Abstract
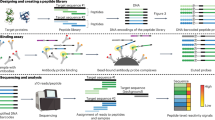
We developed a new, silicon-based peptide array for a broad range of biological applications, including potential development as a real-time point-of-care platform. We used photolithography on silicon wafers to synthesize microarrays (Intel arrays) that contained every possible overlapping peptide within a linear protein sequence covering the N-terminal tail of human histone H2B. These arrays also included peptides with acetylated and methylated lysine residues, reflecting post-translational modifications of H2B. We defined minimum binding epitopes for commercial antibodies recognizing the modified and unmodified H2B peptides. We further found that this platform is suitable for the highly sensitive characterization of methyltransferases and kinase substrates. The Intel arrays also revealed specific H2B epitopes that are recognized by autoantibodies in individuals with systemic lupus erythematosus who have elevated disease severity. By combining emerging nonfluorescence-based detection methods with an underlying integrated circuit, we are now poised to create a truly transformative proteomics platform with applications in bioscience, drug development and clinical diagnostics.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Cortese, D.A. A vision of individualized medicine in the context of global health. Clin. Pharmacol. Ther. 82, 491–493 (2007).
Robinson, W.H. et al. Autoantigen microarrays for multiplex characterization of autoantibody responses. Nat. Med. 8, 295–301 (2002).
Robinson, W.H. et al. Protein microarrays guide tolerizing DNA vaccine treatment of autoimmune encephalomyelitis. Nat. Biotechnol. 21, 1033–1039 (2003).
Neuman de Vegvar, H.E. et al. Microarray profiling of antibody responses against simian-human immunodeficiency virus: postchallenge convergence of reactivities independent of host histocompatibility type and vaccine regimen. J. Virol. 77, 11125–11138 (2003).
Chan, S.M., Ermann, J., Su, L., Fathman, C.G. & Utz, P.J. Protein microarrays for multiplex analysis of signal transduction pathways. Nat. Med. 10, 1390–1396 (2004).
Kattah, M.G., Coller, J., Cheung, R.K., Oshidary, N. & Utz, P.J. HIT: a versatile proteomics platform for multianalyte phenotyping of cytokines, intracellular proteins and surface molecules. Nat. Med. 14, 1284–1289 (2008).
Haab, B.B. Antibody arrays in cancer research. Mol. Cell. Proteomics 4, 377–383 (2005).
Ray, S. et al. Classification and prediction of clinical Alzheimer's diagnosis based on plasma signaling proteins. Nat. Med. 13, 1359–1362 (2007).
Britschgi, M. et al. Neuroprotective natural antibodies to assemblies of amyloidogenic peptides decrease with normal aging and advancing Alzheimer's disease. Proc. Natl. Acad. Sci. USA 106, 12145–12150 (2009).
Vigh-Conrad, K.A., Conrad, D.F. & Preuss, D. A protein allergen microarray detects specific IgE to pollen surface, cytoplasmic, and commercial allergen extracts. PLoS ONE 5, e10174 (2010).
Bailey, R.C. Grand challenge commentary: informative diagnostics for personalized medicine. Nat. Chem. Biol. 6, 857–859 (2010).
Fodor, S.P. et al. Light-directed, spatially addressable parallel chemical synthesis. Science 251, 767–773 (1991).
Gao, X., Zhou, X. & Gulari, E. Light directed massively parallel on-chip synthesis of peptide arrays with t-Boc chemistry. Proteomics 3, 2135–2141 (2003).
Shin, D.S. et al. Automated maskless photolithography system for peptide microarray synthesis on a chip. J. Comb. Chem. 12, 463–471 (2010).
Khorasanizadeh, S. The nucleosome: from genomic organization to genomic regulation. Cell 116, 259–272 (2004).
Kouzarides, T. Chromatin modifications and their function. Cell 128, 693–705 (2007).
Monestier, M., Decker, P., Briand, J.P., Gabriel, J.L. & Muller, S. Molecular and structural properties of three autoimmune IgG monoclonal antibodies to histone H2B. J. Biol. Chem. 275, 13558–13563 (2000).
van Bavel, C.C. et al. Apoptosis-associated acetylation on histone H2B is an epitope for lupus autoantibodies. Mol. Immunol. 47, 511–516 (2009).
Barone, A.D. et al. Photolithographic synthesis of high-density oligonucleotide probe arrays. Nucleosides Nucleotides Nucleic Acids 20, 525–531 (2001).
Lee, M.S. & Kerns, E.H. LC/MS applications in drug development. Mass Spectrom. Rev. 18, 187–279 (1999).
Dhayalan, A., Kudithipudi, S., Rathert, P. & Jeltsch, A. Specificity analysis–based identification of new methylation targets of the SET7/9 protein lysine methyltransferase. Chem. Biol. 18, 111–120 (2011).
Cheung, W.L. et al. Apoptotic phosphorylation of histone H2B is mediated by mammalian sterile twenty kinase. Cell 113, 507–517 (2003).
von Mühlen, C.A. & Tan, E.M. Autoantibodies in the diagnosis of systemic rheumatic diseases. Semin. Arthritis Rheum. 24, 323–358 (1995).
Baechler, E.C. et al. Interferon-inducible gene expression signature in peripheral blood cells of patients with severe lupus. Proc. Natl. Acad. Sci. USA 100, 2610–2615 (2003).
Bauer, J.W. et al. Elevated serum levels of interferon-regulated chemokines are biomarkers for active human systemic lupus erythematosus. PLoS Med. 3, e491 (2006).
Obermoser, G. & Pascual, V. The interferon-α signature of systemic lupus erythematosus. Lupus 19, 1012–1019 (2010).
