ABSTRACT
Using a combination of hybridization of PAC to a cDNA library and RACE technique, we isolated a novel cDNA, designated as C17orf25 (Chromosome 17 open reading frame 25, previously named it HC71A), from the deletion region on chromosome 17p13.3. The cDNA encodes a protein of 313 amino acids with a calculated molecular mass of 34.8 kDa. C17orf25 is divided into 10 exons and 9 introns, spanning 23 kb of genomic DNA. Northern blot analysis showed that the mRNA expression of C17orf25 was decreased in hepatocellular carcinoma samples as compared to adjacent noncancerous liver tissues from the same patients. The transfection of C17orf25 into the hepatocellular carcinoma cell SMMC7721 and overexpression could inhibit the cell growth. The above results indicate that C17orf25 is a novel human gene, and the cloning and preliminary characterization of C17orf25 is a prerequisite for further functional analysis of this novel gene in human hepatocellular carcinoma.
Similar content being viewed by others
INTRODUCTION
Loss of heterozygosity (LOH) at chromosomal loci associated with tumor suppressor genes has been implicated in the genesis of many types of human malignancies. On the basis of frequent LOH in tumors, coupled with linkage analysis in some hereditary cancer syndromes, a number of tumor suppressor genes, such as RB1, DCC2, NF23, VHL4, MTS15, DPH2L/OVCA16, and PTEN/MMAC17 have been successively isolated.
It has been reported that LOH occurred at 17p in various types of human cancers, including colorectal carcinoma8, breast cancer9, hepatocellular carcinoma (HCC)10, 11, 12, medulloblastoma13, primitive neuroectodermal tumor 14, astrocytoma15, and prostate carcinoma16. Previous studies in our laboratory implied that some new tumor suppressor(s) or cancer-related gene(s) might exist on chromosome 17p13.3 distinct from the p53 gene and LOH at 17p13.3 in HCC were not obligatorily accompanied by mutation or LOH of p5310. Since the same region has shown frequent LOH in many different human cancers, we anticipated that some novel genes residing on chromosome 17p13.3 might be involved in the development of many cancers. Through detailed deletion mapping analysis in a large number of HCC samples, we have defined a 500 kb deletion region at 17p13.3, flanked by loci D17S1840 and D17S654. Subsequently, the bacterial artificial chromosomes (BACs) and bacteriophage P1-derived artificial chromosomes (PACs) covering the 500 kb region have been isolated (published in a separate article, manuscript in preparation). In this paper, we report the cloning and preliminary characterization of C17orf25 isolated from the one of PACs, PAC579, containing the locus D17S926 which has the most frequent LOH in HCC in the deletion region on human chromosome 17p13.3.
MATERIALS AND METHODS
Isolation, labeling of PAC579 and screening of cDNA library
The PAC579 containing the locus D17S926 in the deletion region of chromosome 17p13.3 was obtained from the GenomeSystem Inc. cDNA clones were isolated by screening the human liver cDNA library (Clontech) using PAC579 DNA as a probe. A full-length cDNA, designated as C17orf25, was obtained by rapid amplification of cDNA end technique.
PAC579 DNA was isolated from the NS3516 bacteria17, 18. The insert of the human genomic DNA in PAC579 was prepared for being used as probes. 1 μg of PAC579 DNA was digested with the restriction endonuclease Not I to release the genomic DNA insert. DNA samples were loaded onto 1% agarose gels and fractionated by pulsed field gel electrophoresis (PFGE) in 0.25×TBE for 30 h using a CHEF-DR II DRIVE MODULE (Bio-Rad) at 200 V with pulse times of 5 to 10 s19, 20. The insert size of the PAC579 was also determined by pulsed field gel electrophoresis. The insert of the PAC579 was excised from the gel and DNA concentration estimated by fluorescence examination. Approximately 150 ng of purified PAC579 DNA was radioactively labeled using a random-priming kit (Amersham) to get a specific radioactivity of 1×109 cpm/mg. Labeled probes were then purified by spin column of Sephadex G-50. Probes were denatured and allowed to prehybridize in 5×SSC with 1 μg/μl denatured Cot1 DNA and 0.36 μg/μl denatured sheared pPAC vector DNA in a total volume of 1 ml for 20 min at 65°C.
