Abstract
Heart failure is a leading cause of death in aging population. Cardiac hypertrophy is an adaptive reaction of the heart against cardiac overloading, but continuous cardiac hypertrophy is able to induce heart failure. We found that the level of miR-541 was decreased in angiotensin II (Ang-II) treated cardiomyocytes. Enforced expression of miR-541 resulted in a reduced hypertrophic phenotype upon Ang-II treatment in cellular models. In addition, we generated miR-541 transgenic mice that exhibited a reduced hypertrophic response upon Ang-II treatment. Furthermore, we found miR-541 is the target of microphthalmia-associated transcription factor (MITF) in the hypertrophic pathway and MITF can negatively regulate the expression of miR-541 at the transcriptional levels. MITFce/ce mice exhibited a reduced hypertrophic phenotype upon Ang-II treatment. Knockdown of MITF also results in a reduction of hypertrophic responses after Ang-II treatment. Knockdown of miR-541 can block the antihypertrophic effect of MITF knockdown in cardiomyocytes upon Ang-II treatment. This indicates that the effect of MITF on cardiac hypertrophy relies on the regulation of miR-541. Our present study reveals a novel cardiac hypertrophy regulating pathway that was composed of miR-541 and MITF. Modulation of their levels may provide a new approach for tackling cardiac hypertrophy.
Similar content being viewed by others
Main
Heart failure is a leading cause of death in aging population. Cardiac hypertrophy is an adaptive reaction of the heart against cardiac overloading, but continuous cardiac hypertrophy can increase cardiac remodeling and heart failure in the future. Nevertheless, the underlying molecular mechanisms of cardiac hypertrophy are still poorly understood. Cardiac hypertrophy is considered to be a therapeutic target for controlling heart failure. There is still a lack of novel approaches to treat cardiac remodeling and heart failure. In order to prevent heart failure, it is necessary to identify and characterize molecules that may regulate cardiac hypertrophy.
MicroRNAs (miRNAs) are small noncoding RNA molecules (∼22 nucleotides) and act as negative regulators of gene expression by inhibiting mRNA translation or promoting mRNA degradation.1 Growing evidences have shown that miRNAs play an important role in the regulation of development, differentiation, proliferation and apoptosis.2, 3, 4 The miRNAs emerged as powerful and dynamic modifiers of cardiovascular diseases. They were reported to participate in the regulation of cardiac hypertrophy.5, 6 In another study of our laboratory, we found miR-541 was significantly downregulated by angiotensin II (Ang-II) treatment in cardiomyocytes that was detected by a microarray assay. So far, miR-541 has never been reported in cardiovascular disease.
Microphthalmia-associated transcription factor (MITF) belongs to a family of transcription factors that contain a basic helix-loop-helix and leucine-zipper (bHLH-LZ) structure.7, 8, 9 MITF is a product of the microphthalmia locus in mice, and it plays important roles in the differentiation of melanocytes, hearing and eye developmental pathways.10, 11 Mutations in this locus cause the Waardenburg syndrome type II in humans that is characterized by sensorineural hearing loss and abnormal pigmentation of the hair and skin.12, 13 MITF regulates gene transcription by binding to E-box motif (CANNTG) in the upstream promoter regions of MITF-responsive genes.14 Beyond the already known genes activated by MITF, other potential targets were identified by using the microarrays in MITF-overexpressing human melanoma cells.15 Recently, a report showed that the mouse strains with MITF mutations were able to decrease the cardiac hypertrophy in response to β-adrenergic stimulation.16 MITF was also reported to be decreased in heart failure sample.17
The aim of our present work is to investigate the role of miR-541 (our present work only focus on the miR-541-5p) in the cardiac hypertrophy pathway. The miR-541 was able to decrease the cardiomyocyte hypertrophy in response to Ang-II stimulation. We generate miR-541 transgenic mouse that was able to antagonize cardiac hypertrophic response of Ang-II. Furthermore, we found that MITF can negatively regulate the expression of miR-541. The ce/ce mutant MITF (MITFce/ce) mice were able to decrease the cardiac hypertrophy in response to Ang-II stimulation. This will provide new insights into understanding the pathogenesis of cardiac hypertrophy.
Results
The miR-541 can inhibit cardiac hypertrophy induced by Ang-II treatment
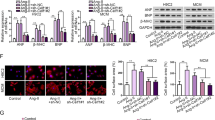
In another study of our laboratory, microarray assays were conducted from neonatal mouse cardiomyocytes that were treated with Ang II for 24 h or untreated (control). The result shows the levels of some miRNAs were significantly altered upon Ang-II treatment (Supplementary Figure 1). The miR-541 was one of the significantly downregulated miRNAs, and it has never been reported in cardiovascular disease. Hence, we wanted to know whether miR-541 can regulate cardiac hypertrophy in response to Ang-II treatment. The level of miR-541 was reduced upon Ang-II treatment in cardiomyocytes, and it occurred in a time-dependent manner (Figure 1a). Enforced expression of miR-541 resulted in a decrease in cell size after Ang-II treatment in cardiomyocytes (Figure 1b). Other cardiac hypertrophic responses including cell surface area (Figure 1c), protein/DNA ratio (Figure 1d) and cardiac hypertrophic markers atrial natriuretic factor (ANF; Figure 1e) and β-myosin heavy chain (β-MHC; Figure 1f) were also attenuated after Ang-II treatment. This indicates that miR-541 can inhibit the cardiomyocyte hypertrophy induced by Ang-II treatment in vitro.
