorineural hearing loss at 4000 and 8000 Hz. The audiogram of patient 264 (age 47 years) showed a bilateral moderate-to-severe sensory neural hearing loss, whereas patient 186 (age 70 years) had a normal hearing sensitivity in the lower frequencies and a precipitously sloping into the moderate hearing loss range beginning at 2000 Hz in both ears.
Discussion
We report here the results of a comprehensive molecular analysis of a cohort of 321 consecutive unrelated albino patients, including 279 Italian subjects and 42 patients from Africa, Asia, South America and other Caucasian countries.
A total of 70 novel mutations were identified. None of these new mutations were present in dbSNP (www.ncbi.nlm.nih.gov/SNP/), TGP (http://www.1000genomes.org/), ExAC (http://exac.broadinstitute.org/) or in HGMD (http://biobase-international.com/hgmd/pro/all.php) databases.
We identified TYR mutations in 184 subjects indicating OCA1 form as the most frequent in our country. Among the 24 new TYR mutations, the 4 frameshift mutations result in premature stop codons (c.69del, p.C24Vfs*7; c.341del, p.E114Gfs*6; c.1072_1075dup, p.Q359Lfs*37; c.1386-1387insAA, p.Y463Nfs*23), whereas the novel splice mutations c.1037-2A>C and c.1184+2T>G lie in exon 2 acceptor splice site and in exon 3 donor one, respectively, abolishing them; all these mutations are predicted to be subject to nonsense-mediated decay.24 The 15 new missense mutations are predicted to be deleterious because of their location in highly conserved residues of known tyrosinase functional domains. The p.F20C, p.C46R, p.C89W, p.L117I, p.K133N and p.T144I mutations are located in the first cysteine-rich region, whereas p.L225P and p.V318A are located in the second one; two of them, p.T144I and p.L225P (pt227), lie in the same allele in CIS p.[(T144I; L225P)] as demonstrated by segregation analysis. The p.V177I, p.Y182C, p.D333Y, p.G346R, p.S375F, p.F397S and p.M426I mutations lie in the tyrosinase catalytic domain containing two copper-binding sites essentials for its catalytic activity.25 We also found one in frame deletion (c.806_808del) causing the removal of a single amino acid (p.F269del) located within the second cysteine-rich region and predicted to be deleterious for the protein. Among the TYR deletions, identified by multiplex ligation-dependent probe amplification analysis, we report two novel ones: the exon 3 deletion identified in homozygous state in a Turkish patient (pt281) with consanguinity and the exon 5 deletion found in four Italian patients, one in homozygous state and three in compound heterozygosity with other TYR mutations. Subsequent DNA analysis of the parents confirmed the segregation of the mutated alleles.
In 39/184 OCA1 patients we found a single severe TYR mutation in compound heterozygosity with the common thermosensitive TYR variant p.R402Q. The causative role of the p.R402Q variant is still controversial. This allele codes for a thermolabile tyrosinase with a decreased catalytic activity at 37 °C and it has been considered for years a nonpathologic variant owing to its high prevalence in the population.26 In fact, in our series also it is the most frequent mutated allele as we found it in 123/321 (38%) albino patients and in 148 alleles. This variant itself is not enough to cause OCA given that individuals homozygous for p.R402Q do not show clinical signs of albinism.27, 28 However, other studies indicated this variant as a possible causative mutation in combination with severe TYR mutations,8, 29, 30 maybe when paired with a not benign genetic background.31 In support of this hypothesis, we noted that p.R402Q is more common in our OCA patients carrying only one TYR mutation (39/46, 84.8%) than in patients with two mutations (51/138, 36.9%). If we were to consider these 39 patients as resolved cases, the percentage of OCA1 patients would rise to 55% from 44% and the percentage of unresolved cases would drop from 19 to 7%. All these subjects were anyway analyzed for our panel of OCA genes and also for OA1 in male patients, and in one patient (pt489) one SLC45A2 mutation has been also found, whereas in the other patients no additional OCA mutations have been identified.
