Abstract
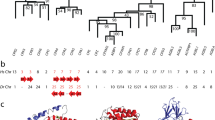
THE sequences of proteins from ancient organisms can be reconstructed from the sequences of their descendants by a procedure that assumes that the descendant proteins arose from the extinct ancestor by the smallest number of independent evolutionary events (‘parsimony’)1,2. The reconstructed sequences can then be prepared in the laboratory and studied3,4. Thirteen ancient ribonucleases (RNases) have been reconstructed as intermediates in the evolution of the RNase protein family in artiodactyls (the mammal order that includes pig, camel, deer, sheep and ox)5. The properties of the reconstructed proteins suggest that parsimony yields plausible ancient sequences. Going back in time, a significant change in behaviour, namely a fivefold increase in catalytic activity against double-stranded RNA, appears in the RNase reconstructed for the founding ancestor of the artiodactyl lineage, which lived about 40 million years ago6. This corresponds to the period when ruminant digestion arose in the artiodactyls, suggests that contemporary artiodactyl digestive RNases arose from a non-digestive ancestor, and illustrates how evolutionary reconstructions can help in the understanding of physiological function within a protein family7–9.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
References
Pauling, I. & Zuckerkandl, E. Acta chem. scand. 17 (Suppl. 1), S9—S16 (1963).
Fitch, W. M. Syst. Zool. 20, 406–416 (1971).
Stackhouse, J., Presnell, S. R., McGeehan, G. M., Nambiar, K. P. & Benner, S. A. FEBS Lett. 262, 104–106 (1990).
Malcolm, B. A., Wilson, K. P., Matthews, B. W., Kirsch, J. F. & Wilson, A. C. Nature 345, 86–89 (1990).
Carroll, R. L. Vertebrate Paleontology and Evolution (Freeman, New York, 1988).
Rose, K. D. Science 216, 621–623 (1982).
Nambiar, K. P. et al. Science 223, 1299–1301 (1984).
Adey, N. B., Tollefsbol, T. O., Sparks, A. B., Edgell, M. H. & Hutchinson, C. A. Proc. natn. Acad. Sci. U.S.A. 91, 1569–1573 (1994).
Hillis, D. M., Huelsenbeck, J. P. & Cunningham, C. W. Science 264, 671–677 (1994).
Soucek, J., Chudomel, V., Potmesilova, I. & Novak, J. T. Nat. Immun. Cell Growth Reg. 5, 250–258 (1986).
Matousek, J. Experientia 29, 858 (1973).
Ardelt, W., Mikulski, S. & Shogen, K. J. biol. Chem. 266, 245–251 (1991).
Strydom, D. J. et al. Biochemistry 24, 5486–5494 (1985).
Okabe, Y. et al. J. Biochem. (Tokyo) 109, 786–790 (1991).
Beintema, J. J., Fitch, W. M. & Carsana, A. Molec. Biol. Evol. 3, 262–275 (1986).
Nambiar, K. P., Stackhouse, J., Presnell, S. R. & Benner, S. A. Eur. J. biochem. 163, 67–71 (1987).
McGeehan, G. M. & Benner, S. A. FEBS Lett. 247, 55–56 (1989).
Trautwein, K. & Benner, S. A. in Proc. Int. Symp. Site Directed Mutagenesis and Protein Engineering (ed. El-Gewely, M. R.) 53–61 (Elsevier, New York, 1991).
Blackburn, P. & Moore, S. The Enzymes 15, 317–434 (1982).
Ipata, P. L. & Felicioli, R. A. FEBS Lett. 1, 29–31 (1968).
Lang, K. & Schmidt, F. X. Eur. J. Biochem. 159, 275–281 (1986).
Jolles, J. et al. J. molec. Evol. 28, 528–533 (1989).
Barnard, E. A. Nature 221, 340–344 (1969).
Watanabe, H. et al. J. Biochem. (Tokyo) 104, 939–945 (1988).
Libonati, M. & Floridi, A. Eur. J. Biochem. 8, 81–87 (1969).
Graur, D. FEBS Lett. 325, 152–159 (1993).
Ciglic, M. diplomarbeit, ETH Zürich (1994).
Stevens, C. E. Comparative Physiology of the Vertebrate Digestive System (Cambridge Univ. Press, Cambridge, UK, 1988).
Sorrentino, S., Carsan̄a, A., Furia, A., Doskocil, J. & Libonati, M. Biochim. biophys. Acta 609, 40–52 (1980).
Breukelman, H. J. et al. J. molec. Evol. 37, 29–35 (1993).
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Jermann, T., Opitz, J., Stackhouse, J. et al. Reconstructing the evolutionary history of the artiodactyl ribonuclease superfamily. Nature 374, 57–59 (1995). https://doi.org/10.1038/374057a0
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/374057a0
This article is cited by
-
Ancestral Sequence Reconstruction: From Chemical Paleogenetics to Maximum Likelihood Algorithms and Beyond
Journal of Molecular Evolution (2021)
-
Building genomes to understand biology
Nature Communications (2020)
-
Evolutionary history of versatile-lipases from Agaricales through reconstruction of ancestral structures
BMC Genomics (2017)
-
A simple method for studying the molecular mechanisms of ultraviolet and violet reception in vertebrates
BMC Evolutionary Biology (2016)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.