The miR-541 can inhibit the cardiomyocyte hypertrophy induced by Ang-II. (a) Cardiomyocytes were treated with Ang-II at the indicated time. The levels of miR-541 were detected by qRT-PCR and the results were normalized to that of U6. *P<0.05 versus control. (b–f) Cardiomyocytes were infected with adenoviral miR-541 or β-gal at the MOI of 80. At 24 h after infection, cells were treated with Ang-II. *P<0.05 versus β-gal+Ang-II. (b) Representative photos of cardiomyocytes revealed by phalloidin-TRITC staining. (c) Analysis of cell surface area. (d) Protein/DNA ratio. (e) The expression levels of ANP were detected by immunoblot and qRT-PCR. (f) The expression levels of β-MHC were detected by immunoblot and qRT-PCR
The miR-541 antagonizes cardiac hypertrophy in the animal model
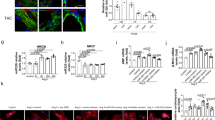
To better understand the function of miR-541 in the heart, we generated transgenic mice with cardiac-specific overexpression of miR-541 (Supplementary Figure 2). Five lines of miR-541 transgenic mice demonstrated a high-level expression of miR-541 in the heart (Figure 2a). Line 2 of miR-541 transgenic mice was used for the follow-up studies. These mice were born normally and developed to adulthood. The miR-541 transgenic mice had no significant alterations in cardiac hypertrophy (Figure 2b), and the cardiac function was also not different as compared with wild-type (WT) mice (Figure 2c). Subsequently, we detected cardiac hypertrophic responses in miR-541 transgenic mice in response to Ang-II treatment. The miR-541 transgenic mice exhibited a reduced hypertrophic phenotype upon Ang-II treatment. The cardiac hypertrophic responses including heart/body weight ratio (Figure 2d), cross-sectional area (Figure 2e) and cardiac hypertrophy markers ANF and β-MHC (Figure 2f) were attenuated compared with WT mice. The level of myocardial fibrosis was significantly reduced in miR-541 transgenic mice upon Ang-II treatment as compared with WT mice (Figure 2g). Concomitantly, the cardiac function was also significantly ameliorated in miR-541 transgenic mice (Figure 2h). Thus, it appears that miR-541 exerts antihypertrophic function in animal model.
The miR-541 transgenic mice can antagonize cardiac hypertrophy induced by Ang-II. (a) Detection of miR-541 levels in miR-541 transgenic (Tg) mice. The expression level of miR-541 in heart was detected by qRT-PCR from wild-type and different lines of miR-541 Tg mice, and the results were normalized to that of U6. (b) The miR-541 Tg mice developed normally without obvious phenotype alterations under basal conditions (low panel, bar=20 μm). The ratios of heart/body weight (n=16). Histological sections were stained with wheat germ agglutinin-FITC conjugate to determine cell size. Cross-sectional areas were analyzed. (c) Cardiac function of wild-type and miR-541 Tg mice were analyzed by echocardiography. LVPWd, left ventricular posterior wall thickness at end-diastole; FS, fractional shortening (n=10 per group). (d–h) Reduced hypertrophic response of miR-541 Tg mice in response to Ang-II infusion. Wild-type (WT) and miR-541 Tg mice were infused with Ang-II. *P<0.05 versus WT+Ang-II. (d) The ratios of heart/body weight. (e) Cross-sectional areas were analyzed by TRITC-conjugated wheat germ agglutinin staining. (f) The expression levels of ANP and β-MHC were detected by qRT-PCR. (g) The miR-541 Tg mice exhibited reduced myocardial fibrosis in response to Ang-II infusion. WT and miR-541 Tg mice were infused with Ang-II. The myocardial fibrosis was determined by Masson trichrome staining. Bar=20 μm. (h) Analysis of cardiac function by echocardiography. IVSd, end-diastolic interventricular septum thickness; LVIDs, left ventricular internal diameters at end-systole (n=8 per group)
The miR-541 is negatively regulated by MITF at the transcriptional level
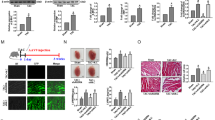
MITF is a bHLH-LZ transcription factor. MITF was reported to be decreased in heart failure sample,17 and the mouse strains with MITF mutations were also able to decrease cardiac hypertrophy in response to β-adrenergic stimulation.16 The simultaneous alterations of miR-541 and MITF made us to consider whether they are related in the cardiac hypertrophic pathway. Hence, we analyzed the promoter region of mouse miR-541 and observed that there was a potential binding site of MITF (Figure 3a). The level of miR-541 was increased in MITFce/ce mice as compared with WT mice (Figure 3b). Next, we tested whether MITF can bind to the promoter region of miR-541. ChIP assay revealed the association of MITF with the binding site in response to Ang-II treatment (Figure 3c). Furthermore, we tested whether MITF can influence the promoter activity of miR-541. The WT miR-541 promoter demonstrated a decreased activity after overexpression of MITF. However, mutations in the MITF consensus binding site could abolish the inhibitory effect of MITF on the promoter activity of miR-541 (Figure 3d). These data suggest that MITF can negatively regulate the expression of miR-541. Ang-II treatment could also decrease the promoter activity of miR-541 (Figure 3e). Concomitantly, knockdown of MITF could attenuate the decrease of miR-541 promoter activity induced by Ang-II in cardiomyocytes (Figure 3f). These data indicate that miR-541 can be transcriptionally repressed by MITF.
The miR-541 is negatively regulated by MITF at the transcriptional level. (a) Mouse miR-541 promoter region contains the potential MITF binding site and the mutant. (b) The levels of miR-541 were detected by qRT-PCR in the heart of MITFce/ce mice and WT mice. The results were normalized to that of U6. *P<0.05. (c) ChIP analysis of MITF binding site in the promoter of miR-541. (d) MITF suppressed miR-541 promoter activity. Cardiomyocytes were infected with adenoviral MITF or β-gal at the MOI of 80. At 24 h after infection, cells were transfected with the constructs of the empty vector (pGL-4.17), the wild-type promoter (wt) or the promoter with mutations in the binding site (mutant), respectively. The luciferase activities were detected (upper panel). MITF levels were detected by immunoblot (lower panel). *P<0.05. (e) Cardiomyocytes were transfected with the constructs of the empty vector (pGL-4.17) or the wt promoter, and then were treated with Ang-II at the indicated time. The luciferase activities were detected. *P<0.05 versus wt alone. (f) Cardiomyocytes were infected with adenoviral MITF-siRNA or MITF-sc at the MOI of 100. At 24 h after infection, cells were transfected with the constructs of the empty vector (pGL-4.17) or the wt promoter, and then were treated with Ang-II. The luciferase activities were detected. *P<0.05
MITF can regulate cardiac hypertrophy in vivo
To test whether MITF is involved in the Ang-II-induced cardiac hypertrophic pathway, we infused MITFce/ce mice with Ang-II, and detected the hypertrophic responses of MITFce/ce mice in response to hypertrophic stimulation. MITFce/ce mice exhibited a reduced hypertrophic phenotype upon Ang-II treatment (Figure 4a). The heart/body weight ratio in MITFce/ce mice group was decreased after Ang-II treatment as compared with the WT group (Figure 4b). We also observed an attenuation of other cardiac hypertrophic responses including cross-sectional area (Figures 4c and d) and cardiac hypertrophic markers ANF and β-MHC (Figure 4e). Concomitantly, the cardiac function in MITFce/ce mice group was significantly ameliorated after Ang-II treatment as compared with the WT group (Figure 4f). Thus, it appears that MITFce/ce mice resist the cardiac hypertrophic response of Ang-II.
