Abstract
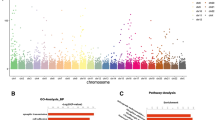
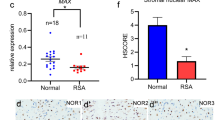
Recurrent pregnancy loss (RPL) is affecting 3–5% of the women who are expecting to have babies annually. However, the molecular changes of RPL patients have not been well characterized before. In the current study, we aimed to discover the differentially expressed genes (DEGs) in decidua from RPL patients by utilizing RNA sequencing (RNA-seq) technology. Four RPL patients and three control women were recruited to conduct RNA-seq, and 153 genes comprising 69 were upregulated and 84 downregulated showed different expression levels between the health control and RPL groups. Gene Ontology (GO) and the Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway enrichment analysis were performed to identify potential biological processes associated with RPL. We recognized that type I interferon signaling pathway and TNF signaling pathway are significantly upregulated in women with RPL. Upregulated expression of MX1, IFI27, and ISG15 in type I interferon signaling pathway and TNFRSF21 in TNF signaling pathway were validated by an extended sample of 15 RPL patients and 12 control women, by quantitative real-time polymerase chain reaction (qRT-PCR). In conclusion, the results of the current study revealed molecular changes in decidua which may reflect the intrauterine circumstance when pregnancy loss happens and potentially contributes to RPL.




Similar content being viewed by others
Data Availability
The datasets used and/or analyzed during the current study are available from the corresponding author on reasonable request.
Code Availability
Not applicable.
Abbreviations
- RPL:
-
Recurrent pregnancy loss
- RIF:
-
Recurrent implantation failure
- DEG:
-
Differentially expressed gene
- RNA-seq:
-
RNA sequencing
- GO:
-
Gene Ontology
- KEGG:
-
Kyoto Encyclopedia of Genes and Genomes
- PCA:
-
Principal component analysis
- qRT-PCR:
-
Quantitative real-time polymerase chain reaction
- TNFRSF21:
-
Tumor necrosis factor receptor superfamily member 21
References
ESHRE Guideline Group on RPL, Bender Atik R, Christiansen OB, Elson J, Kolte AM, Lewis S, et al. ESHRE guideline: recurrent pregnancy loss. Hum Reprod Open. 2018;2018(2):hoy004. https://doi.org/10.1093/hropen/hoy004.
Weimar CHE, Macklon NS, Post Uiterweer ED, Brosens JJ, Gellersen B. The motile and invasive capacity of human endometrial stromal cells: implications for normal and impaired reproductive function. Hum Reprod Update. 2013;19(5):542–57.
Lucas ES, Salker MS, Brosens JJ. Uterine plasticity and reproductive fitness. Reprod BioMed Online. 2013;27(5):506–14.
Brosens JJ, Salker MS, Teklenburg G, Nautiyal J, Salter S, Lucas ES, et al. Uterine selection of human embryos at implantation. Sci Rep. 2014;4:3894.
Altmäe S, Martínez-Conejero JA, Salumets A, Simón C, Horcajadas JA, Stavreus-Evers A. Endometrial gene expression analysis at the time of embryo implantation in women with unexplained infertility. Mol Hum Reprod. 2010;16(3):178–87.
Tong J, Zhao W, Lv H, Li W-P, Chen Z-J, Zhang C. Transcriptomic profiling in human decidua of severe preeclampsia detected by RNA sequencing. J Cell Biochem. 2018;119(1):607–15.
Huang J, Qin H, Yang Y, Chen X, Zhang J, Laird S, et al. A comparison of transcriptomic profiles in endometrium during window of implantation between women with unexplained recurrent implantation failure and recurrent miscarriage. Reproduction. 2017;153(6):749–58.
Chen M-Y, Liao G-D, Zhou B, Kang L-N, He Y-M, Li S-W. Genome-wide profiling of long noncoding RNA expression patterns in women with repeated implantation failure by RNA sequencing. Reprod Sci. 2019;26(1):18–25.
Fu M, Mu S, Wen C, Jiang S, Li L, Meng Y, et al. Whole-exome sequencing analysis of products of conception identifies novel mutations associated with missed abortion. Mol Med Rep. 2018;18(2):2027–32.
