Abstract
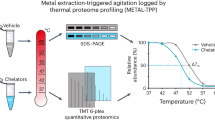
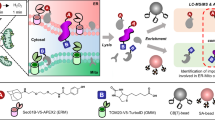
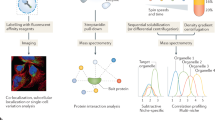
Zinc signaling and dynamics play significant roles in many physiological responses and diseases. To understand the physiological roles of zinc in detail, comprehensive identification of proteins under high concentration of mobile zinc ion is crucial. We developed a 'conditional proteomics' approach to identify proteins involved in zinc homeostasis based on a chemical proteomic strategy that utilizes designer zinc-responsive labeling reagents to tag such proteins and quantitative mass spectrometry for their identification. We used this method to elucidate zinc dyshomeostasis induced by nitric-oxide-triggered oxidative stress in glioma cells, and we unveiled dynamic changes of the zinc-related proteomes. Moreover, we characterized unknown zinc-rich vesicles generated by oxidative stress as endoplasmic-reticulum- and Golgi-related vesicles.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Frederickson, C.J., Koh, J.Y. & Bush, A.I. The neurobiology of zinc in health and disease. Nat. Rev. Neurosci. 6, 449–462 (2005).
Sensi, S.L., Paoletti, P., Bush, A.I. & Sekler, I. Zinc in the physiology and pathology of the CNS. Nat. Rev. Neurosci. 10, 780–791 (2009).
Chang, C.J. Searching for harmony in transition-metal signaling. Nat. Chem. Biol. 11, 744–747 (2015).
Komatsu, K., Kikuchi, K., Kojima, H., Urano, Y. & Nagano, T. Selective zinc sensor molecules with various affinities for Zn2+, revealing dynamics and regional distribution of synaptically released Zn2+ in hippocampal slices. J. Am. Chem. Soc. 127, 10197–10204 (2005).
Nolan, E.M. et al. Zinspy sensors with enhanced dynamic range for imaging neuronal cell zinc uptake and mobilization. J. Am. Chem. Soc. 128, 15517–15528 (2006).
Domaille, D.W., Que, E.L. & Chang, C.J. Synthetic fluorescent sensors for studying the cell biology of metals. Nat. Chem. Biol. 4, 168–175 (2008).
Yuet, K.P. et al. Cell-specific proteomic analysis in Caenorhabditis elegans. Proc. Natl. Acad. Sci. USA 112, 2705–2710 (2015).
Woo, C.M., Iavarone, A.T., Spiciarich, D.R., Palaniappan, K.K. & Bertozzi, C.R. Isotope-targeted glycoproteomics (IsoTaG): a mass-independent platform for intact N- and O-glycopeptide discovery and analysis. Nat. Methods 12, 561–567 (2015).
Hulce, J.J., Cognetta, A.B., Niphakis, M.J., Tully, S.E. & Cravatt, B.F. Proteome-wide mapping of cholesterol-interacting proteins in mammalian cells. Nat. Methods 10, 259–264 (2013).
Lanning, B.R. et al. A road map to evaluate the proteome-wide selectivity of covalent kinase inhibitors. Nat. Chem. Biol. 10, 760–767 (2014).
Niphakis, M.J. et al. A global map of lipid-binding proteins and their ligandability in cells. Cell 161, 1668–1680 (2015).
Rhee, H.W. et al. Proteomic mapping of mitochondria in living cells via spatially restricted enzymatic tagging. Science 339, 1328–1331 (2013).
Lam, S.S. et al. Directed evolution of APEX2 for electron microscopy and proximity labeling. Nat. Methods 12, 51–54 (2015).
Fujishima, S.H., Yasui, R., Miki, T., Ojida, A. & Hamachi, I. Ligand-directed acyl imidazole chemistry for labeling of membrane-bound proteins on live cells. J. Am. Chem. Soc. 134, 3961–3964 (2012).
Miki, T. et al. LDAI-based chemical labeling of intact membrane proteins and its pulse-chase analysis under live cell conditions. Chem. Biol. 21, 1013–1022 (2014).
Finney, L.A. & O'Halloran, T.V. Transition metal speciation in the cell: insights from the chemistry of metal ion receptors. Science 300, 931–936 (2003).
Wei, G., Hough, C.J., Li, Y. & Sarvey, J.M. Characterization of extracellular accumulation of Zn2+ during ischemia and reperfusion of hippocampus slices in rat. Neuroscience 125, 867–877 (2004).
Cuajungco, M.P. & Lees, G.J. Nitric oxide generators produce accumulation of chelatable zinc in hippocampal neuronal perikarya. Brain Res. 799, 118–129 (1998).
Frederickson, C.J., Maret, W. & Cuajungco, M.P. Zinc and excitotoxic brain injury: a new model. Neuroscientist 10, 18–25 (2004).
Colvin, R.A., Holmes, W.R., Fontaine, C.P. & Maret, W. Cytosolic zinc buffering and muffling: their role in intracellular zinc homeostasis. Metallomics 2, 306–317 (2010).
Sensi, S.L. et al. Modulation of mitochondrial function by endogenous Zn2+ pools. Proc. Natl. Acad. Sci. USA 100, 6157–6162 (2003).
Bossy-Wetzel, E. et al. Crosstalk between nitric oxide and zinc pathways to neuronal cell death involving mitochondrial dysfunction and p38-activated K+ channels. Neuron 41, 351–365 (2004).
Haase, H. & Beyersmann, D. Intracellular zinc distribution and transport in C6 rat glioma cells. Biochem. Biophys. Res. Commun. 296, 923–928 (2002).