An oligo(dT) plus random prime human liver cDNA library constructed in λgt11 phage vector system was used in this study (Clontech). It was plated at a density of no more than 40,000 plaques per 15 cm plate and lysis was allowed to continue until plaques were almost confluent during the first screening cycle. Plaque lift filters were prehybridized and hybridized in the solution with 6×SSC, 5×Denhardts, 0.5% SDS, 200 mg/ml denatured sheared salmon sperm DNA and 100 μg/ml denatured sheared human placental DNA. Filters were prehybridized for 12-16 h and hybridized for 18 h. Post-hybridization washes were 2×SSC/0.1% SDS for 30 min at 37°C, and 0.5×SSC/0.1% SDS until the radioactivity measured by a b-ray counter (Mini-Instruments series 900 mini-monitor) was less than 10 cps. Autoradiography was conducted at −70°C for 3-5 d. Duplicate plaque lifts were only used in the first screening cycle.
5′- RACE technique
A ∼ 1.6 kb fragment of C17orf25 cDNA was obtained by screening the human liver cDNA library using PAC579 DNA as a probe. The cDNA was sequenced and found to be lacking the 5′-untranslated region (5′-UTR) and ATG start codon. To isolate the 5′ sequence of the C17orf25 cDNA, primers for 5′-RACE, R1 and R2, were designed from the 3′-UTR region of the C17orf25 cDNA (Fig 1). The primary 5′-RACE PCR was performed by using primers AP1 (Adaptor Primer 1, 5′-CCATCCTAATACGAC TCACTATAGGGC-3′) and R1 (5′-GCGTGCAGCAACGTCACACACTC-3′) for 37 cycles of touchdown PCR with the Human Liver Marathon kit (Clontech) and the Advantage cDNA polymerase (Clontech). The cycling conditions that we used were as follows: 94°C 1 min, 1 cycle; 94°C 30 s, 72°C 4 min, 5 cycles; 94oC 30 s, 70°C 4 min, 5 cycles; 94°C 20 s, 68°C 4 min, 27 cycles. The nested RACE PCR was done by using primers AP2 (Adaptor Primer 2, 5′-ACTCACTATAGGGCTCGAGCGG C-3′) and R2 (5′-GTATCGTCAGGCGCTGGGAATGG-3′) under the same conditions as the primary 5′-RACE PCR. RACE amplification product was then purified and cloned into the vector pT-Adv (Clontech) and sequenced by M13 reverse and forward primers. After 5′-RACE PCR, a full-length cDNA was then obtained and designated as C17orf25.
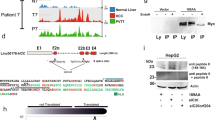
Nucleotide sequence of human C17orf25 cDNA and its deduced amino acid sequence. In the nucleotide sequence, the underlined triplet refers to the in-frame stop codon that lies upstream to the first ATG; 5′ RACE sequence is indicated by the dotted line (bp 1-170); two putative poly(A) signal (AATAAA) are identified by the letters highlighted in gray; the asterisk indicates the stop codon; the arrows indicate the sequence and orientation of 5′ RACE primers. In the deduced protein sequence, the boxed letters indicate a putative protein kinase C phosphorylation site; the double-boxed letters indicate a putative casein kinase II phosphorylation site; the boxed letters in dotted line indicate a N-glycosylation site; the boxed letters highlighted in gray indicate the four putative N-myristoylation sites.
PAC579 DNA sequencing and analysis
The genomic sequence of PAC579 was determined using the shotgun sequencing approach. This work was accomplished by Shanghai GeneCore Biotechnologies Inc. PAC579 DNA sequencing was performed by utilizing the ABI Prism Dye Terminator Cycle Sequencing kit (Perkin-Elmer), and the reactions were analyzed by an ABI Prism 377 DNA Sequencer (Perkin-Elmer). DNA and the predicted amino acid sequences were analyzed by the Genetics Computer Group (GCG) Sequence Analysis Software Package (Wisconsin Package™ version).
