Abstract
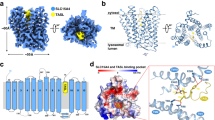
Nucleic acid–sensing Toll-like receptors (TLRs) play a pivotal role in innate immunity by recognizing foreign DNA and RNA. Compartmentalization of these TLRs in the endosome limits their activation by self-derived nucleic acids and reduces the possibility of autoimmune reactions. Although chaperone Unc-93 homolog B1, TLR signaling regulator (UNC93B1) is indispensable for the trafficking of TLRs from the endoplasmic reticulum to the endosome, mechanisms of UNC93B1-mediated TLR regulation remain largely unknown. Here, we report two cryo-EM structures of human and mouse TLR3–UNC93B1 complexes and a human TLR7–UNC93B1 complex. UNC93B1 exhibits structural similarity to the major facilitator superfamily transporters. Both TLRs interact with the UNC93B1 amino-terminal six-helix bundle through their transmembrane and luminal juxtamembrane regions, but the complexes of TLR3 and TLR7 with UNC93B1 differ in their oligomerization state. The structural information provided here should aid in designing compounds to combat autoimmune diseases.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
Cryo-EM maps are deposited in the Electron Microscopy Data Bank under accession codes EMD-30293 (human TLR3–UNC93B1), EMD-30294 (mouse TLR3–UNC93B1) and EMD-30501 (human TLR7–UNC93B1). Structure coordinates are deposited in the wwPDB under accession codes PDB 7C76 (human TLR3–UNC93B1), PDB 7C77 (mouse TLR3–UNC93B1) and PDB 7CYN (human TLR7–UNC93B1). Source data are provided with this paper.
References
Takeuchi, O. & Akira, S. Pattern recognition receptors and inflammation. Cell 140, 805–820 (2010).
Palm, N. W. & Medzhitov, R. Pattern recognition receptors and control of adaptive immunity. Immunological Rev. 227, 221–233 (2009).
Bell, J. K. et al. Leucine-rich repeats and pathogen recognition in Toll-like receptors. Trends Immunol. 24, 528–533 (2003).
Ohto, U. Conservation and divergence of ligand recognition and signal transduction mechanisms in Toll-like receptors. Chem. Pharm. Bull. (Tokyo) 65, 697–705 (2017).
Miyake, K. et al. Mechanisms controlling nucleic acid-sensing Toll-like receptors. Int. Immunol. 30, 43–51 (2018).
Majer, O., Liu, B. & Barton, G. M. Nucleic acid-sensing TLRs: trafficking and regulation. Curr. Opin. Immunol. 44, 26–33 (2017).
Roers, A., Hiller, B. & Hornung, V. Recognition of endogenous nucleic acids by the innate immune system. Immunity 44, 739–754 (2016).
Ewald, S. E. & Barton, G. M. Nucleic acid sensing Toll-like receptors in autoimmunity. Curr. Opin. Immunol. 23, 3–9 (2011).
Kim, Y. M., Brinkmann, M. M., Paquet, M. E. & Ploegh, H. L. UNC93B1 delivers nucleotide-sensing toll-like receptors to endolysosomes. Nature 452, 234–238 (2008).
Tabeta, K. et al. The Unc93b1 mutation 3d disrupts exogenous antigen presentation and signaling via Toll-like receptors 3, 7 and 9. Nat. Immunol. 7, 156–164 (2006).
Casrouge, A. et al. Herpes simplex virus encephalitis in human UNC-93B deficiency. Science 314, 308–312 (2006).
Fukui, R. et al. Unc93B1 restricts systemic lethal inflammation by orchestrating Toll-like receptor 7 and 9 trafficking. Immunity 35, 69–81 (2011).
Fukui, R. et al. Unc93B1 biases Toll-like receptor responses to nucleic acid in dendritic cells toward DNA- but against RNA-sensing. J. Exp. Med. 206, 1339–1350 (2009).
Majer, O. et al. Release from UNC93B1 reinforces the compartmentalized activation of select TLRs. Nature 575, 371–374 (2019).
