Abstract
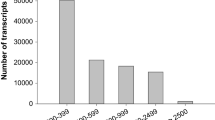
Molecular genetics has identified dozens of genes that regulate flower development in Arabidopsis. However, the complexity of flower development suggests that many other genes are yet to be uncovered. To identify floral genes that are expressed at low levels in the flower, we have sequenced 1587 cDNA fragments from a subtractive floral cDNA library. A total of 1222 unique genes represented by these ESTs are distributed on all five chromosomes with similar frequencies as all predicted genes in the genome. Among these, 17 genes were shown to be expressed anywhere for the first time because they were not found in previous EST and full-length cDNA datasets. Furthermore, 724 of the genes revealed by this library were not definitively shown to be expressed in the flower by previous floral EST datasets. In addition, 49 transcriptional regulators, 31 protein kinases, 12 zinc-finger proteins and other signaling proteins were found to be present in floral buds. Moreover, the EST sequences likely extended the transcribed regions of 26 previously annotated genes, and may have uncovered several previously unrecognized genes. To obtain additional clues about possible gene function, we hybridized cDNA microarray with probes derived from wild-type Arabidopsis rosette leaves and floral buds. We estimated that over 50% of genes were expressed at levels lower than 1/30 of the highest detectable signal intensity, indicating that many floral genes are expressed at low levels. Furthermore, 97 genes were found to be expressed at a higher level in the flower than the leaf by the Significance Analysis of Microarray (SAM) method with a 1.0% false discovery rate (FDR). Further RT-PCR analyses of selected genes support the microarray results. We suggest that the genes encoding putative regulatory proteins and at least some proteins with currently unknown functions might play important roles during flowering.
Similar content being viewed by others
References
AGI. 2000. Analysis of the genome sequence of the flowering plant Arabidopsis thaliana. Nature 408: 796–815.
Allen, M., Qin, W., Moreau, F. and Moffatt, B. 2002. Adenine phosphoribosyltransferase isoforms of Arabidopsis and their potential contributions to adenine and cytokinin metabolism. Physiol. Plant. 115: 56–68.
Amagai, M., Ariizumi, T., Endo, M., Hatakeyama, K., Kuwata, C., Shibata, D., Toriyama, K. and Watanabe, M. 2002. Identification of anther-specific genes in a cruciferous model plant, Arabidopsis thaliana, by using a combination of Arabidopsis macroarray and mRNA from Brassica oleracea. Sex. Plant Reprod. 15: 213–220.
Azumi, Y., Liu, D., Zhao, D., Li, W., Wang, G., Hu, Y. and Ma, H. 2002. Homolog interaction during meiotic prophase I in Arabidopsis requires the SOLO DANCERS gene encoding a novel cyclin-like protein. EMBO J. 21: 3081–3095.
Blazquez, M. 2000. Flower development pathways. J. Cell Sci. 113: 3547–3548.
Bowman, J.L., Smyth, D.R. and Meyerowitz, E.M. 1989. Genes directing flower development in Arabidopsis. Plant Cell 1: 37–52.
Capella, A.N., Menossi, M., Arruda, P. and Benedetti, C.E. 2001. COI1 affects myrosinase activity and controls the expression of two flower-specific myrosinase-binding protein homologues in Arabidopsis. Planta 213: 691–699.
Channeliere, S., Riviere, S., Scalliet, G., Szecsi, J., Jullien, F., Dolle, C., Vergne, P., Dumas, C., Bendahmane, M., Hugueney, P. and Cock, J.M. 2002. Analysis of gene expression in rose petals using expressed sequence tags. FEBS Lett. 515: 35–38.
Chen, C., Marcus, A., Li, W., Hu, Y., Calzada, J.P., Grossniklaus, U., Cyr, R.J. and Ma, H. 2002. The Arabidopsis ATK1 gene is required for spindle morphogenesis in male meiosis. Development 129: 2401–2409.