Northern blot analysis
A multiple tissue northern (MTN) blot (Clontech) and a northern blot with 8 pairs of hepatocellular carcinoma (HCC) tissues and adjacent noncancerous liver tissues from the same patients were hybridized with the DNA insert (1.6 kb) of the C17orf25 fragment. The MTN blot contains poly (A)-selected RNAs from eight different human tissues. The C17orf25 cDNA probe was labeled with [α-32P]dCTP by random priming method. The RNA hybridization buffer contained 6×SSC, 5×Denhardt's solution, 50% formamide, 0.5% SDS and 100 μg/ml of denatured sheared salmon sperm DNA. The blots were hybridized at 42°C overnight and washed with 2×SSC/1% SDS twice at 37°C for 30 min each, followed by autoradiography21.
Colony formation assay
The C17orf25 cDNA containing the complete ORF (bp 1-1384) was cloned into a eukaryotic expression vector pCMV-Script with a strong CMV promoter (neo+, Stratagene). The plasmid DNA of pCMV-Script/C17orf25 and vector DNA of pCMV-Script were extracted and purified by using QIAGEN plasmid purification system. SMMC7721 cells (a gift of Shanghai Second Military Medical University) grown in 60 mm dishes were transfected with pCMV-Script control or pCMV-Script/C17orf25 (6 μg each per 60mm dish) by means of lipofectamine under conditions recommended by the manufacturer (Gibco BRL). The medium was replaced with fresh, complete medium at 8 hours following transfection. On the next day, the transfected cells were selected in the medium containing 800 μg/ml of G418. Cells were incubated for 14 d, fixed with 10% acetate/10% methanol for 15 min, and stained with 0.4% crystal violet in 20% ethanol for 15 min to examine the colonies23. The effect on colony formation of hepatoma cells was examined as compared with vector control.
RESULTS
cDNA isolation from PAC579 and cloning of C17orf25
To identity novel genes from the common deletion region on chromosome 17p13.3, a PAC clone designated as PAC579 was isolated from a human PAC library using STS-specific primers of D17S926 locus which has the most frequent LOH in human HCC (GenomeSystem Inc). PAC579 contained a 110 kb artificial chromosome recombinant DNA according to results of pulsed field gel electrophoresis (data not shown). The DNA of PAC579 was used as a probe to isolate specific cDNA clones from the human liver cDNA library in λgt11 vector (Clontech). Initially, 78 cDNA clones have been isolated after three rounds of hybridization screening. All these cDNA clones were partially sequenced at both ends and the sequencing data were analyzed and compared with genomic sequence of PAC579. Non-specific and duplicate or overlapping clones were discarded. Finally, 13 unique, and non-overlapping cDNA clones were isolated from the genomic DNA region of PAC579. In this report, we describe one of these clones, a novel gene, designated as C17orf25. This original cDNA of C17orf25 was sequenced and found to be missing the partial 5′ end sequence (bp 1 ∼ 170, Fig 1).
To isolate the 5′ sequences of C17orf25, primers (R1 and R2) were designed and used for 5′ RACE experiments. The reaction extended the sequence of C17orf25 to the 5-untranslated sequence (5′ UTR), as shown in Fig 1. The composite sequence is 1814 bp with a single ORF of 313 amino acids (MW=34791). The first ATG codon (bp 25 ∼ 27) with flanking sequences in consistent with Kozak consensus was designated as the methionine initiation codon. There were two polyadenylation signals (AATAAA) in the 3′-untranslated sequences.
Organization of the human C17orf25
The genomic organization of C17orf25 was determined by comparison and analysis of the cDNA sequence of C17orf25 (Genbank accession No. AF177342) and PAC579 genomic sequence (Genbank accession No. AF258545). C17orf25 comprised 10 exons and 9 introns, and spanning ∼ 23 kb (Tab 1).
Homologies of C17orf25 with other proteins
C17orf25 sequence has been registered in Genbank database (Aug 12, 1999). Amino acid sequence comparison of C17orf25 using BLAST in SwissProt database revealed that the protein of C17orf25 is partially homologous to hypothetical 32kd protein C16C10.10 in chromosome III, a putative homologue of glyoxalase, in caenorhabditis elegans. Percentage of identity of amino acid sequences were found to be 42% between C17orf25 and the hypothetical glyoxalase in caenorhabditis elegans, and 27% between C17orf25 and glyoxalase in brassica olerecea (Fig 2). Interestingly, the protein of C17orf25 is not homologous to human glyoxalase I and II. Since only partial identity has been found at a low percentage of identity between C17orf25 and the hypothetical protein in caenorhabditis elegans or brassica oleracea, it seems that C17orf25 might be a gene encoding a human protein with other function than glyoxalase.
