Abstract
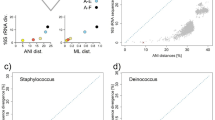
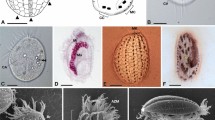
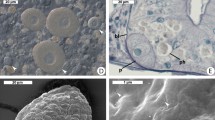
THE use of Koch's technique to isolate bacteria in pure cultures has enabled thousands of bacterial species to be characterized. But for the many microorganisms that have never been cultivated, DNA amplification in vitro using the polymerase chain reaction is now making their genes accessible1–3. Here we use this technique to study bacteria of the genus Holospora, which live in ciliates4 and whose phylogenetic relationship has remained unknown because they are impossible to cultivate. Species of Holospora are highly infectious5–7 and live in the nuclei of their specific host cells: H. elegans and H. undulata infect micronuclei of Paramecium caudatum8, whereas H. obtusa infects the macronucleus in other strains of the same host species9; Holospora species have a common developmental cycle10–13. We have amplified, cloned and sequenced gene fragments encoding ribosomal RNA of H. obtusa. The phylogenetic position of H. obtusa in the α group of Proteobacteria was determined by 16S rRNA sequence analysis. The sequences were then used to design species- as well as genus-specific rRNA hybridization probes, which enabled us to detect and differentiate individual cells of the endosymbionts in situ. The large amount of rRNA in the cells indicates a high physiological activity of the endosymbionts in the host nuclei.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
References
Giovannoni, S. J., Britschgi, T. B., Moyer, C. L. & Field, K. G. Nature 345, 60–63 (1990).
Ward, D., Weller, R. & Bateson, M. M. Nature 345, 63–65 (1990).
Saiki, R. K. et al. Science 239, 487–491 (1988).
Preer, J. R. Jr & Preer, L. B. Int. J. syst. Bact. 32, 140–141 (1984).
Hafkine, M. W. A. Rev. Inst. Pasteur 4, 148–162 (1890).
Gromov, B. V. & Ossipov, D. V. Int. J. Syst. Bact. 31, 348–352 (1981).
Preer, J. R. Jr & Preer, L. B. In Bergeys Manua. of Systematic Bacteriology (eds Sneath, P. H. A., Mair, N. S., Sharpe, M. E. & Holt, J. G.) 795–813 (Williams & Wilkins, Baltimore, 1984).
Görtz, H. D. & Dieckmann, J. Protistologica 16, 591–603 (1980).
Ossipov, D. V., Skoblo, I. I. & Rautian, M. S. Acta protozool. 14, 263–280 (1975).
Görtz, H. D. Int. Rev. Cytol. 102, 169–213 (1986).
Görtz, H. D., Ahlers, H. & Robenek, H. J. gen. Microbiol. 135, 3079–3085 (1989).
Görtz, H. D., Lellig, S., Miosga, O. & Wiemann, M. J. Bact. 172, 5664–5669 (1990).
Fujishima, M., Nagahara, K. & Kojima, Y. Zool. Sci. 7, 849–860 (1990).
Woese, C. R. Microbiol. Rev. 51, 221–271 (1987).
Stackebrandt, E., Murray, R. G. E. & Trüper, H. G. Int. J. syst. Bact. 38, 321–325 (1988).
Yang, D., Oyaizu, Y., Oyaizu, K., Olsen, G. J. & Woese, C. R. Proc. natn. Acad. Sci. U.S.A. 82, 4443–4447 (1985).
Edman, J. C. et al. Nature 334, 519–522 (1988).
Giovannoni, S. J., DeLong, E. F., Olsen, G. J. & Pace, N. R. J. Bact. 170, 720–726 (1988).
DeLong, E. F., Wickham, G. S. & Pace, N. R. Science 243, 1360–1363 (1989).
Amann, R. I., Krumholz, L. & Stahl, D. A. J. Bact. 172, 762–770 (1990).
Amann, R. I. et al. Appl. environ. Microbiol. 56, 1919–1925 (1990).
Torsvik, V., Goksoyr, J. & Daae, F. L. Appl. envir. Microbiol. 56, 782–787.
Hori, H. & Osawa, S. Proc. natn. Acad. Sci. U.S.A. 76, 381–385 (1979).
Saitou, N. & Nei, M. Molec. Biol. Evol. 4, 406–425 (1987).
Neefs, J. M., Van de Peer, Y., Hendriks, L. & De Wachter, R. Nucleic Acids Res. 18, 2237–2318 (1990).
Dryden, S. C. & Kaplan, S. Nucleic Acids Res. 18, 7267–7277 (1990).
Schmidt, H. J., Freiburg, M. & Görtz, H. D. Microbios. 49, 189–197 (1987).
Chen, E. Y. & Seeburg, P. H. DNA 4, 165–170 (1985).
Woese, C. R., Kandler, O. & Wheelis, H. L. Proc. natn. Acad. Sci. U.S.A. 87, 4576–4579 (1990).
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Amann, R., Springer, N., Ludwig, W. et al. Identification in situ and phylogeny of uncultured bacterial endosymbionts. Nature 351, 161–164 (1991). https://doi.org/10.1038/351161a0
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/351161a0
This article is cited by
-
Comparative Analysis Between Paramecium Strains with Different Syngens Using the RAPD Method
Microbial Ecology (2022)
-
Clarification of the Taxonomic Position of Paramecium caudatum Micronucleus Symbionts
Current Microbiology (2021)
-
Response of Anaerobic Protozoa to Oxygen Tension in Anaerobic System
International Microbiology (2019)
-
A House for Two—Double Bacterial Infection in Euplotes woodruffi Sq1 (Ciliophora, Euplotia) Sampled in Southeastern Brazil
Microbial Ecology (2016)
-
Evaluation of Enrichment Protocols for Bacterial Endosymbionts of Ciliates by Real-Time PCR
Current Microbiology (2016)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.