MITF can regulate cardiac hypertrophy in vivo. Wild-type (WT) and MITFce/ce mice were infused with Ang-II or saline for 14 days. *P<0.05 versus WT+Ang-II. (a) Gross hearts (upper panel, bar=2 mm) and heart sections stained with hematoxylin and eosin in lower panel (bar=20 μm). (b) The ratios of heart/body weight (n=16). (c) Typical picture of histological sections stained with TRITC-conjugated wheat germ agglutinin. (d) Cross-sectional areas were analyzed by TRITC-conjugated wheat germ agglutinin staining. (e) The expression levels of ANP and β-MHC were detected by qRT-PCR. (f) Analysis of cardiac function by echocardiography. LVPWd, left ventricular posterior wall thickness at end-diastole; IVSd, end-diastolic interventricular septum thickness; LVIDs, left ventricular internal diameters at end-systole; FS, fractional shortening (n=8 per group)
MITF can regulate cardiac hypertrophy in vitro
Next, we attempted to understand the function of MITF in hypertrophic machinery in cardiomyocytes. We constructed the adenovirus harboring MITF siRNA that could significantly reduce the level of MITF in cardiomyocytes (Figure 5a). Knockdown of MITF resulted in a reduction of hypertrophic responses revealed by the cell surface area measurement (Figure 5b) and the analysis of protein/DNA ratio (Figure 5c) after Ang-II treatment. The cardiomyocyte size (Figure 5d) and cardiac hypertrophic markers ANF (Figure 5e) and β-MHC (Figure 5f) were also reduced after knockdown of MITF by siRNA as compared with that in scramble control. These data suggest that MITF participates in antagonizing hypertrophy in cardiomyocytes.
MITF can regulate cardiac hypertrophy in vitro. (a) Cardiomyocytes were infected with adenoviral MITF-siRNA or its scramble form (MITF-sc) at the indicated MOI. The levels of MITF were detected by immunoblot. (b–f) Knockdown of MITF results in reduced hypertrophic responses induced by Ang-II. Cardiomyocytes were infected with adenoviral MITF-siRNA or MITF-sc at the MOI of 100. At 24 h after infection, cells were treated with Ang-II. *P<0.05 versus MITF-sc+Ang-II. (b) Analysis of MITF levels by immunoblot (upper panel); cell surface area measurement (lower panel). (c) Protein/DNA ratio. (d) Representative photos of cardiomyocytes revealed by phalloidin-TRITC staining. (e) The expression levels of ANP were detected by immunoblot and qRT-PCR. (f) The expression levels of β-MHC were detected by immunoblot and qRT-PCR
MITF regulates cardiac hypertrophy through targeting miR-541
Next, we wanted to further verify whether MITF exerts its antihypertrophic function through targeting miR-541. We used chemically synthesized antagomir18 (anta-541) to knock down endogenous miR-541. The antagomir-NC (anta-NC) was used as a negative control. The anta-541 could efficiently counteract the increase of miR-541 after knockdown of MITF in cardiomyocytes in response to Ang-II (Figure 6a). The anta-541 can also compensate the decrease of cardiac hypertrophic responses including cell surface area (Figure 6b), protein/DNA ratio (Figure 6c) and cardiac hypertrophic marker β-MHC (Figure 6d) after knockdown of MITF in cardiomyocytes in response to Ang-II. These data indicate that knockdown of miR-541 can block the antihypertrophic effect of knockdown of MITF in cardiomyocytes in response to Ang-II. The effect of MITF on cardiac hypertrophy may rely on the regulation of miR-541. Ang-II treatments have been reported to influence the phosphorylation of Akt and Erk1/2 in rat cardiac fibroblasts and endothelial progenitor cells.19, 20 A recent report has shown that miR-541 can negatively regulate the expression of Her2 and Erk1/2 in breast cancer cells.21 Hence, we also detected Akt, Erk1/2 and Her2 in response to Ang-II with or without anti-miR-541. The results showed that Ang-II treatment increased the phosphorylation of Akt and Erk1/2; meanwhile knockdown of miR-541 by antagomir could enhance these effects (Figure 6e). Both Ang-II treatment and knockdown of miR-541 did not influence the expression of Her2 (Figure 6e).