Majer, O., Liu, B., Kreuk, L. S. M., Krogan, N. & Barton, G. M. UNC93B1 recruits syntenin-1 to dampen TLR7 signalling and prevent autoimmunity. Nature 575, 366–370 (2019).
Huh, J. W. et al. UNC93B1 is essential for the plasma membrane localization and signaling of Toll-like receptor 5. PNAS 111, 7072–7077 (2014).
Kim, J. et al. Acidic amino acid residues in the juxtamembrane region of the nucleotide-sensing TLRs are important for UNC93B1 binding and signaling. J. Immunol. 190, 5287–5295 (2013).
Brinkmann, M. M. et al. The interaction between the ER membrane protein UNC93B and TLR3, 7, and 9 is crucial for TLR signaling. J. Cell Biol. 177, 265–275 (2007).
Liu, L. et al. Structural basis of Toll-like receptor 3 signaling with double-stranded RNA. Science 320, 379–381 (2008).
Choe, J., Kelker, M. S. & Wilson, I. A. Crystal structure of human Toll-like receptor 3 (TLR3) ectodomain. Science 309, 581–585 (2005).
Yan, N. Structural biology of the major facilitator superfamily transporters. Annu. Rev. Biophys. 44, 257–283 (2015).
Perland, E., Bagchi, S., Klaesson, A. & Fredriksson, R. Characteristics of 29 novel atypical solute carriers of major facilitator superfamily type: evolutionary conservation, predicted structure and neuronal co-expression. Open Biol. 7, 170142 (2017).
Zhang, B. et al. Structure of a proton-dependent lipid transporter involved in lipoteichoic acids biosynthesis. Nat. Struct. Mol. Biol. 27, 561–569 (2020).
Dang, S. Y. et al. Structure of a fucose transporter in an outward-open conformation. Nature 467, 734–738 (2010).
Pedersen, B. P. et al. Crystal structure of a eukaryotic phosphate transporter. Nature 496, 533–536 (2013).
Zhang, Z. et al. Structural analysis reveals that Toll-like receptor 7 is a dual receptor for guanosine and single-stranded RNA. Immunity 45, 737–748 (2016).
Tanji, H., Ohto, U., Shibata, T., Miyake, K. & Shimizu, T. Structural reorganization of the Toll-like receptor 8 dimer induced by agonistic ligands. Science 339, 1426–1429 (2013).
Pelka, K. et al. The chaperone UNC93B1 regulates Toll-like receptor stability independently of endosomal TLR transport. Immunity 48, 911–922.e7 (2018).
Morales-Perez, C. L., Noviello, C. M. & Hibbs, R. E. Manipulation of subunit stoichiometry in heteromeric membrane proteins. Structure 24, 797–805 (2016).
Kim, J. H. et al. High cleavage efficiency of a 2A peptide derived from porcine teschovirus-1 in human cell lines, zebrafish and mice. PLoS ONE 6, e18556 (2011).
Goehring, A. et al. Screening and large-scale expression of membrane proteins in mammalian cells for structural studies. Nat. Protoc. 9, 2574–2585 (2014).
Mastronarde, D. N. Automated electron microscope tomography using robust prediction of specimen movements. J. Struct. Biol. 152, 36–51 (2005).
Zivanov, J. et al. New tools for automated high-resolution cryo-EM structure determination in RELION-3. Elife 7, e42166 (2018).
Zheng, S. Q. et al. MotionCor2: anisotropic correction of beam-induced motion for improved cryo-electron microscopy. Nat. Methods 14, 331–332 (2017).
Rohou, A. & Grigorieff, N. CTFFIND4: fast and accurate defocus estimation from electron micrographs. J. Struct. Biol. 192, 216–221 (2015).
Kucukelbir, A., Sigworth, F. J. & Tagare, H. D. Quantifying the local resolution of cryo-EM density maps. Nat. Methods 11, 63–65 (2014).
Pettersen, E. F. et al. UCSF Chimera—a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D Biol. Crystallogr. 60, 2126–2132 (2004).
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D Biol. Crystallogr. 66, 213–221 (2010).