Robbins SM, Thimm MA, Valle D, Jelin AC. Genetic diagnosis in first or second trimester pregnancy loss using exome sequencing: a systematic review of human essential genes. J Assist Reprod Genet. 2019;36(8):1539–48.
Sõber S, Rull K, Reiman M, Ilisson P, Mattila P, Laan M. RNA sequencing of chorionic villi from recurrent pregnancy loss patients reveals impaired function of basic nuclear and cellular machinery. Sci Rep. 2016;6:38439.
Trapnell C, Williams BA, Pertea G, Mortazavi A, Kwan G, van Baren MJ, et al. Transcript assembly and quantification by RNA-Seq reveals unannotated transcripts and isoform switching during cell differentiation. Nat Biotechnol. 2010;28(5):511–5.
Kong L, Zhang Y, Ye Z-Q, Liu X-Q, Zhao S-Q, Wei L, et al. CPC: assess the protein-coding potential of transcripts using sequence features and support vector machine. Nucleic Acids Res. 2007;35(Web Server issue):W345–W9.
Szklarczyk D, Gable AL, Lyon D, Junge A, Wyder S, Huerta-Cepas J, et al. STRING v11: protein-protein association networks with increased coverage, supporting functional discovery in genome-wide experimental datasets. Nucleic Acids Res. 2019;47(D1):D607–D13.
Livak KJ, Schmittgen TD. Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) Method. Methods. 2001;25(4):402–8.
Kim D, Langmead B, Salzberg SL. HISAT: a fast spliced aligner with low memory requirements. Nat Methods. 2015;12(4):357–60.
Wang H, Cao Q, Ge J, Liu C, Ma Y, Meng Y, et al. LncRNA-regulated infection and inflammation pathways associated with pregnancy loss: genome wide differential expression of lncRNAs in early spontaneous abortion. Am J Reprod Immunol. 2014;72(4):359–75.
Saifi B, Aflatoonian R, Tajik N, Ahmadpour ME, Vakili R, Amjadi F, et al. T regulatory markers expression in unexplained recurrent spontaneous abortion. J Matern Fetal Neonatal Med. 2015;00(00):1–6.
Cai J, Li M, Huang Q, Fu X, Wu H. Differences in Cytokine Expression and STAT3 Activation between healthy controls and patients of Unexplained Recurrent Spontaneous Abortion (URSA) during early pregnancy. PLoS One. 2016;11(9):e0163252.
Fu B, Wei H. Decidual natural killer cells and the immune microenvironment at the maternal-fetal interface. Sci China Life Sci. 2016;59(12):1224–31.
Fensterl V, Sen GC. Interferons and viral infections. BioFactors. 2009;35(1):14–20.
Moreno I, Franasiak JM. Endometrial microbiota-new player in town. Fertil Steril. 2017;108(1):32–9.
Kushnir VA, Solouki S, Sarig-Meth T, Vega MG, Albertini DF, Darmon SK, et al. Systemic inflammation and autoimmunity in women with chronic endometritis. Am J Reprod Immunol. 2016;75(6):672–7.
Cicinelli E, Matteo M, Tinelli R, Pinto V, Marinaccio M, Indraccolo U, et al. Chronic endometritis due to common bacteria is prevalent in women with recurrent miscarriage as confirmed by improved pregnancy outcome after antibiotic treatment. Reprod Sci. 2014;21(5):640–7.
Cicinelli E, Matteo M, Tinelli R, Lepera A, Alfonso R, Indraccolo U, et al. Prevalence of chronic endometritis in repeated unexplained implantation failure and the IVF success rate after antibiotic therapy. Hum Reprod. 2015;30(2):323–30.
Moreno I, Cicinelli E, Garcia-Grau I, Gonzalez-Monfort M, Bau D, Vilella F, et al. The diagnosis of chronic endometritis in infertile asymptomatic women: a comparative study of histology, microbial cultures, hysteroscopy, and molecular microbiology. Am J Obstet Gynecol. 2018;218(6):602.e1–e16.