MITF regulates cardiac hypertrophy through targeting miR-541. (a–d) Cardiomyocytes were infected with adenoviral MITF-siRNA or MITF-sc at the MOI of 100. At 24 h after infection, cells were transfected with miR-541 antagomir (anta-541) or the antagomir control (anta-NC), and then were treated with Ang-II. *P<0.05. (a) The levels of miR-541 were detected by qRT-PCR and the results were normalized to that of U6. (b) Analysis of cell surface area. (c) Protein/DNA ratio. (d) The expression levels of β-MHC were detected by qRT-PCR. (e) Cardiomyocytes were transfected with miR-541 antagomir (anta-541) or the antagomir negative control (anta-NC), and then were treated with Ang-II. The levels of phospho-Akt, Akt, phospho- Erk1/2, Erk1/2 and Her2 were detected by immunoblot
Discussion
Cardiac hypertrophy is an adaptive reaction of the heart in response to a variety of physiological as well as pathophysiological stimuli, and the continuous cardiac hypertrophy can eventually lead to heart failure. Heart failure is one of the leading causes of hospitalization and death worldwide. Antagonizing the maladaptive cardiac hypertrophy is considered to be a therapeutic target for treating heart failure. Hence, it is necessary to elucidate the molecular mechanisms of cardiac hypertrophy, and discover effective therapeutic targets for suppressing maladaptive hypertrophy and the consequent heart failure. In another study of our laboratory regarding how Ang-II altered the expression of miRNAs, a miRNA microarray analysis from cardiomyocytes identified the significant downregulation of nine miRNAs in response to Ang-II treatment. The miR-541 is one of them, and has never been reported in cardiovascular disease. Our present work identified that MITF is a pro-hypertrophic transcriptional factor. It can negatively regulate the expression of miR-541 at the transcriptional levels. We also produced miR-541 transgenic mice and these mice are resistant to Ang-II-induced cardiac hypertrophy. Our data further show that MITF exerts its pro-hypertrophic function through targeting miR-541. Our results will provide new insights into understanding the pathogenesis of cardiac hypertrophy.
Recent works about miRNAs demonstrate that a number of miRNAs are able to influence cardiac hypertrophy pathway. For example, overexpression of miR-1 can reduce the cell size and attenuates the expression of hypertrophic markers, whereas silencing of miR-1 leads to the hypertrophic phenotype.22 The in vivo inhibition of miR-199b by a specific antagomir reduces nuclear NFAT activity and causes marked inhibition and even reversal of hypertrophy and fibrosis in mouse models of heart failure.23 The miRNAs usually have multiple functions. Some miRNAs exert pro-hyertrophic function, whereas others are able to inhibit hypertrophy. Our present work found that miR-541 is a target of MITF, and the level of miR-541 was also decreased in response to Ang-II treatment. The miR-541 may be the downstream factor of MITF in cardiac hypertrophy pathway. Hence, we constructed the transgenic mice with cardiac-specific overexpression of miR-541.
It is noteworthy that miR-541 transgenic mice exhibit no obvious alterations in phenotype under physiological condition. However, these mice are resistant to cardiac hypertrophy under pathological stimulation with Ang-II. This is similar to a previous report that alteration of a miRNA may not influence the phenotype under physiological condition, and may function only under pathological condition.24, 25 Other reports also show that basal cell size can be reduced by overexpression of a miRNA. For instance, overexpression of miR-98 can reduce cell size in the absence of pathological insult.26 Cardiac-specific overexpression of miR-208a can induce cardiac hypertrophic growth in mice, resulting in a pronounced repression of target genes including thyroid hormone-associated protein 1 and myostatin (two negative regulators of muscle growth and hypertrophy).27 Those prohypertrophic miRNAs may induce cardiac hypertrophic responses in the absence of hypertrophic stimulation.27 Because physiological cell growth and the pathological hypertrophy can be regulated by distinct cellular machinery, the underlying mechanism by which a miRNA regulates cell size under physiological condition is an interesting topic for future studies. Our present work also shows that the miR-541 transgenic mice exhibited less myocardial fibrosis in miR-541 transgenic mice upon Ang-II treatment as compared with WT mice. The above data indicate that miR-541 transgenic mice tolerate Ang-II-stimulated cardiac hypertrophy.
Recently, a report has shown that Her2 and Erk1/2 may be the targets of miR-541 in breast cancer cells.21 Our present results showed that knockdown of miR-541 by antagomir enhanced the phosphorylation of Akt and Erk1/2 after Ang-II treatment in cardiomyocyte. This indicates that the antihypertrophic function of miR-541 may rely on Akt and Erk1/2 signaling pathways. Ang-II treatment and knockdown of miR-541 did not alter the total levels of Akt and Erk1/2. This suggests that alteration of Akt and Erk1/2 are not the direct effects of miR-541. Our results also showed that Ang-II treatment and knockdown of miR-541 did not influence the expression level of Her2 in mouse cardiomyocytes. This is inconsistent with the previous report in human breast cancer cells,21 and this may be because of species differences.
MITF resides at the microphthalmia locus in mice, and belongs to a family of transcription factors that contain the bHLH-LZ structure.7, 8, 9 Mutations of this gene result in deafness, bone loss, small eyes and poorly pigmented eyes and skin.28 In humans, mutation in this gene causes Waardenburg syndrome type, II,12, 13 and it plays an important role in the development and the function of several cell types, including melanocytes and mast cells.11, 29, 30 MITF is transcribed from several alternative promoters, and has generated several splice transcripts at transcriptional levels.11 This may exert potentially distinct biological functions. MITF is also abundantly expressed in cardiomyocytes, and only the MITF-H isoform is expressed in cardiomyocytes.31 Studies show that the expression of myosin light-chain 1a (MLC-1a) is regulated by MITF-H.31 Recently reports also show that the mouse strains with MITF mutations were able to decrease cardiac hypertrophy in response to β-adrenergic stimulation,16 and MITF was also decreased in heart failure sample.17 This indicates that MITF may play an important role in the regulation of cardiac hypertrophy. In the present study, we used MITFce/ce mice and found that MITFce/ce mice can resist Ang-II-induced cardiac hypertrophic response. In addition, we also knocked down the MITF in cardiomyocytes in vitro, and these cells can also antagonize hypertrophy in response to Ang-II treatment. The above data demonstrate that MITF is an important regulator of cardiac hypertrophy.
The present work shows that MITF can negatively regulate the expression of miR-541 at the transcriptional levels. This is consistent with our previous report that Foxo3a can negatively regulate the expression of miR-874.32 In addition, we found that knockdown of miR-541 can block the antihypertrophic effect of knockdown of MITF in cardiomyocytes in response to Ang-II. This indicates that the effect of MITF on cardiac hypertrophy relies on the regulation of miR-541. The miRNAs usually regulate gene expression at the post-transcriptional level through both translational inhibition and mRNA degradation. Which gene is the target of miR-541 in cardiac hypertrophy pathway? This requires further study.
