Abstract
The tumour suppressor p53 plays an important role in somatic cell reprogramming. While wild-type p53 reduces reprogramming efficiency, mutant p53 exerts a gain of function activity that leads to increased reprogramming efficiency. Furthermore, induced pluripotent stem cells expressing mutant p53 lose their pluripotency in vivo and form malignant tumours when injected in mice. It is therefore of great interest to identify targets of p53 (wild type and mutant) that are responsible for this phenotype during reprogramming, as these could be exploited for therapeutic use, that is, formation of induced pluripotent stem cells with high reprogramming efficiency, but no oncogenic potential. Here we studied the transcriptional changes of microRNA in a series of mouse embryonic fibroblasts that have undergone transition to induced pluripotent stem cells with wild type, knock out or mutant p53 status in order to identify microRNAs whose expression during reprogramming is dependent on p53. We identified a number of microRNAs, with known functions in differentiation and carcinogenesis, the expression of which was dependent on the p53 status of the cells. Furthermore, we detected several uncharacterised microRNAs that were regulated differentially in the different p53 backgrounds, suggesting a novel role of these microRNAs in reprogramming and pluripotency.
Similar content being viewed by others
Main
The tumour suppressor p53 is the most frequently mutated or deregulated gene in human cancers.1, 2, 3, 4, 5, 6, 7 Often referred to as the guardian of the genome, its role in protecting the cell from accumulation of DNA damage by inducing DNA repair or cell death is well-studied.8, 9, 10, 11, 12 However, p53 has also been implicated in a vast variety of other cell pathways, including metabolism,13 autophagy,14, 15 mitochondrial function16, 17, 18 and also cell differentiation and pluripotency.19, 20 Interestingly, p53 mutations, in addition to disrupting the protein's wild-type function, result in additional activities that lead to increased tumour malignancy, usually referred to as gain of function (GOF).21, 22
Recently, p53 is emerging as a key regulator in the process of reprogramming from somatic to induced pluripotent stem (iPS) cells as well as being involved in stem cell maintenance.23, 24, 25, 26, 27, 28, 29, 30 Stem cells are characterised by high genomic stability, which is crucial to minimise tumorigenesis following stem cell expansion.31, 32, 33 p53 is an important factor that protects this genomic integrity and has the ability to counteract somatic reprogramming by inducing cell cycle arrest and apoptosis.23, 25, 26, 34, 35, 36 In contrast to somatic cells, p53 does not induce apoptosis in embryonic stem cells (ESCs) following DNA damage, but promotes differentiation of ESC by several mechanisms including transcriptional repression of the pluripotency factors Nanog and Oct4.37, 38, 39, 40 After differentiation p53 activates the expression of genes that lead to cell death or senescence by classical p53 pathways. Thus, p53 plays an important role in maintaining a pool of stem cells with an intact genome and moreover prevents of reprogramming cells with faulty genome.27
We have previously studied the reprogramming efficiency of a series of MEFs with different p53 status, that is, p53 wt, p53 knock out (KO) and mutant p53R172H cells.27 p53R172H (R175H in human) is a conformational mutant that results in a misfolded p53 protein. This study showed that p53 depletion or the expression mutant p53 increases reprogramming efficiency.27 However, cells expressing p53R172H in addition to their augmented pluripotency in vitro exhibited carcinogenic potential in vivo. When injected into nude mice, p53R172H expressing iPS cells lost their differentiation capacity and gave rise to aggressive sarcomas, while p53 KO iPS cells maintained pluripotency and led to the formation of benign teratomas, thus displaying a novel GOF for mutant p53.27
It is of great interest to generate iPS cells with a high reprogramming efficiency, but low tumorigenic potential for therapeutic use. As p53 was shown to be important in both reprogramming and maintaining genomic integrity of iPS cell, it provides an interesting target for manipulation of the reprogramming pathway. It is thus of interest to dissect the mechanisms and players regulated by p53 in these pathways. In addition to controlling the expression of protein coding genes, p53 was shown to control the transcription of a number of microRNAs (miRNAs). Expression of miRNAs is altered in many pathological conditions including cancer, where different miRNAs exhibit oncogenic and tumour suppressive properties. Moreover, miRNAs are key regulators of development; for example, miR-34a is fundamental for neuronal and muscle differentiation,41, 42, 43 but also influence reprogramming of stem cells and the maintenance of an undifferentiated cellular stage.44, 45
In this study, we set out to examine miRNAs that are differentially regulated in cells during reprogramming depending on their p53 status, aiming to identify miRNAs that play a role in this process and that could be directly targeted to help optimise iPS cells. This would allow the generation of cells that have intact p53, which protects their genomic integrity, but at the same time exhibit high reprogramming efficiency. To this end, we performed a microarray screening of miRNA expression before and after three factors driven reprogramming of wt, KO and mutant p53 cells and identified several miRNAs whose expression is dependent on the p53 status of the cell.