Comparison of the amino acid sequence of the human C17orf25, putative glyoxalase of caenorhabditis elegans, and putative glyoxalase I of brassica oleracea. Identical amino acid residues were highlighted in black, and similar amino acid residues in gray. The amino acid sequence is represented by single-letter abbreviations. GXL Ce: Glyoxalase of C. elegans; GXL Bo: Glyoxalase of B. oleracea.
Expression of human C17orf25
A ∼ 1.6 kb fragment of C17orf25 cDNA (bp 170-1814) was hybridized to the human multiple-tissue northern blot. Result showed a single transcript of ∼ 1.8 kb in human heart, brain, liver, kidney, pancreas and placenta, but no expression in skeletal muscle and lung tissues (Fig 3). These data also indicated that the full-length cDNA sequence (1814 bp) of C17orf25 was correspondence with the size of C17orf25 mRNA transcript. Northern blot analysis in HCC tissues showed that the expression of C17orf25 was decreased in HCC samples as compared to adjacent noncancerous liver tissues in patient D24, G2, G11 and G16 (4/8) (Fig 4). To determine the possible occurrence of mutation present in C17orf25 from HCC, we sequenced the PCR products from the full coding region of C17orf25 from 20 pairs of genomic DNA extracted from HCC and adjacent noncancerous liver tissues. However, no mutations/alterations have so far been identified in these samples.
Tissue expression pattern of C17orf25. Northern blot analysis was performed using human MTN blots, which contained 2 μg of poly(A) RNA from different human tissues in each lane. The human tissues from which mRNAs were isolated were shown at the top. The size of internal RNA standards (kb) was indicated on the left. A transcript of ∼1.8 kb has been observed in heart, brain, placenta, liver, kidney, pancreas. Lower panel: blot was reprobed with β actin.
C17orf25 expression analysis in hepatocellular carcinoma. Northern blots containing RNAs from tumor (T) and adjacent noncancerous liver (N) tissues in patients with primary hepatocellular carcinoma from China were probed with 32P-labeled C17orf25 cDNA. The patient numbers are shown above the blots. Lower panel: blot was reprobed with GAPDH.
Growth suppression effect of C17orf25 to HCC SMMC7721 cell
We examined the growth suppressive effect of C17orf25 on the colony formation efficiency on human HCC SMMC7721 cells by colony formation assay. After 14 days of selection in G418, C17orf25 suppressed the colony formation of the transfected SMMC7721 cells. The amounts of colonies in SMMC7721 cells transfected with C17orf25 were decreased as compared with the result in cells transfected with pCMV-Script vector control (Fig 5).
DISCUSSION
In this study, we have isolated C17orf25 in the deletion region of chromosome 17p13.3 by a combination of cDNA library screening using PAC 579 as a probe and RACE method. The gene is distributed in a genomic area about 23 kb, comprising 10 exons in total. C17orf25 is a novel gene, since the same sequence has not been found with any known human genes at either nucleic acid or protein level in database. A putative full length human gene transcript, CGI-15024 , was also registered in Genbank database (Genbank accession No. AF151908; May 18, 20000). CGI-150 was obtained by assembling in silico cloning method through the comparative gene identification (CGI) with caenorhabditis elegans. After sequence comparison between CGI-150 and C17orf25, we found that the CGI-150 was an incorrect composite sequence misassembled by the full-length C17orf25 cDNA and the partial fragment of another gene. Sequence analysis between PAC579 genomic sequence and CGI-150 also revealed that the CGI-150 was the misconnected sequence at the 5′ end of C17orf25 cDNA with the other gene in PAC579. The expression pattern of C17orf25 and its size of mRNA transcript[Fig 3] also demonstrated that the size of CGI-150 (2580 bp) was incorrect as an artifact. Therefore, the CGI-150 registered in Genbank was an incorrect composite sequence misassembled by the entire C17orf25 cDNA with a fragment from another gene. Lai et al also admitted in their article that the comparative gene identification (CGI) method had some limitations inherent from dbEST. Gene transcripts are often fragmented and not easily assembled into one continuous contig by using nucleotide sequences as the primary alignment basis, such as UniGene and HGI database. It seems essential for the strategy of comparative gene identification to eliminate nucleotide sequence errors in database.