Haase, H. & Beyersmann, D. Uptake and intracellular distribution of labile and total Zn(II) in C6 rat glioma cells investigated with fluorescent probes and atomic absorption. Biometals 12, 247–254 (1999).
Dayon, L. et al. Relative quantification of proteins in human cerebrospinal fluids by MS/MS using 6-plex isobaric tags. Anal. Chem. 80, 2921–2931 (2008).
Takamori, S. et al. Molecular anatomy of a trafficking organelle. Cell 127, 831–846 (2006).
Hwang, J.J., Lee, S.J., Kim, T.Y., Cho, J.H. & Koh, J.Y. Zinc and 4-hydroxy-2-nonenal mediate lysosomal membrane permeabilization induced by H2O2 in cultured hippocampal neurons. J. Neurosci. 28, 3114–3122 (2008).
Liuzzi, J.P., Guo, L., Yoo, C. & Stewart, T.S. Zinc and autophagy. Biometals 27, 1087–1096 (2014).
Lee, S.J., Cho, K.S. & Koh, J.Y. Oxidative injury triggers autophagy in astrocytes: the role of endogenous zinc. Glia 57, 1351–1361 (2009).
Brandizzi, F. & Barlowe, C. Organization of the ER-Golgi interface for membrane traffic control. Nat. Rev. Mol. Cell Biol. 14, 382–392 (2013).
Allan, B.B., Moyer, B.D. & Balch, W.E. Rab1 recruitment of p115 into a cis-SNARE complex: programming budding COPII vesicles for fusion. Science 289, 444–448 (2000).
Hamlin, J.N.R. et al. Scyl1 scaffolds class II Arfs to specific subcomplexes of coatomer through the γ-COP appendage domain. J. Cell Sci. 127, 1454–1463 (2014).
Ben-Tekaya, H., Miura, K., Pepperkok, R. & Hauri, H.P. Live imaging of bidirectional traffic from the ERGIC. J. Cell Sci. 118, 357–367 (2005).
Que, E.L. et al. Quantitative mapping of zinc fluxes in the mammalian egg reveals the origin of fertilization-induced zinc sparks. Nat. Chem. 7, 130–139 (2015).
Jeong, J. et al. Promotion of vesicular zinc efflux by ZIP13 and its implications for spondylocheiro dysplastic Ehlers-Danlos syndrome. Proc. Natl. Acad. Sci. USA 109, E3530–E3538 (2012).
Roh, H.C., Collier, S., Guthrie, J., Robertson, J.D. & Kornfeld, K. Lysosome-related organelles in intestinal cells are a zinc storage site in C. elegans. Cell Metab. 15, 88–99 (2012).
Ishihama, Y., Rappsilber, J., Andersen, J.S. & Mann, M. Microcolumns with self-assembled particle frits for proteomics. J. Chromatogr. A 979, 233–239 (2002).
Olsen, J.V. et al. Parts per million mass accuracy on an Orbitrap mass spectrometer via lock mass injection into a C-trap. Mol. Cell. Proteomics 4, 2010–2021 (2005).
Olsen, J.V., Ong, S.E. & Mann, M. Trypsin cleaves exclusively C-terminal to arginine and lysine residues. Mol. Cell. Proteomics 3, 608–614 (2004).
Acknowledgements
We thank S. Kitagawa and D. Umeyama (Kyoto University, Japan) for X-ray structure determination. This work was supported by the Japan Science and Technology Agency (JST) Core Research for Evolutional Science and Technology (CREST) of Molecular Technology for I.H. and a Research Fellowship from the Japan Society for the Promotion of Science (JSPS) for Young Scientists for T.M. (25-4988) and Y.N. (16J10316).
Author information
Authors and Affiliations
Contributions
T.M. and I.H. designed the project. T.M., M.A., and Y.N. performed synthesis and chemical labeling. T.M., M.W., and Y.I. performed LC–MS/MS analysis. T.M. and S.K. constructed the expression plasmids. The manuscript was written by T.M. and I.H., according to discussion with all authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–19 and Supplementary Note (PDF 14425 kb)
Supplementary Table 1
The list of labeled peptides by AIZin-1 in HeLa cell lysates (XLSX 37 kb)
Supplementary Table 2
The protein list and statistical analyses for proteomic data of labeled proteins in NO-stimulated C6 cells. (XLSX 366 kb)
Supplementary Table 3
The peptide sequences identified in the proteomic analysis of labeled proteins in NO-stimulated C6 cells. (XLSX 507 kb)
Supplementary Table 4
The protein list for proteomic analysis of labeled proteins in NO-stimulated C6 cells in the presence or absence of TPEN. (XLSX 84 kb)
Source data
Rights and permissions
About this article
Cite this article
Miki, T., Awa, M., Nishikawa, Y. et al. A conditional proteomics approach to identify proteins involved in zinc homeostasis. Nat Methods 13, 931–937 (2016). https://doi.org/10.1038/nmeth.3998
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth.3998
This article is cited by
-
Zinc homeostasis governed by Golgi-resident ZnT family members regulates ERp44-mediated proteostasis at the ER-Golgi interface
Nature Communications (2023)
-
Organelle membrane-specific chemical labeling and dynamic imaging in living cells
Nature Chemical Biology (2020)
-
Split-BioID a conditional proteomics approach to monitor the composition of spatiotemporally defined protein complexes
Nature Communications (2017)
-
Trail-blAIZin new directions for conditional proteomics
Nature Methods (2016)