Results
Identification of microRNAs that are modulated during the MEF to iPS cell transition depending on cell's p53 status
To identify miRNAs that are targeted by either wt or mutant p53 during reprogramming, we performed microarray analysis on a series of iPS cells and their parental MEFs with different p53 (wt, p53, KO and p53R172H). The MEFs were reprogrammed by introduction of Oct4, Sox2, Klf4 as described in our previous study by Sarig et al.27
All Cq values above 32 were considered noise background and excluded from the analysis. The results for a given sample were normalised by the geometric mean of the relative quantities of all targets expressed in the same sample (global mean procedure). Targets that were up- or downregulated more than twofold were considered to be changed in the analysis.
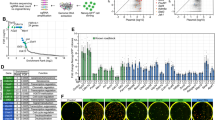
In a first step we looked at the miRNAs that were up- or downregulated in all three conditions during reprogramming (Figure 1). A high number of the detected miRNAs are known to play a role in stemness and differentiation, that is, let7 family, miR-125b, miR-126, miR-136, miR-143, miR-145, miR-152, suggesting that we indeed identified miRNAs that are important in reprogramming using our experimental set up.
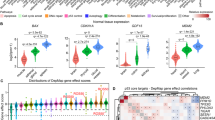
A number of miRNAs involved in differentiation are up/downregulated during the MEF to iPS cell transition independent of the cell's p53 status. MiRNAs levels were examined upon transition from MEFs to iPS cell using microarray analysis. Selected miRNAs that were induced (a) or decreased (b) in all genetic backgrounds, that is, p53 wt, p53R172H and p53 KO (NRQ=normalised relative quantities)
Next, to study the role of p53, both wt and mutant, during reprogramming, we were interested in miRNAs that were specifically regulated depending on the cell's p53 status (see Supplementary Table S1 for summary of relevant miRNAs identified). Figure 2a shows those that were upregulated specifically by wt p53 in iPS cells, while not significantly changed in the other conditions. Of these, miR-200c was previously shown to be a transcriptional target of p53.46, 47 Furthermore, several of these miRNAs have been implicated in both stemness and cancer (Supplementary Table S1). MiR-182, for example, is an important factor in the development of the inner ear and retina, T-cell development and osteogenesis and has also been implicated in cancer development and metastasis,48 while miR-497 is a tumour suppressor that has been shown to induce quiescence in skeletal muscle stem cells.49, 50, 51
Several miRNAs are regulated exclusively in p53 wt cells. MiRNAs levels were examined upon transition from MEFs to iPS cells using microarray analysis. Selected miRNAs that were induced exclusively in p53 wt cells (a), only detected in p53 wt cells (b) and downregulated in p53 wt cells (c) are shown
Additionally, we identified a number of small non-coding RNAs that were only detectable in iPS cells bearing functional p53 (Figure 2b), most of which have not yet been characterised. Furthermore, we identified two miRNAs, miR-27a, see also refs 52, 53, 54, and miR-33 that were downregulated following reprogramming exclusively in p53 wt cells (Figure 2c).
Both p53 KO and p53 mutant cells displayed increased reprogramming efficiency compared with wt p53, notably p53 mutant cells underwent reprogramming more effectively than p53 KO cells. We were thus interested in hits that were regulated in these conditions and found several miRNA that are downregulated (Figure 3a) or upregulated (Figure 3b) in p53 KO and mutant cells, but not changed in p53 wt cells. For example, we found that p53 KO and p53 mutant iPS cells induced the expression of miR-186. Furthermore, a high number of miRNAs that were downregulated in p53 compromised iPS cells convey tumour suppressive functions, that is, miR-30a-5p,55 miR-31,56, 57 miR-335,58 miR-38259 and miR-503.60, 61
Levels of some miRNAs change solely in p53R172H and p53 KO cells during the MEF to iPS cell transition. Analysis of miRNAs levels upon transition from MEFs to iPS cells highlighted miRNAs that were regulated differentially depending on the cell's p53 status. Here miRNAs that were induced (a) or decreased (b) in p53R172H and p53 KO cells, while unchanged in a p53 wt background are shown
Mutant p53 regulates a specific pool of microRNAs during reprogramming that might be linked to its GOF activity
In previous studies we demonstrated a GOF activity of mutant p53 in reprogramming, manifested not only by more efficient process, but also by acquiring in cells with tumorigenic properties.27 We were therefore particularly interested to identify miRNA that are specifically modulated in p53R172H cells during MEF to iPS cell transition. We identified several miRNAs that are strongly upregulated (Figure 4a) or downregulated (Figure 4b) upon reprogramming in p53R172H cells, while unchanged or less strongly regulated in the other samples. As mutant p53 cells have a high reprogramming efficiency, these miRNAs could facilitate the reprograming process and it would be interesting to study their role in more detail. Indeed several of the miRNA regulated by mutant p53 during transition from MEF to iPS cells have been implicated with stemness as well as carcinogenesis (Supplementary Table S1). For example, miR-194 and miR-206 were previously found to be involved in osteoblast62 and muscle differentiation,63 respectively, while miR-101, whose expression was decreased in p53R172H iPS cells, is a well-characterised tumour suppressor that inhibits tumour growth and metastasis.64, 65, 66, 67, 68
Complex regulation of miRNAs during the induced transition from MEF to iPS cells depending on p53. Analysis of miRNAs levels upon transition from MEFs to iPS cell revealed a group of miRNAs that was upregulated (a) or downregulated (b) only in p53R172H iPS cell, while unchanged in the other groups, and miRNAs whose expression was induced in p53 wt while reduced in KO or p53 mutant cells (c) or vice versa (d)
Reversely regulated microRNAs in p53 wt and p53 mutant cells
Interestingly, we identified several miRNAs whose expression was opposite in p53 wt and p53 mutant cells, that is, increased expression during reprogramming in wt cells, yet decreased in mutant cells or vice versa (Figures 4c and d). This suggests that these miRNAs are regulated by p53 during reprogramming by complex mechanisms. An example is miR-10a that on the one hand is known to promote the differentiation of human mesenchymal stem cells69 and on the other contributes to cancer development.70, 71 This miR is downregulated during reprogramming of p53 wt cells, but upregulated in cells expressing mutant p53. Mir-199b shows upregulation in p53R172H iPS. The same is true for miR-218, an miRNA that seems to acts as a tumour suppressor72, 73 and promotes stem cell differentiation.74, 75, 76 The tumour suppressive miR-708 and miR-126 on the other hand are upregulated during reprogramming in p53 wt cells and reduced in p53R172H cells.
