Abstract
Studies have highlighted the significant role of focal adhesion signaling in cancer. Nevertheless, its specific involvement in the pathogenesis of endometrial cancer and its clinical significance remains uncertain. We analyzed TCGA-UCEC and GSE119041 datasets with corresponding clinical data to investigate focal adhesion-related gene expression and their clinical significance. A signature, "FA-riskScore," was developed using LASSO regression in the TCGA cohort and validated in the GSE dataset. The FA-riskScore was compared with four existing models in terms of their prediction performance. We employed univariate and multivariate Cox regression analyses towards FA-riskScore to assess its independent prognostic value. A prognostic evaluation nomogram based on our model and clinical indexes was established subsequently. Biological and immune differences between high- and low-risk groups were explored through functional enrichment, PPI network analysis, mutation mining, TME evaluation, and single-cell analysis. Sensitivity tests on commonly targeted drugs were performed on both groups, and Connectivity MAP identified potentially effective molecules for high-risk patients. qRT-PCR validated the expressions of FA-riskScore genes. FA-riskScore, based on FN1, RELN, PARVG, and PTEN, indicated a poorer prognosis for high-risk patients. Compared with published models, FA-riskScore achieved better and more stable performance. High-risk groups exhibited a more challenging TME and suppressive immune status. qRT-PCR showed differential expression in FN1, RELN, and PTEN. Connectivity MAP analysis suggested that BU-239, potassium-canrenoate, and tubocurarine are effective for high-risk patients. This study introduces a novel prognostic model for endometrial cancer and offers insights into focal adhesion's role in cancer pathogenesis.
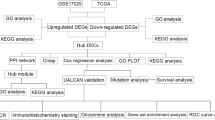
Graphical Abstract









Similar content being viewed by others
Data Availability
Publicly available datasets were analyzed in this study. This data can be found here: (1) https://portal.gdc.cancer.gov/. (2) https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE119041.
Code Availability
All codes used in the study are available from the respective authors upon reasonable request.
References
Makker V, et al. Endometrial cancer. Nat Rev Dis Primers. 2021;7(1):88.
Sung H, et al. Global Cancer Statistics 2020: GLOBOCAN Estimates of Incidence and Mortality Worldwide for 36 Cancers in 185 Countries. CA Cancer J Clin. 2021;71(3):209–49.
Siegel RL, Miller KD, Jemal A. Cancer statistics, 2018. CA Cancer J Clin. 2018;68(1):7–30.
Brooks RA, et al. Current recommendations and recent progress in endometrial cancer. CA Cancer J Clin. 2019;69(4):258–79.
Janiszewska M, Primi MC, Izard T. Cell adhesion in cancer: Beyond the migration of single cells. J Biol Chem. 2020;295(8):2495–505.
Zhang Z, et al. Functional and clinical characteristics of focal adhesion kinases in cancer progression. Front Cell Dev Biol. 2022;10:1040311.
McLean GW, et al. The role of focal-adhesion kinase in cancer - a new therapeutic opportunity. Nat Rev Cancer. 2005;5(7):505–15.
Zhang P, et al. CPNE8 Promotes Gastric Cancer Metastasis by Modulating Focal Adhesion Pathway and Tumor Microenvironment. Int J Biol Sci. 2022;18(13):4932–49.
Tsujioka M, et al. Identification of a novel type of focal adhesion remodelling via FAK/FRNK replacement, and its contribution to cancer progression. Cell Death Dis. 2023;14(4):256.
Chen L, et al. LIM domain-containing 2 (LIMD2) promotes the progress of ovarian cancer via the focal adhesion signaling pathway. Bioengineered. 2021;12(2):10089–100.
Hu X, et al. Collagen triple helix repeat containing 1 promotes endometrial cancer cell migration by activating the focal adhesion kinase signaling pathway. Exp Ther Med. 2020;20(2):1405–14.
Alowayed N, et al. LEFTY2 Controls Migration of Human Endometrial Cancer Cells via Focal Adhesion Kinase Activity (FAK) and miRNA-200a. Cell Physiol Biochem. 2016;39(3):815–26.
Li Z, Gou J, Xu J. Down-regulation of focal adhesion signaling in response to cyclophilin A knockdown in human endometrial cancer cells, implicated by cDNA microarray analysis. Gynecol Oncol. 2013;131(1):191–7.
Gabriel B, et al. Expression of focal adhesion kinase in patients with endometrial cancer: a clinicopathologic study. Int J Gynecol Cancer. 2009;19(7):1221–5.