Chen, Y., Dougherty, E.R. and Bittner, M.L. 1997. Ratio-based decisions and the quantitative analysis of cDNAmicroarray images. J. Biomed. Optics 2: 364–374.
Diatchenko, L., Lau, Y.F., Campbell, A.P., Chenchik, A., Moqadam, F., Huang, B., Lukyanov, S., Lukyanov, K., Gurskaya, N., Sverdlov, E.D. and Siebert, P.D. 1996. Suppression subtractive hybridization: a method for generating differentially regulated or tissue-specific cDNA probes and libraries. Proc. Natl. Acad. Sci. USA 93: 6025–6030.
Diatchenko, L., Lukyanov, S., Lau, Y.F. and Siebert, P.D. 1999. Suppression subtractive hybridization: a versatile method for identifying differentially expressed genes. Meth. Enzymol. 303: 349–380.
Endo, M., Kokubun, T., Takahata, Y., Higashitani, A., Tabata, S. and Watanabe, M. 2000. Analysis of expressed sequence tags of flower buds in Lotus japonicus. DNA Res. 7: 213–216.
Fourgoux-Nicol, A., Drouaud, J., Haouazine, N., Pelletier, G. and Guerche, P. 1999. Isolation of rapeseed genes expressed early and specifically during development of the male gametophyte. Plant Mol. Biol. 40: 857–872.
Gamboa, A., Paez-Valencia, J., Acevedo, G.F., Vazquez-Moreno, L. and Alvarez-Buylla, R.E. 2001. Floral transcription factor AGAMOUS interacts in vitro with a leucine-rich repeat and an acid phosphatase protein complex. Biochem. Biophys. Res. Commun. 288: 1018–1026.
Girke, T., Todd, J., Ruuska, S., White, J., Benning, C. and Ohlrogge, J. 2000. Microarray analysis of developing Arabidopsis seeds. Plant Physiol. 124: 1570–1581.
Goda, H., Shimada, Y., Asami, T., Fujioka, S. and Yoshida, S. 2002. Microarray analysis of brassinosteroid-regulated genes in Arabidopsis. Plant Physiol. 130: 1319–1334.
Gu, Q., Ferrandiz, C., Yanofsky, M.F. and Martienssen, R. 1998. The FRUITFULL MADS-box gene mediates cell differentiation during Arabidopsis fruit development. Development 125: 1509–1517.
Guterman, I., Shalit, M., Menda, N., Piestun, D., Dafny-Yelin, M., Shalev, G., Bar, E., Davydov, O., Ovadis, M., Emanuel, M., Wang, J., Adam, Z., Pichersky, E., Lewinsohn, E., Zamir, D., Vainstein, A. and Weiss, D. 2002. Rose scent: genomics approach to discovering novel floral fragrance-related genes. Plant Cell 14: 2325–2338.
Hofte, H., Desprez, T., Amselem, J., Chiapello, H., Rouze, P., Caboche, M., Moisan, A., Jourjon, M.F., Charpenteau, J.L., Berthomieu, P. et al. 1993. An inventory of 1152 expressed sequence tags obtained by partial sequencing of cDNAs from Arabidopsis thaliana. Plant J. 4: 1051–1061.
Honma, T. and Goto, K. 2001. Complexes of MADS-box proteins are sufficient to convert leaves into floral organs. Nature 409: 525–529.
Huang, H., Mizukami, Y., Hu, Y. and Ma, H. 1993. Isolation and characterization of the binding sequences for the product of the Arabidopsis floral homeotic gene AGAMOUS. Nucl. Acids Res. 21: 4769–4776.
Jofuku, K.D., den Boer, B.G., Van Montagu, M. and Okamuro, J.K. 1994. Control of Arabidopsis flower and seed development by the homeotic gene APETALA2. Plant Cell 6: 1211–1225.
Kamalay, J.C. and Goldberg, R.B. 1980. Regulation of structural gene expression in tobacco. Cell 19: 935–946.
Kieffer, M. and Davies, B. 2001. Developmental programmes in floral organ formation. Semin. Cell Dev. Biol. 12: 373–380.