MicroRNAs specifically upregulated in WT p53 iPS cell encode for a p53 responsive element
To further characterise the miRNA that we identified as upregulated solely in p53 wt cells, we decided to screen for predicted responsive elements in the promoter region of those miRNAs that were induced upon reprogramming in the presence of wt p53 (Figure 2). We identified four miRNAs that encode a p53 responsive element with high matrix similarity in their promoter region (Supplementary Table S2). Of all the targets that were specifically upregulated during reprogramming of wt cells, only miR-142-3p lacked a predicted p53 responsive element in its promoter altogether.
Next, we investigated whether any of the miRNAs upregulated specifically in p53 wt cells are predicted to target genes known to be important for reprogramming and pluripotency. While we did not identify any of the key regulators of reprogramming, such as Nanog, Sox2, Oct4/3, ERK2 or β-catenin/Ctnnb1 as predicted targets, our predictive analysis highlighted factors, for example, N-Myc, Akt1/2/3 and Smad 2/2, involved in iPS cell generation and pluripotency that encode a responsive element in their 3′UTR for one or several of the miRNAs that we identified in our screen (Supplementary Table S3).
Discussion
In this study we used our previously published iPS cell model27 to investigate miRNAs that are under control of p53, wild type or mutant during reprogramming. The fact that p53 mutant cells exhibit an increased reprogramming efficiency, while p53 wt iPS cells have high genomic stability and thereby low carcinogenic potential, makes this transcription factor and its targets of great interest in the aim to optimise pluripotent stem cell formation.
Using a microarray approach we were able to identify several miRNAs that are specifically regulated by p53 during the process of reprogramming. Interestingly, we did not only find miRNAs upregulated by wt p53, but also specifically by p53R172H. We have previously shown that this p53 mutant exhibits a GOF activity during reprogramming27 and it would be very interesting to further investigate whether these miRNAs are responsible for this phenotype. Among the miRNAs that are regulated by p53R172H during the MEF to iPS cell transition, several have known functions in stemness and differentiation, for example miR-186, miR-194 and miR-206. Additionally, several miRNAs that are known to exhibit tumour suppressive functions, like miR-30a-5p, miR-31, miR-335, miR-382 and miR-503, were downregulated in the p53R172H cells upon reprograming to iPS cells. This is in line with our earlier study that showed occurrence of malignant tumours after injection of iPS cells with p53R172H in mice.27 On the other hand, we also identified tumour suppressive miRNAs, like miR-218 that are upregulated in p53 mutant cells, and in the case of miR-218 downregulated in p53 wt cells. The roles of these miRs in the reprogrammed cells remains to be studied.
Interestingly, miR-15b that was markedly induced exclusively by mutant p53 during reprogramming. Mir-15b promotes osteoblast differentiation77 whilst it reduces invasion and metastasis78, 79 and its deletion was shown to promote B-cell malignancies;80 these distinct effects suggests a possible complex context-dependent function. Furthermore, we detected a decrease of miR-155 in both p53 KO and mutant cells during reprogramming, while its levels remained high in p53 wt cells, which is at variance with previous studies.81,82
Notably, we also identified miRNAs that are induced in p53 KO and p53 mutant iPS cells, but not in p53 wt iPS cells, suggesting that wt p53 inhibits their up-regulation during reprogramming. It remains to be investigated how these miRNAs effect differentiation and/or oncogenesis in vivo.
We identified several miRNAs that are increased specifically in p53 wt cells during reprogramming from MEF to iPS cells. These miRNAs could be involved in ensuring high genome stability of reprogrammed cells. MiR-709,83 for example, is upregulated specifically in the transition from p53 wt MEFs to iPS cells, while it is downregulated when p53 is not functional. Another interesting hit that is strongly upregulated in p53 wt iPS cells, but very low expressed in p53 KO and mutant iPS cells is miR-142-3p.84,85 We were unable to identify whether a p53 responsive element is its promoter, suggesting that p53 regulates the expression of this miRNA by an indirect mechanism.