In summary, our present work identified that miR-541 exerts antihypertrophic function in both cellular and animal models. Furthermore, we reveal that miR-541 is the target of MITF in the hypertrophic pathway, and MITF can negatively regulate the expression of miR-541 at the transcriptional levels. MITF is also able to influence cardiac hypertrophy in cellular and animal models. MITF regulates cardiac hypertrophy through targeting miR-541. Our results provide novel molecular pathway regulating hypertrophy in the heart. Modulation of miR-541 or MITF levels may provide an intriguing approach for tackling cardiac hypertrophy.
Materials and Methods
Animal
All mice lines were held and propagated in a specific pathogen-free environment. MITFsp/ce mice were purchased from the Jackson Laboratory (Bar Harbor, ME, USA), and bred to produce MITFce/ce mice for experiments. The MITF encoded by the mutated mouse allele (ce/ce) lack the LZ domain of MITF because of a stop codon between the HLH and LZ domains.10 Littermates of MITFce/ce mice were distinguished by coat color.
For generation of cardiac-specific overexpression of miR-541 transgenic mice, a 414-bp DNA fragment containing murine miR-541 was cloned to the pαMHC-clone26 vector (kindly provided by Dr. Zhongzhou Yang, Model Animal Research Center of Nanjing University, Nanjing, China), under the control of the α-MHC promoter. The primers used to generate miR-541 transgenic mice include: forward primer 5′- CACAGGTCAGTTTCCAGAACA-3′; reverse primer: 5′-CGGTGATGTCATAGCAAGAA-3′. Microinjection was performed following standard protocols. The primers for genotyping miR-541 transgenic mice include: forward primer in the α-MHC promoter, 5′-CAGAAATGACAGACAGATCCCTCC-3′; the reverse primer in the miR-541 DNA, 5′- CCCTTGTGCCAGTGTGTAT-3′.
Animal experiments
C57BL/6 mice (WT mice) were purchased from Institute of Laboratory Animal Science of Chinese Academy of Medical Sciences (Beijing, China). The adult male mice (8 weeks old) were used throughout the study. WT mice, MITFce/ce mice and miR-541 transgenic mice were infused with Ang-II (0.6 mg/kg/day dissolved in 0.9% NaCl, Sigma, St. Louis, MO, USA); saline-infused mice served as controls. All mice were infused with Ang-II for 2 weeks. The infusions were executed with implanted osmotic minipumps (Alzet model 1002, Alza Corp, Mountain View, CA, USA) as we have described previously.33 All animal experimental procedures were approved by the Animal Ethics Committee of Institute of Zoology, Chinese Academy of Sciences.
Cardiomyocyte culture and treatment
Cardiomyocytes were isolated from 1- to 2-day-old mice as we have described previously.34 Briefly, after dissection, hearts were washed and minced in HEPES-buffered saline solution. Tissues were then dispersed in a series of incubations at 37°C in HEPES-buffered saline solution containing 1.2 mg/ml pancreatin and 0.14 mg/ml collagenase (Worthington, Lakewood, NJ, USA). After centrifugation, cells were resuspended in Dulbecco’s modified Eagle’s medium/F-12 (GIBCO, Invitrogen, San Diego, CA, USA) containing 5% heat-inactivated horse serum, 0.1 mM ascorbate, insulin-transferring-sodium selenite media supplement (Sigma), 100 U/ml penicillin, 100 μg/ml streptomycin and 0.1 mM bromodeoxyuridine. The dissociated cells were preplated at 37°C for 1 h. The cells were then diluted to 1 × 106 cells/ml and plated in 10 μg/ml laminin-coated different culture dishes according to the specific experimental requirements. Cells were treated with Ang-II at 150 nM for 24 h, except as otherwise indicated elsewhere.
Echocardiographic assessment of cardiac dimensions and function
Transthoracic echocardiography was performed on lightly anesthetized mice by using a Vevo 770 high-resolution system (Visualsonics, Toronto, ON, Canada) equipped with a 40-MHz RMV 704 scanhead. Two-dimensional guided M-mode tracings were recorded in both parasternal long and short axis views at the level of papillary muscles. Left ventricular posterior wall thickness at end-diastole (LVPWd), end-diastolic interventricular septum thickness (IVSd), left ventricular internal diameters at end-systole (LVIDs) and fractional shortening (FS) were calculated with the established standard equation. All the measurements were made from more than three beats and averaged.
Immunoblot
Immunoblot was performed as we described previously.35 In brief, cells were lysed for 1 h at 4°C in a lysis buffer (20 mM Tris (pH 7.5), 2 mM EDTA, 3 mM EGTA, 2 mM DTT, 250 mM sucrose, 0.1 mM PMSF, 1% Triton X-100 and a protease inhibitor cocktail). Samples were subjected to 12% SDS-PAGE and transferred to nitrocellulose membranes. Blots were probed using antibodies. The antibodies against MITF, Akt and phospho-Akt were from Abcam (Cambridge, UK), antibodies against ERK1/2, phospho-ERK1/2 and Her2 were from Cell Signaling Technology (Beverly, MA, USA) and horseradish peroxidase-conjugated goat anti-mouse IgG was purchased from Santa Cruz Biotechnology (Santa Cruz, CA, USA). Probe using β-actin antibody (Santa Cruz Biotechnology) was used as loading control. Proteins were visualized by enhanced chemiluminescence.