We identified 54/65 subjects with two mutations in the OCA2 gene, indicating this gene as the second most frequent in our population (17%). We found 23 novel mutations including 2 partial homozygous OCA2 gene deletions: one spanning from exons 12 to 18 (Italian patient 115) whose inheritance from the parents was confirmed, as well as no consanguinity indicating this as a regional private mutation, and the second one from exons 7 to 8 (Pakistani patient 354) from consanguineous parents. In addition, 6/54 patients carried one OCA2 mutation with the p.R305W variant on the other allele and one patient (pt171) resulted homozygous for this mutation. The p.R305W was initially described as a coding alteration responsible for phenotypic variations in human eye color,32 but a recent computational study identifies it as most deleterious and disease-associated variant,33 and we agree that this mutation in homozygous state or in association with another OCA2 mutation could explain the albino phenotypes of patients. We also found a novel synonymous transition (c.1842G>A, p.K614K) involving the last nucleotide of exon 17. The in silico analysis performed by computer-assisted software for splice-site prediction NNSPLICE 0.9 detected the wild-type donor splice site with a score of 0.94, whereas in the mutant sequence, it was no longer identified. A similar mutation, falling in the last nucleotide of exon 20 (c.2139G>A, p.K713K), has been recently demonstrated to be causative through an in vitro hybrid-minigene approach by our group.34 All the novel OCA2 missense mutations p.S216P, p.A355E, p.T453P, p.L460F, p.G479R, p.I492T, p.L639R, p.V677A, p.A733P and p.C793Y are well conserved during evolution and predicted to be damaging by in silico analysis. We also identified a new nonsense (p.R165*) and two frameshifts (c.2404dup, p.Y802Lfs*48; c.1841dup, p.H615Afs*2), all predicted to result in premature truncation of the P protein and to be subjected to nonsense-mediated decay. The effect of the seven novel OCA2 splice-site mutations (c.228-1G>C, IVS2-1G>C; c.574-1G>A, IVS5-1G>A; c.647-2_-13del, IVS6-2_-13del; c.1045-1G>C, IVS9-1G>C; c.1117-5T>G, IVS10-5T>G; c.1503+2T>C, IVS14+2T>C; c.2244+2T>G, IVS21+2T>G) were analyzed by NNSPLICE 0.9 program. For all these mutations the software predicted the disruption of the native splice sites and aberrant splicings.
In our series we also found three Italian patients with an apparently digenic OCA inheritance (pt307, pt476 and pt489). The first one (pt307) presented a mild phenotype with only one known OCA2 mutation and the p.Y438* TYRP1 mutation, whereas the other two carried one TYR and one TYRP1 mutation (pt476) and one TYR and one SLC45A2 mutation (pt489) with a quite classical OCA phenotype. Although some authors reported about patients with digenic albinism,35 we, together with many other authors,36, 37 usually do not believe in OCA digenic inheritance as the evidence in familial segregation analysis of identified healthy parents or sons with digenic OCA mutations. This is confirmed by our data for OCA1/OCA2, OCA1/OCA4 and OCA2/OCA3 genotypes, but to date, we have not enough data to rule out a possible additive effect of OCA1 and OCA3 gene mutations as parents of patient 476 carrying one mutation each. Moreover, we know that Tyr and Tyrp1 interact in a unique melanogenic protein complex and that a mutation of one of these proteins may affect the maturation and/or degradation of the other,38 and hence we cannot totally reject the hypothesis of a causative digenic interaction of OCA1/OCA3. Of course, the apparent digenic OCA inheritance could also be explained by the presence of genetic alterations not detectable by our actual molecular methods (see discussion below).
Four patients were found to be mutated in TYRP1 (OCA3), and one out of the seven mutations identified, the p.G267E, was novel. It was found in homozygosis in patient 266 and predicted to be disease causing by in silico analysis involving a highly conserved amino acid residue.