Of note, our microarray screening did not highlight miR-34a as upregulated during reprogramming, as previously reported.86, 87, 88 We think this could be due to the fact that in the previous work, iPS cells were generated with four reprogramming factors (Oct4, Sox2, Klf4 and c-Myc), while in this study the iPS cells were generated using only three reprogramming factors (Oct4, Sox2 and Klf4) since c-Myc has been correlated with carcinogenesis of iPS cells in vivo.89
In conclusion, our study highlighted a group of miRNAs driven by wt and mutant p53 that might be important for reprogramming, tumorigenesis and loss of genomic integrity. While a high number of miRNAs identified in our screen have already been associated with cancer development or differentiation and stemness (Supplementary Table S1), others are less well characterised and provide interesting targets to study in more details. It would be of particular interest to dissect their role during reprogramming and how this is controlled by p53 in either its wild type or mutant form. Thus, further studies will be necessary to further validate these results and pinpoint the exact functions of these miRNAs during the formation of iPS cells.
Materials and Methods
Cell culture and iPS cell generation
Primary MEFs were prepared from wt, p53 mutant or p53 KO E13.5 embryos as reported previously.27 Briefly, MEFs were prepared from 13.5 days postcoitum embryos and maintained in DME, supplemented with 10% FCS, 1 mM sodium pyruvate, 2 mM l-glutamine, 0.1 mM nonessential amino acids, 0.1 mM β-mercaptoethanol and antibiotics. Reprogramming was induced using the three factor protocol (Oct4, Sox2, Klf4) as described previously.27
RNA extraction and microarray analysis
RNA was extracted using Trizol (Invitrogen, USA), following manufacturer’s instruction.
Microarray analysis was performed by Biogazelle (Gent, Belgium). All Cq values above 32 were excluded as noise background. The results for a given sample were normalised by the geometric mean of the relative quantities of all targets that are expressed in that sample (global mean normalisation procedure). Three biological samples (n=3) were collected and pooled together for technical reasons; these were analysed with three technical replicates, obtaining always a mean error lower than 5% of the measure. Because this variation derives from a technical measure, and not from a biological replicate, it has not been reported on the histograms. Therefore, we did not show any statistics on these qualitative results.
Predictive analysis
P53 responsive element in microRNAs promoters were predicted using the programme MatInspector that allows identification of transcription factors binding sites in nucleotide sequences using a large library of weight matrices (Genomatix, Germany).
MicroRNAs targets were predicted by using the miRanda Software (microrna.org). Only target sites of conserved miRNAs and with good mirSVR score were taken into consideration.
References
Soussi T, Wiman KG . TP53: an oncogene in disguise. Cell Death Differ 2015; 22: 1239–1249.
Zhang B, Rotelli M, Dixon M, Calvi BR . The function of Drosophila p53 isoforms in apoptosis. Cell Death Differ 2015; 22: 2058–2067.
Kroemer G, Bravo-San Pedro JM, Galluzzi L . Novel function of cytoplasmic p53 at the interface between mitochondria and the endoplasmic reticulum. Cell Death Dis 2015; 6: e1698.
Laezza C, D'Alessandro A, Di Croce L, Picardi P, Ciaglia E, Pisanti S et al. p53 regulates the mevalonate pathway in human glioblastoma multiforme. Cell Death Dis 2015; 6: e1909.
Puzio-Kuter AM, Laddha SV, Castillo-Martin M, Sun Y, Cordon-Cardo C, Chan CS et al. Involvement of tumor suppressors PTEN and p53 in the formation of multiple subtypes of liposarcoma. Cell Death Differ 2015; 22: 1785–1791.
Slatter TL, Hung N, Bowie S, Campbell H, Rubio C, Speidel D et al. Delta122p53, a mouse model of Delta133p53alpha, enhances the tumor-suppressor activities of an attenuated p53 mutant. Cell Death Dis 2015; 6: e1783.
te Raa GD, Moerland PD, Leeksma AC, Derks IA, Yigittop H, Laddach N et al. Assessment of p53 and ATM functionality in chronic lymphocytic leukemia by multiplex ligation-dependent probe amplification. Cell Death Dis 2015; 6: e1852.
Dashzeveg N, Taira N, Lu ZG, Kimura J, Yoshida K . Palmdelphin, a novel target of p53 with Ser46 phosphorylation, controls cell death in response to DNA damage. Cell Death Dis 2014; 5: e1221.
Manzl C, Fava LL, Krumschnabel G, Peintner L, Tanzer MC, Soratroi C et al. Death of p53-defective cells triggered by forced mitotic entry in the presence of DNA damage is not uniquely dependent on Caspase-2 or the PIDDosome. Cell Death Dis 2013; 4: e942.