Quantitative reverse transcription-PCR (qRT-PCR)
Stem-loop qRT-PCR for mature miR-541-5p was performed as described previously.36 Total RNA was extracted using Trizol reagent. After DNAse I (Takara, Otsu, Japan) treatment, RNA was reverse transcribed with reverse transcriptase (ReverTra Ace, Toyobo, Osaka, Japan). The SYBR Premix Ex Taq II Kit (Takara) was used for amplification. Quantitative PCR was performed on a CFX96 Real-Time PCR Detection System (Bio-Rad, Richmond, CA, USA). The results of qRT-PCR were normalized to that of U6. The sequences of U6 primers were: forward: 5′-GCTTCGGCAGCACATATACTAA-3′; and reverse: 5′-AACGCTTCACGAATTTGCGT-3′. The quantitative detection of ANP and β-MHC by qRT-PCR was performed using the SYBR Premix Ex Taq II Kit (Takara) on a CFX96 Real-Time PCR Detection System (Bio-Rad). The qRT-PCR primers for ANP and β-MHC, the sequences of ANP primers were forward: 5′-CTCCGATAGATCTGCCCTCTTGAA-3′ and reverse: 5′- GGTACCGGAAGCTGTTGCAGCCTA-3′. β-MHC forward primer was 5′-CAGACATAGAGACCTACCTTC-3′ and reverse was 5′-CAGCATGTCTAGAAGCTCAGG-3′. The results were standardized to control values of glyceraldehyde-3-phosphate dehydrogenase (GAPDH). GAPDH forward primer was 5′-TGTGTCCGTCGTGGATCTGA-3′ and reverse was 5′-CCTGCTTCACCACCTTCTTGA-3′. The specificity of the PCR amplification was confirmed by melting curve analysis.
Determinations of cell surface areas and protein/DNA ratio
Cell surface area of F-actin-stained cells or unstained cells was measured as we have described previously.35 In each group, 100–200 cardiomyocytes in 30–50 fields were examined. For staining of filamentous actin, the cardiomyocytes were fixed in 3.7% formaldehyde in PBS. Cells were dehydrated with acetone for 3 min and treated with 0.1% Triton X-100 for 20 min. They were then stained with a 50 μg/ml fluorescent Phalloidin-TRITC (Sigma) for 45 min at room temperature, and visualized by a laser confocal microscopy (Zeiss LSM 510 META, Oberkochen, Germany). To measure protein/DNA ratio, total protein and DNA contents were analyzed as we have described previously.34
Histological analysis
Histological analysis of the hearts was carried out as we have described previously.33 Briefly, hearts were excised, fixed in 10% formalin, embedded in paraffin and sectioned into 7 μm slices, and stained with hematoxylin–eosin (HE). To measure the cross-sectional area of the cardiomyocytes, the sections were stained with FITC-conjugated wheat germ agglutinin (Sigma) according to the method previously described.37 The myocardial fibrosis was determined by standard Masson trichrome staining in the heart sections according to the manufacturer’s instructions (Sigma).
Chromatin immunoprecipitation (ChIP) assay
ChIP assay was performed as we have described previously.33 In brief, cells were washed with PBS and incubated for 10 min with 1% formaldehyde at room temperature. The crosslinking was quenched with 0.1 M glycine for 5 min. Cells were washed twice with PBS and lysed for 1 h at 4°C in a lysis buffer. The cell lysates were sonicated into chromatin fragments with an average length of 500 to 800 bp as assessed by agarose gel electrophoresis. The samples were precleared with Protein-A agarose (Roche, Penzberg, Germany) for 1 h at 4°C on a rocking platform, and 5 μg MITF antibodies (Abcam, Cambridge, UK) were added and rocked overnight at 4°C. Immunoprecipitates were captured with 10% (vol/vol) Protein-A agarose for 4 h. Before use, Protein-A agarose was blocked twice at 4°C with salmon sperm DNA (2 μg/ml) overnight. DNA fragments were purified with a QIAquick Spin Kit (Qiagen, Hilden, Germany). The purified DNA was used as a template and amplified with the following primer sets: 5′-CCCTTCCTCTCCTCAACTCA-3′ and 5′-GTAAACGGGATGAAATGCTG-3′.
Constructions of mouse miR-541 promoter and its mutant
We downloaded 3-kb region upstream sequences of mouse miR-541, and we found potential MITF binding sites in −1441 to −1446. The miR-541 promoter fragment was amplified from mouse genome using PCR. The forward primer was 5′-AGAGGTTGCCCTTGGTGAA-3′ and the reverse primer was 5′-CAAGAGTTCACACAGGCTTCTA-3′. A 1895-bp fragment containing the binding sites was cloned into the pGL4.17 vector (Promega, Madison, WI, USA). The introduction of mutations in the putative MITF binding site was performed with the QuikChange II XL Site-Directed Mutagenesis Kit (Stratagene, La Jolla, CA, USA) using the WT vector as a template. The construct was sequenced to check that only the desired mutations had been introduced.
Luciferase activity assay was performed using the Dual-Luciferase Reporter Assay System (Promega) according to the manufacturer’s instructions. Cells were transfected with the constructs of the empty vector (pGL-4.17), the WT promoter (wt) or the promoter with mutations in the binding site (mutant), respectively. At 48 h after transfection, cells were lysed and luciferase activity was measured.
Adenoviral constructions and infection
The codine region of mouse MITF-H was amplified from mouse heart total cDNA using the following primer set: 5′-ATGGAGGCGCTTAGATTTGAG-3′ and 5′-CTAACACGCATGCTCCGTTTC-3′. The adenoviruses harboring MITF were constructed using the Adeno-X expression system (Clontech, Tokyo, Japan). The adenovirus containing β-galactosidase (β-gal) is as we have described elsewhere.38 To construct adenoviruses encoding miR-541, mouse genomic sequence harboring the pre-miR-541 was amplified using the following primer set: 5′-CACAGGTCAGTTTCCAGAACA-3′ and 5′-CGGTGATGTCATAGCAAGAA-3′, and then subcloned into the Adeno-X expression system (Clontech). The adenoviruses were produced according to the Kit’s instructions.