Twenty-three patients had two mutations in the SLC45A2 gene (OCA4). In total, we found 24 OCA4 mutations, 11 of which were novel. All the 5 novel missense mutations (p.A54E, p.L65V, p.G148S, p.C481Y and p.G627E) affect highly conserved amino acid residues. The three novel frameshift mutations identified, c.391_394del (p.I131Lfs*11), c.511_520del (p.C171Rfs*27) and c.614_617dup (p.E206Dfs*35), together with the nonsense mutations p.E282* and p.Y319* are predicted to result in the premature termination of the polypeptide chain and can therefore be considered pathogenic. The c.1369-3C>A substitution was identified in the consensus sequence of the acceptor splice site of intron 6 (IVS6-3C>A); NNSPLICE 0.9 analysis predicted a score reduction in the mutated sequence (0.96 to 0.58) and the possible activation of one or two stronger cryptic acceptor splice sites (one located 31 bp upstream, the second 212 bp downstream the physiologic one). This mutation was found in compound heterozygosity with mutations p.H94D and p.G100S lying in CIS on the same allele p.[(H94D; G100S)]. Noteworthy, this already reported pathological allele39 has also been found in two other unrelated Italian patients (Table 1).
Recently, we introduced OCA6 mutational screening to implement OCA molecular diagnosis and we analyzed the SLC24A5 gene in patients who resulted negative or heterozygous for one mutation in the OCA1-4 gene. We identified the first Italian albino patient (pt162) with two OCA6 pathogenic mutations, the c.216T>G (p.Tyr72*) and the c.385+2T>G (IVS3+2T>G), from 321 patients (0.3%), indicating OCA6 as the less frequent OCA gene in our series.40
Finally, we identified 17 male patients with mutations in the GPR143 gene associated with X-linked ocular albinism, and 11 of these have novel OA1 mutations. Four frameshift mutations, c.42_61del, c.580del, c.327_355dup and c.723dup, cause the premature termination of the amino acid chain p.D15Afs*79, p.H194Tfs*24, p.S119Kfs*36 and p.I242Dfs*23, respectively, whereas the c.768-2_769del includes and abolishes the acceptor splice site of intron 6 and the first two nucleotides of exon 7, and can therefore be considered pathogenic. The c.548+1G>C mutation lies in the invariant dinucleotide GT of the donor splice site of GPR143 intron 4 (IVS4+1G>C). The in silico analysis of these mutant sequences did not identify any splice-site activity.
We identified four novel missense mutations, p.L34P, p.G172R, p.N260K and p.N299R, all lying in OA1 transmembrane domains and near amino acid positions (p.G35D, p.A173D, p.I261N and p.W292G) that have been well characterized at the functional level. In fact, it has been demonstrated that these OA1 mutants were retained in the endoplasmic reticulum, showing defective glycosylation and reduced yield.41 We also found the nucleotide substitution c.727G>A (p.K243E) on the same allele with the c.723dup frameshift mutation. We consider that this further substitution should not create additional problems as the protein is anyway interrupted, but we are not able to give a pathological significance to this variant alone.
Diagnosis of oculocutaneous albinism is principally based on clinical examination and may be classified according to the hypopigmentation trait (relative to other family members) and to the ophthalmic findings. When possible (for many outpatients we have no phenotype information), we collected detailed ophthalmological and dermatological data. This study also tried to provide information about OCA and OA1 genotypes and degree of ocular problems, but we did not observe clear genotype–phenotype correlations. Furthermore, we noticed variable intra- and interfamilial degrees of eye disorder gravity. It is still unclear why the same type of mutations leads to different degrees of ocular disorders. This variability may reflect the effect of modifier genes and/or environmental factors leading to incomplete penetrance and variable expression of OCA alleles. It could be hypothesized that additional mutations or polymorphisms in other interacting genes, as described about the crucial role of TYRP1 in stabilizing tyrosinase activity,38 may influence the extent of eye disorder gravity. Characterization of such modifier genes will be essential for improving OCA phenotype prediction.