Nair BC, Krishnan SR, Sareddy GR, Mann M, Xu B, Natarajan M et al. Proline, glutamic acid and leucine-rich protein-1 is essential for optimal p53-mediated DNA damage response. Cell Death Differ 2014; 21: 1409–1418.
Phesse TJ, Myant KB, Cole AM, Ridgway RA, Pearson H, Muncan V et al. Endogenous c-Myc is essential for p53-induced apoptosis in response to DNA damage in vivo. Cell Death Differ 2014; 21: 956–966.
Levine AJ, Oren M . The first 30 years of p53: growing ever more complex. Nat Rev Cancer 2009; 9: 749–758.
Kruiswijk F, Labuschagne CF, Vousden KH . p53 in survival, death and metabolic health: a lifeguard with a licence to kill. Nat Rev Mol Cell Biol 2015; 16: 393–405.
Garufi A, Pucci D, D'Orazi V, Cirone M, Bossi G, Avantaggiati ML et al. Degradation of mutant p53H175 protein by Zn(II) through autophagy. Cell Death Dis 2014; 5: e1271.
Simon HU, Yousefi S, Schmid I, Friis R . ATG5 can regulate p53 expression and activation. Cell Death Dis 2014; 5: e1339.
Ci Y, Shi K, An J, Yang Y, Hui K, Wu P et al. ROS inhibit autophagy by downregulating ULK1 mediated by the phosphorylation of p53 in selenite-treated NB4 cells. Cell Death Dis 2014; 5: e1542.
Evstafieva AG, Garaeva AA, Khutornenko AA, Klepikova AV, Logacheva MD, Penin AA et al. A sustained deficiency of mitochondrial respiratory complex III induces an apoptotic cell death through the p53-mediated inhibition of pro-survival activities of the activating transcription factor 4. Cell Death Dis 2014; 5: e1511.
Xu J, Wang J, Hu Y, Qian J, Xu B, Chen H et al. Unequal prognostic potentials of p53 gain-of-function mutations in human cancers associate with drug-metabolizing activity. Cell Death Dis 2014; 5: e1108.
Aloni-Grinstein R, Shetzer Y, Kaufman T, Rotter V . p53: the barrier to cancer stem cell formation. FEBS Lett 2014; 588: 2580–2589.
Rivlin N, Koifman G, Rotter V . p53 orchestrates between normal differentiation and cancer. Semin Cancer Biol 2015; 32: 10–17.
Brosh R, Rotter V . When mutants gain new powers: news from the mutant p53 field. Nat Rev Cancer 2009; 9: 701–713.
Zhao Y, Zhang C, Yue X, Li X, Liu J, Yu H et al. Pontin, a new mutant p53-binding protein, promotes gain-of-function of mutant p53. Cell Death Differ 2015; 22: 1824–1836.
Hong H, Takahashi K, Ichisaka T, Aoi T, Kanagawa O, Nakagawa M et al. Suppression of induced pluripotent stem cell generation by the p53-p21 pathway. Nature 2009; 460: 1132–1135.
Kanatsu-Shinohara M, Inoue K, Lee J, Yoshimoto M, Ogonuki N, Miki H et al. Generation of pluripotent stem cells from neonatal mouse testis. Cell 2004; 119: 1001–1012.
Kawamura T, Suzuki J, Wang YV, Menendez S, Morera LB, Raya A et al. Linking the p53 tumour suppressor pathway to somatic cell reprogramming. Nature 2009; 460: 1140–1144.
Marion RM, Strati K, Li H, Murga M, Blanco R, Ortega S et al. A p53-mediated DNA damage response limits reprogramming to ensure iPS cell genomic integrity. Nature 2009; 460: 1149–1153.
Sarig R, Rivlin N, Brosh R, Bornstein C, Kamer I, Ezra O et al. Mutant p53 facilitates somatic cell reprogramming and augments the malignant potential of reprogrammed cells. J Exp Med 2010; 207: 2127–2140.
Takeshita S, Kikuno R, Tezuka K, Amann E . Osteoblast-specific factor 2: cloning of a putative bone adhesion protein with homology with the insect protein fasciclin I. Biochem J 1993; 294 (Pt 1): 271–278.
Velletri T, Xie N, Wang Y, Huang Y, Yang Q, Chen X et al. P53 functional abnormality in mesenchymal stem cells promotes osteosarcoma development. Cell Death Dis 2016; 7: e2015.
Stedman A, Beck-Cormier S, Le Bouteiller M, Raveux A, Vandormael-Pournin S, Coqueran S et al. Ribosome biogenesis dysfunction leads to p53-mediated apoptosis and goblet cell differentiation of mouse intestinal stem/progenitor cells. Cell Death Differ 2015; 22: 1865–1876.
Di Ruscio A, Patti F, Welner RS, Tenen DG, Amabile G . Multiple sclerosis: getting personal with induced pluripotent stem cells. Cell Death Dis 2015; 6: e1806.
Kumar S, Ingle H, Mishra S, Mahla RS, Kumar A, Kawai T et al. IPS-1 differentially induces TRAIL, BCL2, BIRC3 and PRKCE in type I interferons-dependent and -independent anticancer activity. Cell Death Dis 2015; 6: e1758.