Tenenbaum D, Volkening J, Bioconductor Package Maintainer. KEGGREST: Client-side REST access to the Kyoto Encyclopedia of Genes and Genomes (KEGG). 2023. https://doi.org/10.18129/B9.bioc.KEGGREST, R package version 1.42.0, https://bioconductor.org/packages/KEGGREST.
Wu T, et al. clusterProfiler 4.0: A universal enrichment tool for interpreting omics data. Innovation. 2021;2(3):100141.
Castanza AS, et al. Extending support for mouse data in the Molecular Signatures Database (MSigDB). Nat Methods. 2023;20:1619–1620. https://doi.org/10.1038/s41592-023-02014-7
Hanzelmann S, Castelo R, Guinney J. GSVA: gene set variation analysis for microarray and RNA-seq data. BMC Bioinformatics. 2013;14:7.
Szklarczyk D, et al. STRING v11: protein-protein association networks with increased coverage, supporting functional discovery in genome-wide experimental datasets. Nucleic Acids Res. 2019;47(D1):D607–13.
Shannon P, et al. Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res. 2003;13(11):2498–504.
Bader GD, Hogue CW. An automated method for finding molecular complexes in large protein interaction networks. BMC Bioinformatics. 2003;4:2.
Geeleher P, Cox N, Huang RS. pRRophetic: an R package for prediction of clinical chemotherapeutic response from tumor gene expression levels. PLoS One. 2014;9(9):e107468.
Subramanian A, et al. A Next Generation Connectivity Map: L1000 Platform and the First 1,000,000 Profiles. Cell. 2017;171(6):1437–1452 e17.
R Foundation for Statistical Computing. R: A language and environment for statistical computing. RA Lang Environ Stat Comput. 2018.
Wei S, et al. Identification of an integrated kinase-related prognostic gene signature associated with tumor immune microenvironment in human uterine corpus endometrial carcinoma. Front Oncol. 2022;12:944000.
Chen S, et al. Development of Biomarker Signatures Associated with Anoikis to Predict Prognosis in Endometrial Carcinoma Patients. J Oncol. 2021;2021:3375297.
Liu J, et al. Development and clinical validation of novel 8-gene prognostic signature associated with the proportion of regulatory T cells by weighted gene co-expression network analysis in uterine corpus endometrial carcinoma. Front Immunol. 2021;12:788431.
Ruan T, et al. Identification of a Novel Epithelial-Mesenchymal Transition-Related Gene Signature for Endometrial Carcinoma Prognosis. Genes. 2022;13(2):216.
Malpica A. How to approach the many faces of endometrioid carcinoma. Mod Pathol. 2016;29(Suppl 1):S29–44.
Bai JDK, et al. Keratin 17 is a negative prognostic biomarker in high-grade endometrial carcinomas. Hum Pathol. 2019;94:40–50.
Siesser PF, Maness PF. L1 cell adhesion molecules as regulators of tumor cell invasiveness. Cell Adh Migr. 2009;3(3):275–7.
Chaudhary PK, Kim S. An Insight into GPCR and G-Proteins as Cancer Drivers. Cells. 2021;10(12):3288.
Filardo EJ, et al. Distribution of GPR30, a seven membrane-spanning estrogen receptor, in primary breast cancer and its association with clinicopathologic determinants of tumor progression. Clin Cancer Res. 2006;12(21):6359–66.
Yun CC, et al. LPA2 receptor mediates mitogenic signals in human colon cancer cells. Am J Physiol Cell Physiol. 2005;289(1):C2–11.
Komachi M, et al. Orally active lysophosphatidic acid receptor antagonist attenuates pancreatic cancer invasion and metastasis in vivo. Cancer Sci. 2012;103(6):1099–104.
Suzuki-Karasaki M, Ochiai T, Suzuki-Karasaki Y. Crosstalk between mitochondrial ROS and depolarization in the potentiation of TRAIL-induced apoptosis in human tumor cells. Int J Oncol. 2014;44(2):616–28.
Sabharwal SS, Schumacker PT. Mitochondrial ROS in cancer: initiators, amplifiers or an Achilles' heel? Nat Rev Cancer. 2014;14(11):709–21.
Huang T, Zhou J, Wang J. Calcium and calcium-related proteins in endometrial cancer: opportunities for pharmacological intervention. Int J Biol Sci. 2022;18(3):1065–78.
Deng Y, et al. High SPRR1A expression is associated with poor survival in patients with colon cancer. Oncol Lett. 2020;19(5):3417–24.
Yamakawa K, et al. Increased expression of SPRR1A is associated with a poor prognosis in pancreatic ductal adenocarcinoma. PLoS One. 2022;17(5):e0266620.
Zhang Z, et al. Identification of small proline-rich protein 1B (SPRR1B) as a prognostically predictive biomarker for lung adenocarcinoma by integrative bioinformatic analysis. Thorac Cancer. 2021;12(6):796–806.