Unresolved cases and apparently digenic OCA inheritance could be explained by the presence of genetic alterations located in the promoter, in noncoding genomic regions or far away from the locus in relevant transcriptional regulators sequence,42 or by the presence of exonic unrecognized splicing mutations,33, 43, 44 large deletions or complex gene rearrangements45, 46 not detectable by Sanger sequencing. Therefore, in the near future, we consider a next-generation sequencing approach essential for molecular diagnosis of albino patients, including entire OCA/OA genes (exons, introns and flanking regions) in order to identify rearrangements embedded in the introns or in the regulatory regions or gene dosage anomalies with reading depth measurements. The existence of new OCA loci yet to be identified could obviously be considered.
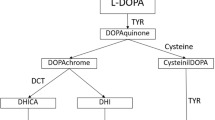
Audiological evaluation of our patients identified 12 adult subjects with different degrees of hearing impairment. As melanin binds calcium, its function in the inner ear seems to be implicated in the endolymph homeostasis. Calcium is known to enter into the cochlear hair cells from endolymph during sound stimulation and its reduced availability in albinism could modify functional properties of these cells.47, 48 It has been demonstrated in a mouse model of human OCA1 that L-DOPA (L-3,4-dihydroxyphenylalanine), which is a melanin precursor equally able to bind calcium, is also capable of preventing age-related and noise-induced hearing loss.49 It is noteworthy that patients who showed hearing impairment are not only OCA1 albino subjects; in fact, patient 187 is OCA2, patient 266 is OCA3 and patients 205 and 186 are OCA4. These results demonstrate that age-related hearing impairment in albino patients could be caused by the lack of the pigment, along with associated metabolites, regardless of their OCA genotypes.
The aim of the dermatological examination was to detect pigmented skin lesions (nevi, angiomas) and sunlight damages in albinos and to give them correct information about sun exposure in current life. Thanks to these dermatological controls, three cases of actinic keratosis and one case of melanoma were identified and treated in four adult patients. Adequate photo protection by sunscreens and dietary supplements were recommended to all patients together with periodic dermatological checks.
An early ophthalmologic diagnosis of albinism allows patients to receive support and visual stimulation programs that could improve their vision beyond the limited associated visual acuity. We would also like to point out that the coordinated efforts of Pediatric Ophtalmology Department, Dermatologists, Department of Otolaryngology and Genetic Units are critical for a timely and accurate diagnostic evaluation of albino patients and families and to care and manage their related clinical needs. Moreover, molecular genetic diagnosis improves the ability of patients and parents to cope with their clinical and genetic condition with consequent better life quality.
References
King, R. A., Hearing, V. J., Creel, D. J. & Oetting, W. S. in The Metabolic and Molecular Basis of Inherited Disease 8th edn Vol. II, 5587–5627 (McGraw-Hill, New York, NY, 2001).
Kirkwood, B. J. Albinism and implications with vision. Insight 34, 13–16 (2009).
Grønskov, K., Ek, J. & Brondum-Nielsen, K. Oculocutaneous albinism. Orphanet J. Rare Dis. 2, 43–50 (2007).
Tomita, Y., Takeda, A., Okinaga, S., Tagami, H. & Shibahara, S. Human oculocutaneous albinism caused by single base insertion in the tyrosinase gene. Biochem. Biophys. Res. Commun. 164, 990–996 (1989).
Rinchik, E. M., Bultman, S. J., Horsthemke, B., Lee, S. T., Strunk, K. M., Spritz, R. A. et al. A gene for the mouse pink-eyed dilution locus and for human type II oculocutaneous albinism. Nature 361, 72–76 (1993).