Noguchi H, Saitoh I, Tsugata T, Kataoka H, Watanabe M, Noguchi Y . Induction of tissue-specific stem cells by reprogramming factors, and tissue-specific selection. Cell Death Differ 2015; 22: 145–155.
Utikal J, Polo JM, Stadtfeld M, Maherali N, Kulalert W, Walsh RM et al. Immortalization eliminates a roadblock during cellular reprogramming into iPS cells. Nature 2009; 460: 1145–1148.
Gatzka M, Tasdogan A, Hainzl A, Allies G, Maity P, Wilms C et al. Interplay of H2A deubiquitinase 2A-DUB/Mysm1 and the p19(ARF)/p53 axis in hematopoiesis, early T-cell development and tissue differentiation. Cell Death Differ 2015; 22: 1451–1462.
Park EK, Lee JC, Park JW, Bang SY, Yi SA, Kim BK et al. Transcriptional repression of cancer stem cell marker CD133 by tumor suppressor p53. Cell Death Dis 2015; 6: e1964.
Loh YH, Wu Q, Chew JL, Vega VB, Zhang W, Chen X et al. The Oct4 and Nanog transcription network regulates pluripotency in mouse embryonic stem cells. Nat Genet 2006; 38: 431–440.
Maimets T, Neganova I, Armstrong L, Lako M . Activation of p53 by nutlin leads to rapid differentiation of human embryonic stem cells. Oncogene 2008; 27: 5277–5287.
Mitsui K, Tokuzawa Y, Itoh H, Segawa K, Murakami M, Takahashi K et al. The homeoprotein Nanog is required for maintenance of pluripotency in mouse epiblast and ES cells. Cell 2003; 113: 631–642.
Lin T, Chao C, Saito S, Mazur SJ, Murphy ME, Appella E et al. p53 induces differentiation of mouse embryonic stem cells by suppressing Nanog expression. Nat Cell Biol 2005; 7: 165–171.
Agostini M, Tucci P, Killick R, Candi E, Sayan BS, Rivetti di Val Cervo P et al. Neuronal differentiation by TAp73 is mediated by microRNA-34a regulation of synaptic protein targets. Proc Natl Acad Sci USA 2011; 108: 21093–21098.
Yu X, Zhang L, Wen G, Zhao H, Luong LA, Chen Q et al. Upregulated sirtuin 1 by miRNA-34a is required for smooth muscle cell differentiation from pluripotent stem cells. Cell Death Differ 2015; 22: 1170–1180.
Agostini M, Tucci P, Steinert JR, Shalom-Feuerstein R, Rouleau M, Aberdam D et al. microRNA-34a regulates neurite outgrowth, spinal morphology, and function. Proc Natl Acad Sci USA 2011; 108: 21099–21104.
Park H, Park H, Pak HJ, Yang DY, Kim YH, Choi WJ et al. miR-34a inhibits differentiation of human adipose tissue-derived stem cells by regulating cell cycle and senescence induction. Differentiation 2015; 90: 91–100.
Ng WL, Chen G, Wang M, Wang H, Story M, Shay JW et al. OCT4 as a target of miR-34a stimulates p63 but inhibits p53 to promote human cell transformation. Cell Death Dis 2014; 5: e1024.
Schubert J, Brabletz T . p53 Spreads out further: suppression of EMT and stemness by activating miR-200c expression. Cell Res 2011; 21: 705–707.
Chang CJ, Chao CH, Xia W, Yang JY, Xiong Y, Li CW et al. p53 regulates epithelial-mesenchymal transition and stem cell properties through modulating miRNAs. Nat Cell Biol 2011; 13: 317–323.
Wei Q, Lei R, Hu G . Roles of miR-182 in sensory organ development and cancer. Thorac Cancer 2015; 6: 2–9.
Guo ST, Jiang CC, Wang GP, Li YP, Wang CY, Guo XY et al. MicroRNA-497 targets insulin-like growth factor 1 receptor and has a tumour suppressive role in human colorectal cancer. Oncogene 2013; 32: 1910–1920.
Lan J, Xue Y, Chen H, Zhao S, Wu Z, Fang J et al. Hypoxia-induced miR-497 decreases glioma cell sensitivity to TMZ by inhibiting apoptosis. FEBS Lett 2014; 588: 3333–3339.
Sato T, Yamamoto T, Sehara-Fujisawa A . miR-195/497 induce postnatal quiescence of skeletal muscle stem cells. Nat Commun 2014; 5: 4597.
Ma Y, Yao N, Liu G, Dong L, Liu Y, Zhang M et al. Functional screen reveals essential roles of miR-27a/24 in differentiation of embryonic stem cells. EMBO J 2015; 34: 361–378.
Miao Y, Li J, Qiu X, Li Y, Wang Z, Luan Y . miR-27a regulates the self renewal of the H446 small cell lung cancer cell line in vitro. Oncol Rep 2013; 29: 161–168.