The mouse MITF-RNAi target sequence is 5′-CACGCACTCTCGAGCGTCG-3′. A nonrelated, scrambled RNAi without any other match in the mouse genomic sequence was used as a control (5′-AGAGCCCTATGCCCTGGCC-3′). The adenoviruses harboring these MITF-RNAi constructs were generated using the pSilencer adeno 1.0-CMV System (Ambion, Austin, TX, USA) according to the Kit’s instructions. Adenoviruses were amplified in HEK293 cells. Adenoviral infection of cardiomyocytes was performed as we have described previously.35
Transfection of antagomir
The miR-541-5p antagomir and the antagomir negative control (antagomir-NC) were purchased from GenePharma Co. Ltd (Shanghai, China). The antagomir sequence is 5′- AGUGUGACCAACAUCAGAAUCCCUU -3′. All the bases were 2′-OMe modified, and the 3′-end was conjugated to cholesterol. Chemically modified oligonucleotides 5′-CAGUACUUUUGUGUAGUACAA-3′ were used as a negative control (antagomir-NC). Cells were transfected with the antagomir or antagomir-NC at 50 nM. The transfection was performed using Lipofectamine 2000 (Invitrogen, San Diego, CA, USA) according to the manufacturer’s instructions.
Statistical analysis
All data were expressed as mean value±S.D. One-way analysis of variance (ANOVA) followed by the Tukey’s Honestly Significant Difference (HSD) multiple range test was used for comparison of group mean values. Data were analyzed by Statistical Package for Social Sciences (SPSS, Chicago, IL, USA) 13.0 software. P<0.05 was considered statistically significant.
Abbreviations
- miRNA:
-
microRNA
- ANF:
-
atrial natriuretic factor
- β-MHC:
-
β-myosin heavy chain
- Ang-II:
-
angiotensin II
- MITF:
-
microphthalmia-associated transcription factor
- qRT-PCR:
-
quantitative reverse transcription polymerase chain reaction
References
Lee Y, Ahn C, Han J, Choi H, Kim J, Yim J et al. The nuclear RNase III Drosha initiates microRNA processing. Nature 2003; 425: 415–419.
Qian L, Van Laake LW, Huang Y, Liu S, Wendland MF, Srivastava D . miR-24 inhibits apoptosis and represses Bim in mouse cardiomyocytes. J Exp Med 2011; 208: 549–560.
Sayed D, He M, Hong C, Gao S, Rane S, Yang Z et al. MicroRNA-21 is a downstream effector of AKT that mediates its antiapoptotic effects via suppression of Fas ligand. J Biol Chem 2010; 285: 20281–20290.
Wang X, Zhang X, Ren XP, Chen J, Liu H, Yang J et al. MicroRNA-494 targeting both proapoptotic and antiapoptotic proteins protects against ischemia/reperfusion-induced cardiac injury. Circulation 2010; 122: 1308–1318.
Care A, Catalucci D, Felicetti F, Bonci D, Addario A, Gallo P et al. MicroRNA-133 controls cardiac hypertrophy. Nat Med 2007; 13: 613–618.
Wang J, Song Y, Zhang Y, Xiao H, Sun Q, Hou N et al. Cardiomyocyte overexpression of miR-27b induces cardiac hypertrophy and dysfunction in mice. Cell Res 2012; 22: 516–527.
Hodgkinson CA, Moore KJ, Nakayama A, Steingrimsson E, Copeland NG, Jenkins NA et al. Mutations at the mouse microphthalmia locus are associated with defects in a gene encoding a novel basic-helix-loop-helix-zipper protein. Cell 1993; 74: 395–404.
Hughes MJ, Lingrel JB, Krakowsky JM, Anderson KP . A helix-loop-helix transcription factor-like gene is located at the mi locus. J Biol Chem 1993; 268: 20687–20690.
Tachibana M, Perez-Jurado LA, Nakayama A, Hodgkinson CA, Li X, Schneider M et al. Cloning of MITF, the human homolog of the mouse microphthalmia gene and assignment to chromosome 3p14.1-p12.3. Hum Mol Genet 1994; 3: 553–557.
Steingrimsson E, Moore KJ, Lamoreux ML, Ferre-D'Amare AR, Burley SK, Zimring DC et al. Molecular basis of mouse microphthalmia (mi) mutations helps explain their developmental and phenotypic consequences. Nat Genet 1994; 8: 256–263.
Steingrimsson E, Copeland NG, Jenkins NA . Melanocytes and the microphthalmia transcription factor network. Annu Rev Genet 2004; 38: 365–411.
Tassabehji M, Newton VE, Read AP . Waardenburg syndrome type 2 caused by mutations in the human microphthalmia (MITF) gene. Nat Genet 1994; 8: 251–255.
Nobukuni Y, Watanabe A, Takeda K, Skarka H, Tachibana M . Analyses of loss-of-function mutations of the MITF gene suggest that haploinsufficiency is a cause of Waardenburg syndrome type 2A. Am J Hum Genet 1996; 59: 76–83.
Morii E, Tsujimura T, Jippo T, Hashimoto K, Takebayashi K, Tsujino K et al. Regulation of mouse mast cell protease 6 gene expression by transcription factor encoded by the mi locus. Blood 1996; 88: 2488–2494.
Hoek KS, Schlegel NC, Eichhoff OM, Widmer DS, Praetorius C, Einarsson SO et al. Novel MITF targets identified using a two-step DNA microarray strategy. Pigment Cell Melanoma Res 2008; 21: 665–676.
Tshori S, Gilon D, Beeri R, Nechushtan H, Kaluzhny D, Pikarsky E et al. Transcription factor MITF regulates cardiac growth and hypertrophy. J Clin Invest 2006; 116: 2673–2681.
Gao Z, Xu H, DiSilvestre D, Halperin VL, Tunin R, Tian Y et al. Transcriptomic profiling of the canine tachycardia-induced heart failure model: global comparison to human and murine heart failure. J Mol Cell Cardiol 2006; 40: 76–86.
Krutzfeldt J, Rajewsky N, Braich R, Rajeev KG, Tuschl T, Manoharan M et al. Silencing of microRNAs in vivo with 'antagomirs'. Nature 2005; 438: 685–689.
Mai J, Qiu Q, Lin YQ, Luo NS, Zhang HF, Wen ZZ et al. Angiotensin II-derived reactive oxygen species promote angiogenesis in human late endothelial progenitor cells through heme oxygenase-1 via ERK1/2 and AKT/PI3K pathways. Inflammation 2014 e-pub ahead of print 18 January 2014 doi:10.1007/s10753-013-9806-9.