Boissy, R. E., Zhao, H., Oetting, W. S., Austin, L. M., Wildenberg, S. C., Boissy, Y. L. et al. Mutation in and lack of expression of tryrosinase-related protein-1 (TRP-1) in melanocytes from an individual with brown oculocutaneous albinism: a new subtype of albinism classified as “OCA3”. Am. J. Hum. Genet. 58, 1145–1156 (1996).
Newton, J. M., Cohen-Barak, O., Hagiwara, N., Gardner, J. M., Davisson, M. T., King, R. A. et al. Mutations in the human orthologue of the mouse underwhite gene (uw) underlie a new form of oculocutaneous albinism, OCA4. Am. J. Hum. Genet. 69, 981–988 (2001).
Simeonov, D. R., Wang, X., Wang, C., Sergeev, Y., Dolinska, M., Bower, M. et al. DNA variations in oculocutaneous albinism: an updated mutation list and current outstanding issues in molecular diagnostics. Hum. Mutat. 34, 827–835 (2013).
Kausar, T., Bhatti, M. A., Ali, M., Shaikh, R. S. & Ahmed, Z. M. OCA5, a novel locus for non-syndromic oculocutaneous albinism, maps to chromosome 4q24. Clin. Genet. 84, 91–93 (2013).
Wei, A. H., Zang, D. J., Zhang, Z., Liu, X. Z., He, X., Yang, L. et al. Exome sequencing identifies SLC24A5 as a candidate gene for nonsyndromic oculocutaneous albinism. J. Invest. Dermatol. 133, 1834–1840 (2013).
Grønskov, K., Dooley, C. M., Ostergaard, E., Kelsh, R. N., Hansen, L., Levesque, M. P. et al. Mutations in C10orf11, a melanocyte-differentiation gene, cause autosomal-recessive albinism. Am. J. Hum. Genet. 92, 415–421 (2013).
Montoliu, L., Grønskov, K., Wei, A. H., Martínez-García, M., Fernández, A., Arveiler, B. et al. Increasing the complexity: new genes and new types of albinism. Pigment Cell Melanoma Res. 27, 11–18 (2014).
Morice-Picard, F., Lasseaux, E., François, S., Simon, D., Rooryck, C., Bieth, E. et al. SLC24A5 mutations are associated with non-syndromic oculocutaneous albinism. J. Invest. Dermatol. 134, 568–571 (2014).
Schiaffino, M. V., Bassi, M. T., Galli, L., Renieri, A., Bruttini, M., De Nigris, F. et al. Analysis of the OA1 gene reveals mutations in only one third of patients with X-linked ocular albinism. Hum. Mol. Genet. 4, 2319–2325 (1995).
Mayeur, H., Roche, O., Vêtu, C., Jaliffa, C., Marchant, D., Dollfus, H. et al. Eight previously unidentified mutations found in the OA1 ocular albinism gene. BMC Med. Genet. 7, 41 (2006).
Jannot, A. S., Meziani, R., Bertrand, G., Gerard, B., Descamps, V., Archimbaud, A. et al. Allele variations in the OCA2 gene (pink-eyed-dilution locus) are associated with genetic susceptibility to melanoma. Eur. J. Hum. Genet. 13, 913–920 (2005).
Ibarrola-Villava, M., Hu, H. H., Guedj, M., Fernandez, L. P., Descamps, V., Basset-Seguin, N. et al. MC1R, SLC45A2 and TYR genetic variants involved in melanoma susceptibility in southern European populations: results from a meta-analysis. Eur. J. Cancer 48, 2183–2191 (2012).
Hayashi, H., Sone, M., Schachern, P. A., Wakamatsu, K., Paparella, M. M. & Nakashima, T. Comparison of the quality of cochlear melanin in young and old C57BL/6 mice. Arch. Otolaryngol. Head Neck Surg. 133, 151–154 (2007).
Ohlemiller, K. K. & Gagnon, P. M. Genetic dependence of cochlear cells and structures injured by noise. Hear. Res. 224, 34–50 (2007).