Tang W, Yu F, Yao H, Cui X, Jiao Y, Lin L et al. miR-27a regulates endothelial differentiation of breast cancer stem like cells. Oncogene 2014; 33: 2629–2638.
Baraniskin A, Birkenkamp-Demtroder K, Maghnouj A, Zollner H, Munding J, Klein-Scory S et al. MiR-30a-5p suppresses tumor growth in colon carcinoma by targeting DTL. Carcinogenesis 2012; 33: 732–739.
Bhatnagar N, Li X, Padi SK, Zhang Q, Tang MS, Guo B . Downregulation of miR-205 and miR-31 confers resistance to chemotherapy-induced apoptosis in prostate cancer cells. Cell Death Dis 2010; 1: e105.
Ivanov SV, Goparaju CM, Lopez P, Zavadil J, Toren-Haritan G, Rosenwald S et al. Pro-tumorigenic effects of miR-31 loss in mesothelioma. J Biol Chem 2010; 285: 22809–22817.
Png KJ, Yoshida M, Zhang XH, Shu W, Lee H, Rimner A et al. MicroRNA-335 inhibits tumor reinitiation and is silenced through genetic and epigenetic mechanisms in human breast cancer. Genes Dev 2011; 25: 226–231.
Xu M, Jin H, Xu CX, Sun B, Mao Z, Bi WZ et al. miR-382 inhibits tumor growth and enhance chemosensitivity in osteosarcoma. Oncotarget 2014; 5: 9472–9483.
Sarkar S, Dey BK, Dutta A . MiR-322/424 and -503 are induced during muscle differentiation and promote cell cycle quiescence and differentiation by down-regulation of Cdc25A. Mol Biol Cell 2010; 21: 2138–2149.
Xu YY, Wu HJ, Ma HD, Xu LP, Huo Y, Yin LR . MicroRNA-503 suppresses proliferation and cell-cycle progression of endometrioid endometrial cancer by negatively regulating cyclin D1. FEBS J 2013; 280: 3768–3779.
Li J, He X, Wei W, Zhou X . MicroRNA-194 promotes osteoblast differentiation via downregulating STAT1. Biochem Biophys Res Commun 2015; 460: 482–488.
Kim HK, Lee YS, Sivaprasad U, Malhotra A, Dutta A . Muscle-specific microRNA miR-206 promotes muscle differentiation. J Cell Biol 2006; 174: 677–687.
Wang HJ, Ruan HJ, He XJ, Ma YY, Jiang XT, Xia YJ et al. MicroRNA-101 is down-regulated in gastric cancer and involved in cell migration and invasion. Eur J Cancer 2010; 46: 2295–2303.
Wang L, Zhang LF, Wu J, Xu SJ, Xu YY, Li D et al. IL-1beta-mediated repression of microRNA-101 is crucial for inflammation-promoted lung tumorigenesis. Cancer Res 2014; 74: 4720–4730.
Xiaoping L, Zhibin Y, Wenjuan L, Zeyou W, Gang X, Zhaohui L et al. CPEB1, a histone-modified hypomethylated gene, is regulated by miR-101 and involved in cell senescence in glioma. Cell Death Dis 2013; 4: e675.
Yan F, Shen N, Pang J, Xie D, Deng B, Molina JR et al. Restoration of miR-101 suppresses lung tumorigenesis through inhibition of DNMT3a-dependent DNA methylation. Cell Death Dis 2014; 5: e1413.
Zhang X, Schulz R, Edmunds S, Kruger E, Markert E, Gaedcke J et al. MicroRNA-101 suppresses tumor cell proliferation by acting as an endogenous proteasome inhibitor via targeting the proteasome assembly factor POMP. Mol Cell 2015; 59: 243–257.
Li J, Dong J, Zhang ZH, Zhang DC, You XY, Zhong Y et al. miR-10a restores human mesenchymal stem cell differentiation by repressing KLF4. J Cell Physiol 2013; 228: 2324–2336.
Bryant A, Palma CA, Jayaswal V, Yang YW, Lutherborrow M, Ma DD . miR-10a is aberrantly overexpressed in Nucleophosmin1 mutated acute myeloid leukaemia and its suppression induces cell death. Mol Cancer 2012; 11: 8.
Yan Y, Wang Q, Yan XL, Zhang Y, Li W, Tang F et al. miR-10a controls glioma migration and invasion through regulating epithelial-mesenchymal transition via EphA8. FEBS Lett 2015; 589: 756–765.
Cheng Y, Yang X, Deng X, Zhang X, Li P, Tao J et al. MicroRNA-218 inhibits bladder cancer cell proliferation, migration, and invasion by targeting BMI-1. Tumour Biol 2015; 36: 8015–8023.
Zhu K, Ding H, Wang W, Liao Z, Fu Z, Hong Y et al. Tumor-suppressive miR-218-5p inhibits cancer cell proliferation and migration via EGFR in non-small cell lung cancer. Oncotarget. Oncotarget 2016; 7: 28075–28085.
Gay I, Cavender A, Peto D, Sun Z, Speer A, Cao H et al. Differentiation of human dental stem cells reveals a role for microRNA-218. J Periodontal Res 2014; 49: 110–120.