As the 17p13.3 region is heterozygously deleted with a high frequency in many types of human cancer tissues and their cell lines, we investigated the change of mRNA expression. The preliminary study of the expression of C17orf25 was reduced obviously as compared with the matched noncancerous liver tissues in HCC samples. The transfection of pCMV-Script plasmid containing C17orf25 into the human HCC SMMC7721 cells and overexpression of C17orf25 shown that C17orf25 was able to inhibit the colony formation of human hepatoma cell in vitro, which provided a clue to link this novel gene to the regulation of cell growth.
Although the C17orf25 protein does share considerable identities with glyoxalase from caenorhabditis elegans and brassica oleracea, but no obvious homology is found between it and human glyoxylase. The observation of partial homology of the C17orf25 protein sequence to C16C10.10 (a hypothetical glyoxalase in caenorhabditis elegans) seems unlikely to provide obvious clues to the function of this gene in vivo. Nevertheless, we could not exclude so far the possibility that C17orf25 would be a distant member of glyoxalse system or glyoxalase-like enzyme family till the function of C17orf25 has been extensively examined. Human glyoxalase I is a glutathione-binding protein involved in the detoxification of methylglyoxal, a by-product of glycolysis, and other a-oxoaldehydes25. This enzyme catalyzes the conversion of the methylglyoxal-glutathione conjugate to S-D lactoylglutathione, which in turn is hydrolyzed by glyoxalase II into D-lactate and glutathione (GSH). This cytosolic enzyme system is among the earliest expressed proteins during the embryogenesis and development and it would persist throughout all the course of maturation, adult life and senescence26. The main physiological substrates for the glyoxalase system are methylglyoxal formed from the Embden-Meyerhof pathway and glyxal formed from lipid peroxidation and glycation reactions27. Aberrations in the expression of human glyoxalase in cancer and diabetes have been reported. Concentrations of methylglyoxal, S-D lactoyl glutathione and D-lactate were found to be elevated in the blood samples of both insulin dependent and independent diabetic patients, compared to normal healthy controls28, 29, 30. Ayoub et al31 described that the glyoxalase metabolite, S-D-lactoyl glutathione, might mediate the anti-proliferative effects. Another study by Di Ilio et al32 measured glyoxalase I and glyoxalase II activities in urogenital tumor and non-tumorous tissues and found decreased glyoxalase I levels in 10 out of 15 kidney tumors when compared with corresponding normal kidney tissue. Elevated levels of glyoxalase I were also reported in human prostate cancer33. Studies from Ranganathan et al34 showed elevations in glyxalase I activities in 16 out of 21 colon tumor samples as compared to corresponding normal colon tissues. Considering about the expression of C17orf25 in multiple types of human tissues and the result of homology analysis between C17orf25 and homologue in caenorhabditis elegans or brassica olerecea, we postulated that C17orf25 was evolutionarily conserved. This study can be regarded as a contribution to the goal of the human genome project to identify and fully characterize all human genes. The detailed mechanism of C17orf25 in the development of cancer and the relationship between C17orf25 and human glyoxylase system remain to be elucidated in further investigation.
References
Friend SH, Bernards R, Rogelj S, Weinberg RA, Rapaport JM, Albert DM, Dryja TP . A human DNA segment with properties of the gene that predisposes to retinoblastoma and osteosarcoma. Nature 1986; 323:643–46.
Fearon ER, Cho KR, Nigro JM, Kern SE, Simons JW, Ruppert JM, Hamilton SR, Preisinger AC, Thomas G, Kinzler KW, Vogelstein B . Identification of a chromosome 18q gene that is altered in colorectal cancers. Science 1990; 247:49–56.
Rouleau GA, Merel P, Lutchman M, Sanson M, Zucman J, Marineau C, Hoang-Xuan K, Demczuk S, Desmaze C, Plougastel B, Pulst SM, Lenoir G, Bijsma E, Fashold R, Dumanski J, de Jong P, Parry D, Eldrige R, Aurias A, Delattre O, Thomas G . Alteration in a new gene encoding a putative membrane-organizing protein causes neuro-fibromatosis type 2. Nature 1993; 363:515–21.