Chaki, M., Mukhopadhyay, A. & Ray, K. Determination of variants in the 3'-region of the tyrosinase gene requires locus specific amplification. Hum. Mutat. 26, 53–58 (2005).
Durham-Pierre, D., Gardner, J. M., Nakatsu, Y., King, R. A., Francke, U., Ching, A. et al. African origin of an intragenic deletion of the human P gene in tyrosinase positive oculocutaneous albinism. Nat. Genet. 7, 176–179 (1994).
Wildeman, M., van Ophuizen, E., den Dunnen, J. T. & Taschner, P. E. Improving sequence variant descriptions in mutation databases and literature using the Mutalyzer sequence variation nomenclature checker. Hum. Mutat. 29, 6–13 (2008).
Schwarz, J. M., Cooper, D. N., Schuelke, M. & Seelow, D. MutationTaster2: mutation prediction for the deep-sequencing age. Nat. Methods 11, 361–362 (2014).
Popp, M. W. & Maquat, L. E. Organizing principles of mammalian nonsense-mediated mRNA decay. Annu. Rev. Genet. 47, 139–165 (2013).
Spritz, R. A., Ho, L., Furumura, M. & Hearing, V. J. Mutational analysis of copper binding by human tyrosinase. J. Invest. Dermatol. 109, 207–212 (1997).
Tripathi, R. K., Giebel, L. B., Strunk, K. M. & Spritz, R. A. A polymorphism of the human tyrosinase gene is associated with temperature-sensitive enzymatic activity. Gene Expr. 1, 103–110 (1991).
Oetting, W. S., Pietsch, J., Brott, M. J., Savage, S., Fryer, J. P., Summers, C. G. et al. The R402Q tyrosinase variant does not cause autosomal recessive ocular albinism. Am. J. Med. Genet. A 149A, 466–469 (2009).
Preising, M. N., Forster, H., Gonser, M. & Lorenz, B. Screening of TYR, OCA2, GPR143, and MC1R in patients with congenital nystagmus, macular hypoplasia, and fundus hypopigmentation indicating albinism. Mol. Vis. 17, 939–948 (2011).
Hutton, S. M. & Spritz, R. A. Comprehensive analysis of oculocutaneous albinism among non-Hispanic Caucasians shows that OCA1 is the most prevalent OCA type. J. Invest. Dermatol. 128, 2442–2450 (2008).
Gargiulo, A., Testa, F., Rossi, S., Di Iorio, V., Fecarotta, S., de Berardinis, T et al. Molecular and clinical characterization of albinism in a large cohort of Italian patients. Invest. Ophthalmol. Vis. Sci. 52, 1281–1289 (2011).
Chiang, P. W., Spector, E. & Tsai, A. C. Oculocutaneous albinism spectrum. Am. J. Med. Genet. A 149A, 1590–1591 (2009).
Rebbeck, T. R., Kanetsky, P. A., Walker, A. H., Holmes, R., Halpern, A. C., Schuchter, L. M. et al. P gene as an inherited biomarker of human eye color. Cancer Epidemiol. Biomarkers Prev. 11, 782–784 (2002).
Kamaraj, B. & Purohit, R. Computational screening of disease-associated mutations in OCA2 gene. Cell Biochem. Biophys. 68, 97–109 (2014).
Rimoldi, V., Straniero, L., Asselta, R., Mauri, L., Manfredini, E., Penco, S. et al. Functional characterization of two novel splicing mutations in the OCA2 gene associated with oculocutaneous albinism type II. Gene 537, 79–84 (2014).
Wei, A. H., Yang, X. M., Lian, S. & Li, W. Genetic analyses of Chinese patients with digenic oculocutaneous albinism. Chin. Med. J. (Engl.) 126, 226–230 (2013).
Rooryck, C., Morice-Picard, F., Elçioglu, N. H., Lacombe, D., Taieb, A. & Arveiler, B. Molecular diagnosis of oculocutaneous albinism: new mutations in the OCA1-4 genes and practical aspects. Pigment Cell Melanoma Res. 21, 583–587 (2008).