Hu K, Xu C, Ni H, Xu Z, Wang Y, Xu S et al. Mir-218 contributes to the transformation of 5-Aza/GF induced umbilical cord mesenchymal stem cells into hematopoietic cells through the MITF pathway. Mol Biol Rep 2014; 41: 4803–4816.
Zhang WB, Zhong WJ, Wang L . A signal-amplification circuit between miR-218 and Wnt/beta-catenin signal promotes human adipose tissue-derived stem cells osteogenic differentiation. Bone 2014; 58: 59–66.
Vimalraj S, Partridge NC, Selvamurugan N . A positive role of microRNA-15b on regulation of osteoblast differentiation. J Cell Physiol 2014; 229: 1236–1244.
Li J, Chen Y, Guo X, Zhou L, Jia Z, Tang Y et al. Inhibition of miR-15b decreases cell migration and metastasis in colorectal cancer. Tumour Biol 2016; 37: 8765–8773.
Zheng X, Chopp M, Lu Y, Buller B, Jiang F . MiR-15b and miR-152 reduce glioma cell invasion and angiogenesis via NRP-2 and MMP-3. Cancer Lett 2013; 329: 146–154.
Lovat F, Fassan M, Gasparini P, Rizzotto L, Cascione L, Pizzi M et al. miR-15b/16-2 deletion promotes B-cell malignancies. Proc Natl Acad Sci USA 2015; 112: 11636–11641.
Neilsen PM, Noll JE, Mattiske S, Bracken CP, Gregory PA, Schulz RB et al. Mutant p53 drives invasion in breast tumors through up-regulation of miR-155. Oncogene 2013; 32: 2992–3000.
Bouamar H, Jiang D, Wang L, Lin AP, Ortega M, Aguiar RC . MicroRNA 155 control of p53 activity is context dependent and mediated by Aicda and Socs1. Mol Cell Biol 2015; 35: 1329–1340.
Maes OC, An J, Sarojini H, Wang E . Murine microRNAs implicated in liver functions and aging process. Mech Ageing Dev 2008; 129: 534–541.
Jin HL, Kim JS, Kim YJ, Kim SJ, Broxmeyer HE, Kim KS . Dynamic expression of specific miRNAs during erythroid differentiation of human embryonic stem cells. Mol Cells 2012; 34: 177–183.
Nimmo R, Ciau-Uitz A, Ruiz-Herguido C, Soneji S, Bigas A, Patient R et al. MiR-142-3p controls the specification of definitive hemangioblasts during ontogeny. Dev Cell 2013; 26: 237–249.
Cheng CY, Hwang CI, Corney DC, Flesken-Nikitin A, Jiang L, Oner GM et al. miR-34 cooperates with p53 in suppression of prostate cancer by joint regulation of stem cell compartment. Cell Rep 2014; 6: 1000–1007.
Choi YJ, Lin CP, Ho JJ, He X, Okada N, Bu P et al. miR-34 miRNAs provide a barrier for somatic cell reprogramming. Nat Cell Biol 2011; 13: 1353–1360.
Menges CW, Kadariya Y, Altomare D, Talarchek J, Neumann-Domer E, Wu Y et al. Tumor suppressor alterations cooperate to drive aggressive mesotheliomas with enriched cancer stem cells via a p53-miR-34a-c-Met axis. Cancer Res 2014; 74: 1261–1271.
Okita K, Ichisaka T, Yamanaka S . Generation of germline-competent induced pluripotent stem cells. Nature 2007; 448: 313–317.
Acknowledgements
We thank Dr Ivano Amelio for constructive criticism and suggestions. This work has been supported by the Medical Research Council, UK; grants from 'Fondazione Roma' (2015), AIRC (2011-IG11955), AIRC 5xmille (MCO #9979) to GM. Prof. Varda Rotter is the incumbent of the Norman and Helen Asher Professorial Chair for Cancer Research at the Weizmann Institute of Science.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Edited by G Raschella
Supplementary Information accompanies this paper on Cell Death and Disease website
Supplementary information
Rights and permissions
Cell Death and Disease is an open-access journal published by Nature Publishing Group. This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Grespi, F., Landré, V., Molchadsky, A. et al. Differential regulated microRNA by wild type and mutant p53 in induced pluripotent stem cells. Cell Death Dis 7, e2567 (2016). https://doi.org/10.1038/cddis.2016.419
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/cddis.2016.419
This article is cited by
-
MALAT-1/p53/miR-155/miR-146a ceRNA circuit tuned by methoxylated quercitin glycoside alters immunogenic and oncogenic profiles of breast cancer
Molecular and Cellular Biochemistry (2022)
-
Targeting mutant p53 for cancer therapy: direct and indirect strategies
Journal of Hematology & Oncology (2021)
-
Interplay between p53 and non-coding RNAs in the regulation of EMT in breast cancer
Cell Death & Disease (2021)
-
p73 Regulates Primary Cortical Neuron Metabolism: a Global Metabolic Profile
Molecular Neurobiology (2018)