Yu L, et al. Identification of SPRR3 as a novel diagnostic/prognostic biomarker for oral squamous cell carcinoma via RNA sequencing and bioinformatic analyses. PeerJ. 2020;8:e9393.
Yoshida S, et al. Fibronectin mediates activation of stromal fibroblasts by SPARC in endometrial cancer cells. BMC Cancer. 2021;21(1):156.
Yadav VK, et al. Computational analysis for identification of the extracellular matrix molecules involved in endometrial cancer progression. PLoS One. 2020;15(4):e0231594.
Jhunjhunwala S, Hammer C, Delamarre L. Antigen presentation in cancer: insights into tumour immunogenicity and immune evasion. Nat Rev Cancer. 2021;21(5):298–312.
Papageorgis P, Stylianopoulos T. Role of TGFbeta in regulation of the tumor microenvironment and drug delivery (review). Int J Oncol. 2015;46(3):933–43.
Kim M, et al. VEGFA links self-renewal and metastasis by inducing Sox2 to repress miR-452, driving Slug. Oncogene. 2017;36(36):5199–211.
Yang X, et al. VEGF-B promotes cancer metastasis through a VEGF-A-independent mechanism and serves as a marker of poor prognosis for cancer patients. Proc Natl Acad Sci USA. 2015;112(22):E2900–9.
Zhao G, et al. Development and validation of focal adhesion-related genes signature in gastric cancer. Front Genet. 2023;14:1122580.
Li H, et al. A Focal Adhesion-Related Gene Signature Predicts Prognosis in Glioma and Correlates With Radiation Response and Immune Microenvironment. Front Oncol. 2021;11:698278.
Xu X, Wang J. Multi-omics analysis reveals focal adhesion characteristic associated tumor immune microenvironment in colon adenocarcinoma. Front Genet. 2023;14:1088091.
Lin Z, et al. A novel focal adhesion related gene signature for prognostic prediction in hepatocellular carcinoma. Aging. 2021;13(7):10724–48.
Choi DS, et al. Endometrial cancer invasion depends on cancer-derived tumor necrosis factor-alpha and stromal derived hepatocyte growth factor. Int J Cancer. 2009;124(11):2528–38.
Park DW, et al. Gonadotropin-releasing hormone (GnRH)-I and GnRH-II induce cell growth inhibition in human endometrial cancer cells: involvement of integrin beta3 and focal adhesion kinase. Reprod Biol Endocrinol. 2009;7:81.
Tan LH, et al. The characteristics of Ishikawa endometrial cancer cells are modified by substrate topography with cell-like features and the polymer surface. Int J Nanomed. 2015;10:4883–95.
Weijiao Y, et al. Immune infiltration and a ferroptosis-associated gene signature for predicting the prognosis of patients with endometrial cancer. Aging. 2021;13(12):16713–32.
Shan L, et al. Identification of Five m6A-Related lncRNA Genes as Prognostic Markers for Endometrial Cancer Based on TCGA Database. J Immunol Res. 2022;2022:2547029.
Gao L, et al. A prognostic model and immune regulation analysis of uterine corpus endometrial carcinoma based on cellular senescence. Front Oncol. 2022;12:1054564.
Gianfrancesco MA, et al. Potential Biases in Machine Learning Algorithms Using Electronic Health Record Data. JAMA Intern Med. 2018;178(11):1544–7.
Ren S, et al. CRPMKB: a knowledge base of cancer risk prediction models for systematic comparison and personalized applications. Bioinformatics. 2022;38(6):1669–76.
Author information
Authors and Affiliations
Contributions
YZ, LS, and RS conceptualized and designed this project. CY and LH established the model and performed further data analysis based on the TCGA dataset. LH and YM validated the established model based on the GEO dataset. JC and YM conducted wet-lab experimental validation. CY, LS, and RS wrote the whole manuscript. YZ contributed to the funding. All the authors have reviewed the manuscript and approved the submission.
Corresponding authors
Ethics declarations
Conflicts of Interest
The authors declare that they have no conflicts of interest.
Ethics Approval and Consent to Participate
The Ethics Committee of Dushu Lake Hospital, Affiliated with Soochow University, reviewed and approved this study involving human participants.
Consent to Participate
All participants have provided their written informed consent to participate in this study.
Consent for Publication
Written informed consent for publication was obtained from all participants.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Yan, C., He, L., Ma, Y. et al. Establishing and Validating an Innovative Focal Adhesion-Linked Gene Signature for Enhanced Prognostic Assessment in Endometrial Cancer. Reprod. Sci. (2024). https://doi.org/10.1007/s43032-024-01564-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s43032-024-01564-1