Lim, C.O., Kim, H.Y., Kim, M.G., Lee, S.I., Chung, W.S., Park, S.H., Hwang, I. and Cho, M.J. 1996. Expressed sequence tags of Chinese cabbage flower bud cDNA. Plant Physiol. 111: 577–588.
Lohmann, J.U. and Weigel, D. 2002. Building beauty: the genetic control of floral patterning. Dev. Cell 2: 135–142.
Ma, L., Li, J., Qu, L., Hager, J., Chen, Z., Zhao, H. and Deng, X.W. 2001. Light control of Arabidopsis development entails coordinated regulation of genome expression and cellular pathways. Plant Cell 13: 2589–2607.
Mandel, M.A. and Yanofsky, M.F. 1995. The Arabidopsis AGL8 MADS box gene is expressed in inflorescence meristems and is negatively regulated by APETALA1. Plant Cell 7: 1763–1771.
Meeks-Wagner, D.R., Dennis, E.S., Tran Thanh Van, K. and Peacock, W.J. 1989. Tobacco genes expressed during in vitro floral initiation and their expression during normal plant development. Plant Cell 1: 25–35.
Moffatt, B.A., McWhinnie, E.A., Agarwal, S.K. and Schaff, D.A. 1994. The adenine phosphoribosyltransferase-encoding gene of Arabidopsis thaliana. Gene 143: 211–216.
Nagai, J., Yamato, K.T., Sakaida, M., Yoda, H., Fukuzawa, H. and Ohyama, K. 1999. Expressed sequence tags from immature female sexual organ of a liverwort, Marchantia polymorpha. DNA Res. 6: 1–11.
Park, H.C., Kang, Y.H., Chun, H.J., Koo, J.C., Cheong, Y.H., Kim, C.Y., Kim, M.C., Chung, W.S., Kim, J.C., Yoo, J.H., Koo, Y.D., Koo, S.C., Lim, C.O., Lee, S.Y. and Cho, M.J. 2002. Characterization of a stamen-specific cDNA encoding a novel plant defensin in Chinese cabbage. Plant Mol. Biol. 50: 59–69.
Pelaz, S., Ditta, G.S., Baumann, E., Wisman, E. and Yanofsky, M.F. 2000. B and C floral organ identity functions require SEPALLATA MADS-box genes. Nature 405: 200–203.
Riechmann, J.L. and Meyerowitz, E.M. 1998. The AP2/EREBP family of plant transcription factors. Biol. Chem. 379: 633–646.
Roberts, M.R., Foster, G.D., Blundell, R.P., Robinson, S.W., Kumar, A., Draper, J. and Scott, R. 1993. Gametophytic and sporophytic expression of an anther-specific Arabidopsis thaliana gene. Plant J. 3: 111–120.
Sablowski, R.W. and Meyerowitz, E.M. 1998. A homolog of NO APICAL MERISTEM is an immediate target of the floral homeotic genes APETALA3/PISTILLATA. Cell 92: 93–103.
Seki, M., Narusaka, M., Abe, H., Kasuga, M., Yamaguchi-Shinozaki, K., Carninci, P., Hayashizaki, Y. and Shinozaki, K. 2001. Monitoring the expression pattern of 1300 Arabidopsis genes under drought and cold stresses by using a full-length cDNA microarray. Plant Cell 13: 61–72.
Seki, M., Ishida, J., Narusaka, M., Fujita, M., Nanjo, T., Umezawa, T., Kamiya, A., Nakajima, M., Enju, A., Sakurai, T., Satou, M., Akiyama, K., Yamaguchi-Shinozaki, K., Carninci, P., Kawai, J., Hayashizaki, Y. and Shinozaki, K. 2002. Monitoring the expression pattern of around 7,000 Arabidopsis genes under ABA treatments using a full-length cDNA microarray. Funct. Integr. Genomics 2: 301.
Sessions, A., Yanofsky, M.F. and Weigel, D. 1998. Patterning the floral meristem. Semin. Cell Dev. Biol. 9: 221–226.