Wang LP, Wang Y, Zhao LM, Li GR, Deng XL . Angiotensin II upregulates K(Ca)3.1 channels and stimulates cell proliferation in rat cardiac fibroblasts. Biochem Pharmacol 2013; 85: 1486–1494.
Leivonen SK, Sahlberg KK, Makela R, Due EU, Kallioniemi O, Borresen-Dale AL et al. High-throughput screens identify microRNAs essential for HER2 positive breast cancer cell growth. Mol Oncol 2014; 8: 93–104.
Sayed D, Hong C, Chen IY, Lypowy J, Abdellatif M . MicroRNAs play an essential role in the development of cardiac hypertrophy. Circ Res 2007; 100: 416–424.
da Costa Martins PA, Salic K, Gladka MM, Armand AS, Leptidis S, el Azzouzi H et al. MicroRNA-199b targets the nuclear kinase Dyrk1a in an auto-amplification loop promoting calcineurin/NFAT signalling. Nat Cell Biol 2010; 12: 1220–1227.
van Rooij E, Sutherland LB, Qi X, Richardson JA, Hill J, Olson EN . Control of stress-dependent cardiac growth and gene expression by a microRNA. Science 2007; 316: 575–579.
Wang JX, Jiao JQ, Li Q, Long B, Wang K, Liu JP et al. miR-499 regulates mitochondrial dynamics by targeting calcineurin and dynamin-related protein-1. Nat Med 2011; 17: 71–78.
Yang Y, Ago T, Zhai P, Abdellatif M, Sadoshima J . Thioredoxin 1 negatively regulates angiotensin II-induced cardiac hypertrophy through upregulation of miR-98/let-7. Circ Res 2011; 108: 305–313.
Callis TE, Pandya K, Seok HY, Tang RH, Tatsuguchi M, Huang ZP et al. MicroRNA-208a is a regulator of cardiac hypertrophy and conduction in mice. J Clin Invest 2009; 119: 2772–2786.
Moore KJ . Insight into the microphthalmia gene. Trends Genet 1995; 11: 442–448.
Ito A, Morii E, Maeyama K, Jippo T, Kim DK, Lee YM et al. Systematic method to obtain novel genes that are regulated by mi transcription factor: impaired expression of granzyme B and tryptophan hydroxylase in mi/mi cultured mast cells. Blood 1998; 91: 3210–3221.
Jippo T, Morii E, Tsujino K, Tsujimura T, Lee YM, Kim DK et al. Involvement of transcription factor encoded by the mouse mi locus (MITF) in expression of p75 receptor of nerve growth factor in cultured mast cells of mice. Blood 1997; 90: 2601–2608.
Tshori S, Sonnenblick A, Yannay-Cohen N, Kay G, Nechushtan H, Razin E . Microphthalmia transcription factor isoforms in mast cells and the heart. Mol Cell Biol 2007; 27: 3911–3919.
Wang K, Liu F, Zhou LY, Ding SL, Long B, Liu CY et al. miR-874 regulates myocardial necrosis by targeting caspase-8. Cell Death Dis 2013; 4: e709.
Lin Z, Murtaza I, Wang K, Jiao J, Gao J, Li PF . miR-23a functions downstream of NFATc3 to regulate cardiac hypertrophy. Proc Natl Acad Sci USA 2009; 106: 12103–12108.
Tan WQ, Wang K, Lv DY, Li PF . Foxo3a inhibits cardiomyocyte hypertrophy through transactivating catalase. J Biol Chem 2008; 283: 29730–29739.
Murtaza I, Wang HX, Feng X, Alenina N, Bader M, Prabhakar BS et al. Down-regulation of catalase and oxidative modification of protein kinase CK2 lead to the failure of apoptosis repressor with caspase recruitment domain to inhibit cardiomyocyte hypertrophy. J Biol Chem 2008; 283: 5996–6004.
Chen C, Ridzon DA, Broomer AJ, Zhou Z, Lee DH, Nguyen JT et al. Real-time quantification of microRNAs by stem-loop RT-PCR. Nucleic Acids Res 2005; 33: e179.
Dolber PC, Bauman RP, Rembert JC, Greenfield JC Jr . Regional changes in myocyte structure in model of canine right atrial hypertrophy. Am J Physiol 1994; 267 (4 Pt 2): H1279–H1287.
Li PF, Dietz R, von Harsdorf R . p53 regulates mitochondrial membrane potential through reactive oxygen species and induces cytochrome c-independent apoptosis blocked by Bcl-2. EMBO J 1999; 18: 6027–6036.
Acknowledgements
This work was supported by National Natural Science Foundation of China (81270160, 81000034) and the National Basic Research Program of China (973 Program, 2011CB965300).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Edited by M Agostini
Supplementary Information accompanies this paper on Cell Death and Disease website
Supplementary information
Rights and permissions
Cell Death and Disease is an open-access journal published by Nature Publishing Group. This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivs 3.0 Unported License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/3.0/
About this article
Cite this article
Liu, F., Li, N., Long, B. et al. Cardiac hypertrophy is negatively regulated by miR-541. Cell Death Dis 5, e1171 (2014). https://doi.org/10.1038/cddis.2014.141
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/cddis.2014.141
Keywords
This article is cited by
-
Altered expression of miR-375 and miR-541 in type 2 diabetes patients with and without coronary artery disease (CAD): the potential of miR-375 as a CAD biomarker
Journal of Diabetes & Metabolic Disorders (2024)
-
Platelet membrane-camouflaged nanoparticles carry microRNA inhibitor against myocardial ischaemia‒reperfusion injury
Journal of Nanobiotechnology (2022)
-
The Tietz syndrome associated with cardiac malformation: a case report with literature review
The Egyptian Journal of Otolaryngology (2021)
-
NRF1-enhanced miR-4458 alleviates cardiac hypertrophy through releasing TTP-inhibited TFAM
In Vitro Cellular & Developmental Biology - Animal (2020)
-
Dgcr8 deletion in the primitive heart uncovered novel microRNA regulating the balance of cardiac-vascular gene program
Protein & Cell (2019)