Xu, Y. et al. Structural basis for signal transduction by the Toll/interleukin-1 receptor domains. Nature 408, 111–115 (2000).
Acknowledgements
We thank A. Tsutsumi, Y. Sakamaki, M. Kikkawa (Cryo-EM facility in the University of Tokyo) and M. Yamamoto (RIKEN RSC Cryo-EM facility) for their help in cryo-EM data collection. We thank R. Fukui and K. Miyake for their helpful discussion on this manuscript. This work was supported by a Grant-in-Aid from the Japanese Ministry of Education, Culture, Sports, Science, and Technology grant nos. 20K16274 (H.I.), 26711002 (U.O.) and 19H00976 (T.S.); CREST, JST (T.S.); the Takeda Science Foundation (U.O. and T.S.); the Mochida Memorial Foundation for Medical and Pharmaceutical Research (U.O.); the Daiichi Sankyo Foundation of Life Science (U.O.); the Uehara Memorial Foundation (U.O. and T.S.); and the Naito Foundation (U.O. and T.S.). This work is partially supported by Platform Project for Supporting Drug Discovery and Life Science Research (Basis for Supporting Innovative Drug Discovery and Life Science Research (BINDS)) from Japan Agency for Medical Research and Development (AMED) under grant number JP19am0101115, JP19am0101070 (to H.S.). This work was supported in part by the RIKEN Dynamic Structural Biology project (to H.S.).
Author information
Authors and Affiliations
Contributions
H.I. and U.O. designed the experiments. H.I. prepared recombinant proteins. H.I. and J.A. performed pull-down experiments. H.I. and U.O. performed cryo-EM analyses with assistance from Z.Z., T.N. and H.S. H.I., U.O. and T.S. wrote the paper with assistance from all the authors. U.O. and T.S. supervised the project.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Structural & Molecular Biology thanks Gregory Barton, Bostjan Kobe and Eicke Latz for their contribution to the peer review of this work. Anke Sparmann was the primary editor on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Sample preparation and cryo-EM analysis of human TLR3-UNC93B1 complex.
a Representative size-exclusion chromatography profile of human TLR3-UNC93B1 complex (left) and SDS-PAGE analysis of the purified sample stained with Coomassie blue (right). Pooled fractions are shown with the bar. Absorbances at 280 nm and 260 nm are shown with solid and dashed lines, respectively. An uncropped image is available as Supplementary Data. b Data processing workflow of cryo-EM analysis of human TLR3-UNC93B1 complex. Representative motion-corrected micrograph, 2D class averages, and 3D class-averages, gold-standard FSC curve of the final 3D reconstruction (resolution cutoff at FSC = 0.143), and the final 3D-map colored according to the local resolution are shown. The 2D class averages were calculated using the refined particles that was used for the final reconstruction. The 3D classes selected for following analysis are indicated with red boxes.
Extended Data Fig. 2 Sample preparation and cryo-EM analysis of mouse TLR3-UNC93B1 complex.
a Representative size-exclusion chromatography profile of mouse TLR3-UNC93B1 complex (left) and SDS-PAGE analysis of the purified sample stained with Coomassie blue (right). Pooled fractions are shown with the bar. Absorbances at 280 nm and 260 nm are shown with solid and dashed lines, respectively. An uncropped image is available as Supplementary Data. b Data processing workflow of cryo-EM analysis of mouse TLR3-UNC93B1 complex. Representative motion-corrected micrograph, 3D class-averages, Gold-standard FSC curve of the final 3D reconstruction (resolution cutoff at FSC = 0.143), and the final 3D-map colored according to the local resolution are shown. The 3D classes selected for following analysis are indicated with red boxes.
Extended Data Fig. 3 Cryo-EM density maps.
a Cryo-EM density maps of human TLR3-UNC93B1 complex around each TM helix of TLR3 and UNC93B1 are shown. b Cryo-EM density map for N-glycans of human TLR3-UNC93B1 complex. The Asn residues to which N-glycans were attached are labeled. c Cryo-EM density map for N-glycans of human TLR7-UNC93B1 complex. The Asn residues to which N-glycans were attached are labeled.