Suryawanshi H, Morozov P, Straus A, Sahasrabudhe N, Max KEA, Garzia A, et al. A single-cell survey of the human first-trimester placenta and decidua. Sci Adv. 2018;4(10):eaau4788.
Alijotas-Reig J, Esteve-Valverde E, Ferrer-Oliveras R, Llurba E, Gris JM. Tumor necrosis factor-alpha and pregnancy: focus on biologics. An updated and comprehensive review. Clin Rev Allergy Immunol. 2017;53(1):40–53.
Nikolaev A, McLaughlin T, O’Leary DD, Tessier-Lavigne M. APP binds DR6 to trigger axon pruning and neuron death via distinct caspases. Nature. 2009;457(7232):981–9. https://doi.org/10.1038/nature07767.
Tam SJ, Richmond DL, Kaminker JS, Modrusan Z, Martin-McNulty B, Cao TC, et al. Death receptors DR6 and TROY regulate brain vascular development. Dev Cell. 2012;22(2):403–17.
Zeng L, Li T, Xu DC, Liu J, Mao G, Cui M-Z, et al. Death receptor 6 induces apoptosis not through type I or type II pathways, but via a unique mitochondria-dependent pathway by interacting with Bax protein. J Biol Chem. 2012;287(34):29125–33.
Benschop R, Wei T, Na S. Tumor necrosis factor receptor superfamily member 21: TNFR-related death receptor-6, DR6. Adv Exp Med Biol. 2009;647:186–94.
Wang J-M, Gu Y, Zhang Y, Yang Q, Zhang X, Yin L, et al. Deep-sequencing identification of differentially expressed miRNAs in decidua and villus of recurrent miscarriage patients. Arch Gynecol Obstet. 2016;293(5):1125–35.
Kai K, Nasu K, Kawano Y, Aoyagi Y, Tsukamoto Y, Hijiya N, et al. Death receptor 6 is epigenetically silenced by histone deacetylation in endometriosis and promotes the pathogenesis of endometriosis. Am J Reprod Immunol. 2013;70(6):485–96.
Vento-Tormo R, Efremova M, Botting RA, Turco MY, Vento-Tormo M, Meyer KB, et al. Single-cell reconstruction of the early maternal-fetal interface in humans. Nature. 2018;563(7731):347–53.
Koot YEM, van Hooff SR, Boomsma CM, van Leenen D, Groot Koerkamp MJA, Goddijn M, et al. An endometrial gene expression signature accurately predicts recurrent implantation failure after IVF. Sci Rep. 2016;6:19411.
Qiao Y, Wen J, Tang F, Martell S, Shomer N, Leung PC, et al. Whole exome sequencing in recurrent early pregnancy loss. Mol Hum Reprod. 2016;22(5):364–72.
Quintero-Ronderos P, Mercier E, Fukuda M, Gonzalez R, Suarez CF, Patarroyo MA, et al. Novel genes and mutations in patients affected by recurrent pregnancy loss. PLoS One. 2017;12(10):e0186149.
Popescu F, Jaslow CR, Kutteh WH. Recurrent pregnancy loss evaluation combined with 24-chromosome microarray of miscarriage tissue provides a probable or definite cause of pregnancy loss in over 90% of patients. Hum Reprod. 2018;33(4):579–87.
Hu S, Yao G, Wang Y, Xu H, Ji X, He Y, et al. Transcriptomic changes during the pre-receptive to receptive transition in human endometrium detected by RNA-Seq. J Clin Endocrinol Metab. 2014;99(12):E2744–E53.
Bhagwat SR, Chandrashekar DS, Kakar R, Davuluri S, Bajpai AK, Nayak S, et al. Endometrial receptivity: a revisit to functional genomics studies on human endometrium and creation of HGEx-ERdb. PLoS One. 2013;8(3):e58419.
Murakami K, Lee YH, Lucas ES, Chan Y-W, Durairaj RP, Takeda S, et al. Decidualization induces a secretome switch in perivascular niche cells of the human endometrium. Endocrinology. 2014;155(11):4542–53.
Borthwick JM, Charnock-Jones DS, Tom BD, Hull ML, Teirney R, Phillips SC, et al. Determination of the transcript profile of human endometrium. Mol Hum Reprod. 2003;9(1):19–33.