Latif F, Tory K, Gnarra J, Yao M, Duh F-M, Orcutt ML, Stackhouse T, Kuzmin I, Modi W, Geil L, Schmidt L, Zhou F, Li H, Wei MH, Chen F, Glenn G, Choyke P, Walther MM, Weng Y, Duan D-SR, Dean M, Glava D, Richards FM, Crossey PA, Ferguson-Smith MA, Paslier DL, Chumakov I, Cohen D, Chinault AC, Maher ER, Linehan WM, Zbar B, Lerman MI . Identification of the von Hippel-Lindau disease tumor suppressor gene. Science 1993; 260:1317–20.
Kamb A, Gruis NA, Weaver-Feldhaus J, Liu Q, Harshman K, Tavtigian SV, Stockert E, Day RS III, Johnson BE, Skolnick MH . A cell cycle regulator potentially involved in genesis of many tumor types. Science 1994; 264:436–40.
Schultz DC, Vanderveer L, Berman DB, Hamilton TC, Wong AJ, Godwin AK . Identification of two candidate tumor suppressor genes on chromosome 17p13.3. Cancer Res 1996; 56:1997–2002.
Li J, Yen C, Liaw D, Podsypanina K, Bose S, Wang SI, Puc J, Miliaresis C, Rodgers L, McCombie R, Bigner SH, Giovanella BC, Ittmann M, Tycko B, Hibshoosh H, Wigler MH, Parsons R . PTEN, a putative protein tyrosine phosphatase gene mutated in human brain, breast, and prostate cancer. Science 1997; 275:1943–47.
Lothe RA, Fossli T, Danielsen HE, Stenwig AE, Nesland JM, Gallie B, Borresen, AL . Molecular genetic studies of tumor suppressor gene regions on chromosome 13 and 17 in colorectal tumors. J Natl Cancer Inst 1992; 84:1100–8.
Stack M, Jones D, White G, Liscia DS, Venesio T, Casey G, Crichton D, Varley J, Mitchell E, Heighway J, Santibanez-Koref M . Detail mapping and loss of heterozygosity analysis suggests a suppressor locus involved in sporadic breast cancer within a distal region of chromosome band 17p13.3. Hum Mol Genet 1995; 4:2047–55.
Li D, Cao Y, He L, Wang NJ, Gu JR . Aberations of p53 gene in human hepatocellular carcinoma from china. Carcinogenesis 1993; 14:169–73.
Nishida N, Fukuda Y, Kokuryu H, Toguchida J, Yandell DW, Ikenega M, Imura H, Ishizaki K . Role and mutational heterogeneity of the p53 gene in hepatocellularb carcinoma. Cancer Res 1993; 53:368–72.
Wang G, Huang CH, Zhao Y, Cai L, Wang Y, Xiu SJ, Jiang ZW, Yang S, Zhao XT, Huang W, Gu JR . Genetic aberration in primary hepatocellular carcinoma: correlation between p53 gene mutation and loss-of-heterozygosity on chromosome 16q21-q23 and 9p21-p23. Cell Res 2000; 10:311–23.
Cogen PH, Daneshvar L, Metzger AK, Duyk G., Edwards MS, Sheffield VC . Involvement of multiple chromosome 17p loci in medulloblastoma tumorigenesis. Am J Hum Genet 1992; 50:584–9.
Biegel JA, Burk CD, Barr FG, Emanuel BS . Evidence for a 17p tumor related locus distinct from p53 in pediatric primitive neuroectodermal tumors. Cancer Res 1992; 52:3391–5.
Saxena A, Clark WC, Robertson JT, Ikejiri B, Oldfield EH, Ali IU . Evidence for the involvement of a potential second tumor suppressor gene on chromosome 17 distinct from p53 in malignant astrocytomas. Cancer Res 1992; 52:6716–21.
Ittmann MM . Loss of heterozygosity on chromosomes 10 and 17 in clinically localized prostate carcinoma. Prostate 1996; 28:275–81.
Ioannou PA, Amemiya CT, Garnes J, Kroisel PM, Shizuya H, Chen C, Batzer MA, de Jong PJ . A new bacteriophge P1-derived vector for the propagation of large human DNA fragment. Nat Genet 1994; 6:84–9.
Birnboim HC, Doly J . A rapid alkaline extraction procedure for screening recombinant plasmid DNA. Nucleic Acids Res 1979; 7:1513–23.