Extended Data Fig. 4 Structural comparisons of TLR3-UNC93B1 complex.
a Structural superposition of human and mouse TLR3-UNC93B1 complexes and sequence alignment of TLR3 from various species. The Cys residues forming conserved disulfide bonds in LRR-CT are highlighted with yellow. b Structural comparison between the cryo-EM structure of UNC93B1-bound TLR3 and the crystal structures of unliganded human TLR320 (PDB 1ZIW) and dsRNA-bound mouse TLR319 (PDB 3CIY). The UNC93B1 molecule in the TLR3-UNC93B1 complex, and the dsRNA and the other protomer of TLR3 in the dsRNA-bound TLR3 are shown in semitransparent representation.
Extended Data Fig. 5 Structural similarity with major facilitator superfamily transporters.
Structural similarity of UNC93B1 with major facilitator superfamily transporters. The UNC93B1 from human TLR3-UNC93B1 complex (this study), lipoteichoic acids flippase23 (PDB 6S7V), fucose/H+ symporter24 (PDB 3O7Q), and phosphate transporter25 (PDB 4J05). The N- and C-termini are indicated. The structures are colored from blue to red from the N to C terminus.
Extended Data Fig. 6 Sample preparation and cryo-EM analysis of human TLR7-UNC93B1 complex.
a Representative size-exclusion chromatography profile of human TLR7-UNC93B1 complex (left) and SDS-PAGE analysis of the purified sample stained with Coomassie blue (right). Pooled fractions are shown with the bar. Absorbances at 280 nm and 260 nm are shown with solid and dashed lines, respectively. Uncropped images are available as Supplementary Data. b Data processing workflow of cryo-EM analysis of human TLR7-UNC93B1 complex. Representative motion-corrected micrograph, 2D class averages, and 3D class-averages, Gold-standard FSC curve of the final 3D reconstruction (resolution cutoff at FSC = 0.143), and the final 3D-map colored according to the local resolution are shown. The 2D class averages were calculated using the refined particles that was used for the final reconstruction. The 3D classes selected for following analysis are indicated with red boxes.
Extended Data Fig. 7 Comparison of LRR-CT, juxtamembrane, and TM regions from different members of TLRs.
a Sequence alignment of LRR-CT, juxtamembrane, and TM regions from different members of TLRs. The conserved Phe and His residues among TLR3, TLR7, TLR8, TLR9 in the TM regions are indicated by red boxes. b Structural comparison of the LRR-CT, juxtamembrane, and TM regions of TLR3 from TLR3-UNC93B1 complex (this study, left), TLR7 from TLR7-UNC93B1 complex (this study, right). The two conserved disulfide bonds in LRR-CT are shown with sticks. c Electrostatic surface potential of the TLR7-UNC93B1 complex. Positive and negative electrostatic potentials are shown in blue and red, respectively.
Supplementary information
Supplementary Information
Supplementary Fig. 1.
Supplementary Data
Unprocessed gel image for Figs. 1b and 2f and Extended Data Figs. 1a, 2a and 6a.
Source data
Source Data Fig. 2
Statistical source data
Rights and permissions
About this article
Cite this article
Ishida, H., Asami, J., Zhang, Z. et al. Cryo-EM structures of Toll-like receptors in complex with UNC93B1. Nat Struct Mol Biol 28, 173–180 (2021). https://doi.org/10.1038/s41594-020-00542-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41594-020-00542-w
This article is cited by
-
An immunoinformatics and structural vaccinology approach to design a novel and potent multi-epitope base vaccine targeting Zika virus
BMC Chemistry (2024)
-
Future opportunities in solute carrier structural biology
Nature Structural & Molecular Biology (2024)
-
Interface Gain-of-Function Mutations in TLR7 Cause Systemic and Neuro-inflammatory Disease
Journal of Clinical Immunology (2024)
-
The role of TLR7 agonists in modulating COVID-19 severity in subjects with loss-of-function TLR7 variants
Scientific Reports (2023)
-
The architecture of transmembrane and cytoplasmic juxtamembrane regions of Toll-like receptors
Nature Communications (2023)