Shiu, S.H. and Bleecker, A.B. 2001. Receptor-like kinases from Arabidopsis form a monophyletic gene family related to animal receptor kinases. Proc. Natl. Acad. Sci. USA 98: 10763–10768.
Siegfried, K.R., Eshed, Y., Baum, S.F., Otsuga, D., Drews, G.N. and Bowman, J.L. 1999. Members of the YABBY gene family specify abaxial cell fate in Arabidopsis. Development 126: 4117–4128.
Smyth, D.R., Bowman, J.L. and Meyerowitz, E.M. 1990. Early flower development in Arabidopsis. Plant Cell 2: 755–767.
Sung, Z.R., Chen, L., Moon, Y.H. and Lertpiriyapong, K. 2003. Mechanisms of floral repression in Arabidopsis. Curr. Opin. Plant Biol. 6: 29–35.
Taipalensuu, J., Falk, A. and Rask, L. 1996. A wound-and methyl jasmonate-inducible transcript coding for a myrosinaseassociated protein with similarities to an early nodulin. Plant Physiol. 110: 483–491.
Takechi, K., Sakamoto, W., Utsugi, S., Murata, M. and Motoyoshi, F. 1999. Characterization of a flower-specific gene encoding a putative myrosinase binding protein in Arabidopsis thaliana. Plant Cell Physiol. 40: 1287–1296.
Takemura, M., Fujishige, K., Hyodo, H., Ohashi, Y., Kami, C., Nishii, A., Ohyama, K. and Kohchi, T. 1999. Systematic isolation of genes expressed at low levels in inflorescence apices of Arabidopsis thaliana. DNA Res. 6: 275–282.
Tusher, V.G., Tibshirani, R. and Chu, G. 2001. Significance analysis of microarrays applied to the ionizing radiation response. Proc. Natl. Acad. Sci. USA 98: 5116–5121.
Utsugi, S., Sakamoto, W., Murata, M. and Motoyoshi, F. 1998. Arabidopsis thaliana vegetative storage protein (VSP) genes: gene organization and tissue-specific expression. Plant Mol. Biol. 38: 565–576.
Weigel, D. 1998. From floral induction to floral shape. Curr. Opin. Plant Biol. 1: 55–59.
Xue, J., Jorgensen, M., Pihlgren, U. and Rask, L. 1995. The myrosinase gene family in Arabidopsis thaliana: gene organization, expression and evolution. Plant Mol. Biol. 27: 911–922.
Yang, Y.H., Dudoit, S., Luu, P., Lin, D.M., Peng, V., Ngai, J. and Speed, T.P. 2002. Normalization for cDNA microarray data: a robust composite method addressing single and multiple slide systematic variation. Nucl. Acids Res. 30: e15.
Zhao, D., Yu, Q., Chen, C. and Ma, H. 2001. Genetic control of reproductive meristems. In: M.T. McManus and B. Veit (Eds.) Annual Plant Review: Meristematic Tissues in Plant Growth and Development, Sheffield Academic Press, Sheffield, UK, pp. 89–142.
Zhao, D.Z., Wang, G.F., Speal, B. and Ma, H. 2002. The excess microsporocytes1 gene encodes a putative leucine-rich repeat receptor protein kinase that controls somatic and reproductive cell fates in the Arabidopsis anther. Genes Dev. 16: 2021–2031.
Zik, M. and Irish, V.F. 2003. Global identification of target genes regulated by APETALA3 and PISTILLATA floral homeotic gene action. Plant Cell 15: 207–222.
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Hu, W., Wang, Y., Bowers, C. et al. Isolation, sequence analysis, and expression studies of florally expressed cDNAs in Arabidopsis . Plant Mol Biol 53, 545–563 (2003). https://doi.org/10.1023/B:PLAN.0000019063.18097.62
Published:
Issue Date:
DOI: https://doi.org/10.1023/B:PLAN.0000019063.18097.62