Grønskov, K., Brondum-Nielsen, K., Birgit, L. & Preising, M. N. Clinical utility gene card for: Oculocutaneous albinism. Eur. J. Hum. Genet. 12, 22 (2014).
Toyofuku, K., Wada, I., Valencia, J. C., Kushimoto, T., Ferrans, V. J. & Hearing, V. J. Oculocutaneous albinism types 1 and 3 are ER retention diseases: mutation of tyrosinase or Tyrp1 can affect the processing of both mutant and wild-type proteins. FASEB J. 15, 2149–2161 (2001).
Mauri, L., Barone, L., Al Oum, M., Del Longo, A., Piozzi, E., Manfredini, E. et al. SLC45A2 mutation frequency in oculocutaneous albinism Italian patients doesn't differ from other European studies. Gene 533, 398–402 (2014).
Veniani, E., Mauri, L., Manfredini, E., Gesu, G. P., Patrosso, M. C., Zelante, L. et al. Detection of the first OCA6 Italian patient in a large cohort of albino subjects. Dermatol. Sci. 81, 208–209.
D'Addio, M., Pizzigoni, A., Bassi, M. T., Baschirotto, C., Valetti, C., Incerti, B. et al. Defective intracellular transport and processing of OA1 is a major cause of ocular albinism type 1. Hum. Mol. Genet. 9, 3011–3018 (2000).
Regales, L., Giraldo, P., García-Díaz, A., Lavado, A & Montoliu, L Identification and functional validation of a 5' upstream regulatory sequence in the human tyrosinase gene homologous to the locus control region of the mouse tyrosinase gene. Pigment Cell Res. 16, 685–692 (2003).
Desmet, F. O. & Beroud, C. Bioinformatics and mutations leading to exon skipping. Methods Mol. Biol. 867, 17–35 (2012).
Straniero, L., Rimoldi, V., Soldà, G., Mauri, L., Manfredini, E., Andreucci, E. et al. Two novel splicing mutations in the SLC45A2 gene cause oculocutaneous albinism type IV by unmasking cryptic splice sites. J. Hum. Genet. 60, 467–471 (2015).
Rooryck, C., Morice-Picard, F., Lasseaux, E., Cailley, D., Dollfus, H., Defoort-Dhellemme, S. et al. High resolution mapping of OCA2 intragenic rearrangements and identification of a founder effect associated with a deletion in Polish albino patients. Hum. Genet. 129, 199–208 (2011).
Morice-Picard, F., Lasseaux, E., Cailley, D., Gros, A., Toutain, J., Plaisant, C. et al. High-resolution array-CGH in patients with oculocutaneous albinism identifies new deletions of the TYR, OCA2, and SLC45A2 genes and a complex rearrangement of the OCA2 gene. Pigment Cell Melanoma Res. 27, 59–71 (2014).
Gill, S. S. & Salt, A. N. Quantitative differences in endolymphatic calcium and endocochlear potential between pigmented and albino guinea pigs. Hear. Res. 113, 191–197 (1997).
Lahne, M. & Gale, J. E. Damage-induced activation of ERK1/2 in cochlear supporting cells is a hair cell death-promoting signal that depends on extracellular ATP and calcium. J. Neurosci. 28, 4918–4928 (2008).
Murillo-Cuesta, S., Contreras, J., Zurita, E., Cediel, R., Cantero, M., Varela-Nieto, I. et al. Melanin precursors prevent premature age-related and noise-induced hearing loss in albino mice. Pigment Cell Melanoma Res. 23, 72–83 (2010).