Schwartz DC, Cantor CR . Separation of yeast chromosome-sized DNAs by pulsed field gradient gel electrophoresis. Cell 1984; 37:67–75.
Vollrath D, Davis RW . Resolution of DNA molecules greater than 5 megabases by contour-clamped homogeneous electric fields. Nucleic Acids Res 1987; 15:7865–7876.
Sambrook J, Fritsch EF, Maniatis T . In: Molecular cloning: a laboratory manual, second edition, Cold Spring Harbor Laboratory Press, Cold Spring Harbor, New York 1989.
Qin X-Q, Chittenden T, Livingston DM, Kaelin WG Jr . Identification of a growth suppression domain within the retinoblastoma gene product. Genes Dev 1992; 6:953–64.
Taniura H, Matsumoto K, Yoshikawa K . Physical and functional interactions of neuronal growth suppressor necdin with p53. J Biol Chem 1999; 274:16242–8.
Lai C-H, Chou C-Y, Ch'ang L-Y, Liu C-S, Lin W-C . Identification of novel human genes evolutionarily conserved in caenorhabditis elegans by comparative proteomics. Genome Res 2000; 10:703–13.
Thornalley PJ . The glyoxalase system: new developments towards functional characterization of a metabolic pathway fundamental to biological life. Biochem J 1990; 269:1–11.
McLellan AC, Thornalley PJ . Glyoxalase activity in human red blood cells 363 fractioned by age. Mech Ageing Dev 1989; 48:63–71.
Thornalley PJ . Cell activation by glycated proteins. AGE receptors, receptor recognition factors and functional classifiction of AGEs. Cell Mol Biol 1998; 44:1013–23.
Thornalley PJ, Hooper NI, Jennings PE, Florkowski CM, Jones AF, Lunec J, Barnett AH . The human red blood cell glyoxalase system in diabetes mellitus. Diabetes Res Clin Pract 1989; 7:115–120.
McLellan AC, Thornalley PJ . Sample storage conditions for the assay of glyoxalase activities in whole blood samples. Ann Clin Biochem 1992; 29:222–3.
McLellan AC, Thornalley PJ, Benn J, Sonksen PH . Modification of the glyoxalase system in clinical diabetes mellitus. Biochem Soc Trans 1993; 21:158S
Ayoub FM, Allen RE, Thornalley PJ . Inhibition of proliferation of human leukaemia 60 cells by methylglyoxal in vitro. Leuk Res 1993; 17:397–401.
Di Ilio C, Angelucci S, Pennelli A, Zezza A, Tenaglia R, Sacchetta P . Glyoxalase activities in tumor and non-tumor human urogenital tissues. Cancer Lett 1995; 96:189–93.
Davidson SD, Cherry JP, Choudhury MS, Tazaki H, Mallouh C, Konno S . Glyoxalase I activity in human prostate cancer: a potential marker and importance in chemotherapy. J Urol 1999; 161:690–1.
Ranganathan S, Tew KD . Analysis of glyoxalase-I from normal and tumor tissue from human colon. Biochim Biophys Acta. 1993; 1182:311–6.
Acknowledgements
This work was supported by the National 863 High Technology Research and Development Program of China (Z19-02-01-01) to Wan DF and the Project of Chinese National Human Genome Center at Shanghai (CNCS-99M-08) to Qin WX.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
QIN, W., WAN, D., SUN, F. et al. Cloning and characterization of a novel gene (C17orf25) from the deletion region on chromosome 17p13.3 in hepatocelular carcinoma. Cell Res 11, 209–216 (2001). https://doi.org/10.1038/sj.cr.7290088
Received:
Revised:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/sj.cr.7290088
Keywords
This article is cited by
-
Immunoscreening of the extracellular proteome of colorectal cancer cells
BMC Cancer (2010)
-
Cloning and characterization of human IC53-2, a novel CDK5 activator binding protein
Cell Research (2003)
-
The inhibition of lung cancer cell growth by intracellular immunization with LC-1 ScFv
Cell Research (2002)
-
The ATF/CREB site is the key element for transcription of the human RNA methyltransferase like 1 (RNMTL1) gene, a newly discovered 17p13.3 gene
Cell Research (2002)
-
The promoter analysis of the human C17orf25 gene, a novel chromosome 17p13.3 gene
Cell Research (2002)