Altmäe S, Koel M, Võsa U, Adler P, Suhorutšenko M, Laisk-Podar T, et al. Meta-signature of human endometrial receptivity: a meta-analysis and validation study of transcriptomic biomarkers. Sci Rep. 2017;7(1):10077.
Díaz-Gimeno P, Ruiz-Alonso M, Sebastian-Leon P, Pellicer A, Valbuena D, Simón C. Window of implantation transcriptomic stratification reveals different endometrial subsignatures associated with live birth and biochemical pregnancy. Fertil Steril. 2017;108(4):703-710.e3. https://doi.org/10.1016/j.fertnstert.2017.07.007.
Huang J, Jin N, Qin H, Shi X, Liu Y, Cheung W, et al. Transcriptomic profiles in peripheral blood between women with unexplained recurrent implantation failure and recurrent miscarriage and the correlation with endometrium: a pilot study. PLoS One. 2017;12(12):e0189159. https://doi.org/10.1371/journal.pone.0189159.
Bastu E, Demiral I, Gunel T, Ulgen E, Gumusoglu E, Hosseini MK, et al. Potential marker pathways in the endometrium that may cause recurrent implantation failure. Reprod Sci. 2019;26(7):879–90.
Kosova G, Stephenson MD, Lynch VJ, Ober C. Evolutionary forward genomics reveals novel insights into the genes and pathways dysregulated in recurrent early pregnancy loss. Hum Reprod. 2015;30(3):519–29.
Kao LC, Tulac S, Lobo S, Imani B, Yang JP, Germeyer A, et al. Global gene profiling in human endometrium during the window of implantation. Endocrinology. 2002;143(6):2119–38.
Mirkin S, Arslan M, Churikov D, Corica A, Diaz JI, Williams S, et al. In search of candidate genes critically expressed in the human endometrium during the window of implantation. Hum Reprod. 2005;20(8):2104–17.
Haouzi D, Mahmoud K, Fourar M, Bendhaou K, Dechaud H, De Vos J, et al. Identification of new biomarkers of human endometrial receptivity in the natural cycle. Hum Reprod. 2009;24(1):198–205.
Schatz F, Guzeloglu-Kayisli O, Arlier S, Kayisli UA, Lockwood CJ. The role of decidual cells in uterine hemostasis, menstruation, inflammation, adverse pregnancy outcomes and abnormal uterine bleeding. Hum Reprod Update. 2016;22(4):497–515.
Acknowledgments
We thank Chaoyang Sun for the great help for bioinformatic analysis during data processing.
Funding
This study was supported by the National Key Research and Development Project (2018YFC1002103).
Author information
Authors and Affiliations
Contributions
Y. L. designed the experiments. Y. L., R. W., and W. H. performed the experiments. Y. L. performed data analysis, constructed the figures, and drafted the manuscript. S. L. and L. J. assisted in the design of the experiments. M. W. and Z. F. revised the manuscript. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Ethics Approval
This study was approved by the Institutional Review Board of Tongji Hospital (No. TJ-IRB20181110).
Consent to Participate
Not applicable.
Consent for Publication
Not applicable.
Competing Interests
The authors declare no competing interests.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Fig. S1
Placental marker genes expressed by each sample. Heat map showing relative expression of placental marker genes in each sequenced sample (X24, X44, and X45 belong to the control group; X46, X47, X49, and X50 belong to the RPL group). The placental marker genes were provided by Suryawanshi et al. [27] (PNG 699 kb).
ESM 1
(XLSX 12 kb).
ESM 2
(XLS 65077 kb).
ESM 3
(XLS 599 kb).
ESM 4
(XLS 36 kb).
ESM 5
(XLS 26 kb).
Rights and permissions
About this article
Cite this article
Li, Y., Wang, R., Wang, M. et al. RNA Sequencing of Decidua Reveals Differentially Expressed Genes in Recurrent Pregnancy Loss. Reprod. Sci. 28, 2261–2269 (2021). https://doi.org/10.1007/s43032-021-00482-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s43032-021-00482-w