Acknowledgements
We thank Clinicians and Geneticists who contributed to the cohort of albino patients: Spaccini L, Medical Genetics, Obstetrics and Gynecology Department, Children’s Hospital V Buzzi, Milano, Italy; Mencarelli MA, Pollazzon M, Renieri A, Department of Medical Genetics, Siena University, Italy; Digilio C, Division of Medical Genetics, IRCCS Bambin Gesù Children’s Hospital, Roma, Italy; Guerrieri F, CNLS–DMISM, Sapienza University, Roma, Italy; Vignola S, Divizia MT, Medical Genetics Unit, Institute G Gaslini, Genova, Italy; De Toni T, Department of Pediatrics, Genova University, Italy; Papa A, Department of Neuropediatrics, Novara Hospital, Italy; Piccione M, UO Medical Genetics, AOU Policlinico, Palermo, Italy; Zelante L, Medical Genetics Unit, IRCCS Casa Sollievo Della Sofferenza, San Giovanni Rotondo, Italy; Marelli S, IRCCS Eugenio Medea, Bosisio Parini, Italy; Naretto G, Neuroscience Department, and Silengo M, Marinosci A, Department of Pediatrics, Torino University, Italy; Grosso E, SCDU Medical Genetics, AOS Giovanni Battista, Torino, Italy; Gasparini P, Institute for Maternal and Child Health, IRCCS ‘Burlo Garofolo’, Trieste, Italy; Tenconi R, Medical Genetics, Department of Pediatrics, Padova, Italy; Sensi A, Medical Genetics, Ferrara University, Italy; Rocchetti L, UO Medical Genetics AV Romagna, Italy; Garavelli L, Clinical Genetics Unit, S Maria Nuova Hospital, Reggio Emilia, Italy; Toschi B, UO Medical Genetics, AOU Pisana, Pisa, Italy; Prontera P, Medical Genetics Unit, AOU Perugia, Italy; Piazzini M, Department of Medical Genetics, Firenze University, Italy; Lapi E, Romano S, Genetics and Molecular Medicine Unit, Meyer Children’s University Hospital, Firenze, Italy; Pelo E, SOD Diagnostica Genetica, and Sodi A, Department of Specialized Surgical Sciences, Eye Clinic, AOU Careggi Hospital, Florence, Italy; Melis D, Department of Pediatrics, and Simonelli F, Multidisciplinary Department, Eye Clinic, AOU Federico II, Napoli, Italy; Maggio PP, UOC Cytogenetics and Molecular Genetics Unit, Sacro Cuore Hospital, Gallipoli, Italy; Di Stefano C, TIN, PO Umberto I, Nocera Inferiore, Italy; Bianca S, Medical Genetics Unit, ARNAS Garibaldi, Catania, Italy; Mandarà GML, UO Medical Genetics, ASP Ragusa, Italy; Kurtozg S Afyon Kocatepe University, Department of Medical Genetics, Afyonkarahisar, Turkey. We thank patients and relatives for their contributions to this study, as without them this research would be impossible. This work was supported by a Lombardy Region Grant Independent Research Project No. 13848 of 11-12-2009.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Rights and permissions
About this article
Cite this article
Mauri, L., Manfredini, E., Del Longo, A. et al. Clinical evaluation and molecular screening of a large consecutive series of albino patients. J Hum Genet 62, 277–290 (2017). https://doi.org/10.1038/jhg.2016.123
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/jhg.2016.123
This article is cited by
-
The experience of albinism in France: a qualitative study on dyads of parents and their adult child with albinism
BMC Medicine (2024)
-
The contribution of common regulatory and protein-coding TYR variants to the genetic architecture of albinism
Nature Communications (2022)
-
Evidence that the Ser192Tyr/Arg402Gln in cis Tyrosinase gene haplotype is a disease-causing allele in oculocutaneous albinism type 1B (OCA1B)
npj Genomic Medicine (2022)
-
Evident hypopigmentation without other ocular deficits in Dutch patients with oculocutaneous albinism type 4
Scientific Reports (2021)
-
Two Novel Homozygous Mutations in Phosphoglucomutase 3 Leading to Severe Combined Immunodeficiency, Skeletal Dysplasia, and Malformations
Journal of Clinical Immunology (2021)