Dieker, J.W. et al. Apoptosis-induced acetylation of histones is pathogenic in systemic lupus erythematosus. Arthritis Rheum. 56, 1921–1933 (2007).
van Bavel, C.C. et al. Apoptosis-induced histone H3 methylation is targeted by autoantibodies in systemic lupus erythematosus. Ann. Rheum. Dis. 70, 201–207 (2011).
Petri, M. Hopkins Lupus Cohort. 1999 update. Rheum. Dis. Clin. North Am. 26, 199–213, v (2000).
Tusher, V.G., Tibshirani, R. & Chu, G. Significance analysis of microarrays applied to the ionizing radiation response. Proc. Natl. Acad. Sci. USA 98, 5116–5121 (2001).
Baker, M. Upping the ante on antibodies. Nat. Biotechnol. 23, 1065–1072 (2005).
Levy, D. et al. Lysine methylation of the NF-κB subunit RelA by SETD6 couples activity of the histone methyltransferase GLP at chromatin to tonic repression of NF-κB signaling. Nat. Immunol. 12, 29–36 (2011).
Yoo, C.B. & Jones, P.A. Epigenetic therapy of cancer: past, present and future. Nat. Rev. Drug Discov. 5, 37–50 (2006).
Volta, U. et al. Deamidated gliadin peptide antibodies as a routine test for celiac disease: a prospective analysis. J. Clin. Gastroenterol. 44, 186–190 (2010).
Lee, D.M. & Schur, P.H. Clinical utility of the anti-CCP assay in patients with rheumatic diseases. Ann. Rheum. Dis. 62, 870–874 (2003).
Hoffman, I.E. et al. Diagnostic performance and predictive value of rheumatoid factor, anti-citrullinated peptide antibodies, and the HLA shared epitope for diagnosis of rheumatoid arthritis. Clin. Chem. 51, 261–263 (2005).
Garren, H. et al. Phase 2 trial of a DNA vaccine encoding myelin basic protein for multiple sclerosis. Ann. Neurol. 63, 611–620 (2008).
Liskamp, R.M.J., Rijkers, D.T.S., Kruijtzer, J.A.W. & Kemmink, J. Peptides and proteins as a continuing exciting source of inspiration for peptidomimetics. ChemBioChem 12, 1626–1653 (2011).
Gaster, R.S., Hall, D.A. & Wang, S.X. nanoLAB: an ultraportable, handheld diagnostic laboratory for global health. Lab Chip 11, 950–956 (2011).
Acknowledgements
We thank H. Lee, J. Zhu and the other members of the peptide array team at Biomedical Life Science at Digital Health Intel for development of the early peptide array process and G. Credo for help with the Online Methods section of the manuscript. We thank D. Hall, K.C. Garcia, S. Sidhu, J. Haddon, J. Jarrell and other members of our laboratories for helpful discussion. J.V.P. was funded by a US National Science Foundation Graduate Research Fellowship (NSF GRFP) and is supported by the Stanford Genome Training Program (SGTP; US National Institutes of Health (NIH), US National Human Genome Research Institute). D.L. is supported by the European Molecular Biology Organization, the Human Frontier Science Program and the Machiah Foundation. O.G. is a recipient of an Ellison Senior Scholar Award and is funded by a grant from the NIH (R01 GM079641). P.J.U. is the recipient of a Donald E. and Delia B. Baxter Foundation Career Development Award and is supported by National Heart, Lung, and Blood Institute (NHLBI) Proteomics contract HHSN288201000034C, Proteomics of Inflammatory Immunity and Pulmonary Arterial Hypertension; grants from the NIH (5 U19-AI082719, 5 U19-AI050864, 5 U19-AI056363, 1 U19 AI090019 and 4 U19 AI090019); Canadian Institutes of Health Research (2 OR-92141); Alliance for Lupus Research (grant number 21858); a gift from the Ben May Trust and a gift from the Floren Family Trust. The research leading to these results has received funding from the European Union Seventh Framework Programme (FP7/2007-2013) under grant agreement number 261382. C.L.L. is a recipient of an NIH National Research Service Award Fellowship (5 F32 AI-080086-02).
Author information
Authors and Affiliations
Contributions
J.V.P. conducted antibody-binding experiments and ELISA, analyzed the data, made the figures and wrote the manuscript. S.T. performed antibody-binding experiments and contributed to data analysis and figure production. G.X. and J.Y. designed and fabricated Intel arrays. D.L. and O.G. contributed the methylation assay. E.C.B. characterized the IFN signature and provided ABCoN patient samples. M.V. supervised the development team at Intel and edited the manuscript. P.J.U. and C.L.L. supervised the project and edited the manuscript.
Corresponding author
Ethics declarations
Competing interests
M.V. and G.X. are employees of Intel.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–6 and Supplementary Methods (PDF 5462 kb)
Rights and permissions
About this article
Cite this article
Price, J., Tangsombatvisit, S., Xu, G. et al. On silico peptide microarrays for high-resolution mapping of antibody epitopes and diverse protein-protein interactions. Nat Med 18, 1434–1440 (2012). https://doi.org/10.1038/nm.2913
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nm.2913
This article is cited by
-
Immunological fingerprint of 4CMenB recombinant antigens via protein microarray reveals key immunosignatures correlating with bactericidal activity
Nature Communications (2020)
-
Electric fields control the orientation of peptides irreversibly immobilized on radical-functionalized surfaces
Nature Communications (2018)
-
Binding patterns of homo-peptides on bare magnetic nanoparticles: insights into environmental dependence
Scientific Reports (2017)
-
A Simple Platform for the Rapid Development of Antimicrobials
Scientific Reports (2017)
-
Multiplex giant magnetoresistive biosensor microarrays identify interferon-associated autoantibodies in systemic lupus erythematosus
Scientific Reports (2016)