Abstract
Accurate testing to detect SARS-CoV-2 RNA is key to counteract the virus spread. Nonetheless, the number of diagnostic laboratories able to perform qPCR tests is limited, particularly in developing countries. We describe the use of a virus-inactivating, denaturing solution (DS) to decrease virus infectivity in clinical specimens without affecting RNA integrity. Swab samples were collected from infected patients and from laboratory personnel using a commercially available viral transport solution and the in-house DS. Samples were tested by RT-qPCR, and exposure to infective viruses was also accessed by ELISA. The DS used did not interfere with viral genome detection and was able to maintain RNA integrity for up to 16 days at room temperature. Furthermore, virus loaded onto DS were inactivated, as attested by attempts to grow SARS-CoV-2 in cell monolayers after DS desalt filtration to remove toxic residues. The DS described here provides a strategy to maintain diagnostic accuracy and protects diagnostic laboratory personnel from accidental infection, as it has helped to protect our lab crew.
Similar content being viewed by others
Introduction
In December 2019, Chinese officials reported cases of respiratory syndrome followed by pneumonia of unknown origin, initially in the city of Wuhan, capital of the Hubei province. The etiological agent behind the upsurge of the new syndrome was identified as a new coronavirus, latter on named SARS-CoV-2 (family Coronaviridae; genus Betacoronavirus, subgenus Sarbecovirus) [1]. The virus spread rapidly throughout China and in less than a month reached other countries in Asia, eventually reaching other continents. On March 11, 2020, a global pandemic was declared by the WHO [2, 3]. To date, more than 88 million cases and almost two million deaths due to the disease caused by SARS-CoV-2, named COVID-19, have been recorded worldwide [4].
While health officials and governments around the world struggle to devise strategies to counteract the pace of the infection’s spread, efforts to implement fast and sensitive approaches for diagnostic have emerged as key steps to control the epidemics as well as part of reopening strategies. Real-time reverse transcriptase-PCR (RT-PCR) testing to detect SARS-CoV-2 RNA on samples collected from the largest possible fraction of the populations became an absolute consensus [5]. Widespread PCR testing has been pointed out as one of the most important elements in the successful COVID-19 containment strategy adopted by countries that have shown positive outcomes, including Taiwan, South Korea, and Germany [6,7,8]. Nonetheless, having test kits available is not the only bottleneck to implement universal testing. The capability to adapt the routines of diagnostic laboratories to cope with manipulation of highly infective clinical samples coming by the thousands is equally essential, especially considering that the RT-PCR diagnostic requires trained laboratory personnel.
After peaking in Asiatic and European countries, the spread of the disease veered towards the Americas and possibly the sub-Saharan African continent. Developing countries in these continents are being hit hard by the pandemic for a number of reasons. One particular troublesome aspect is the limited availability of diagnostic laboratories able to cope with the huge incoming of clinical samples—in terms of either the total number of available laboratories or their capability to deal safely with potentially infective clinical specimens [5]. Therefore, the development of strategies to reduce the infectivity of clinical samples being sent to diagnostic laboratories could be essential to avoid contamination of the limited number of trained personnel and to maintain the operational capability of these laboratories.
The high risk associated with biological samples determines that any clinical samples are to be considered potentially infectious and, therefore, must be treated under strict biosafety protocols. In this regard, national and international guidelines on biosafety must be followed in all circumstances. In the context of the COVID-19 pandemic, there is still limited information regarding laboratory-acquired infections by SARS-CoV-2 affecting health workers involved in diagnostics. The WHO recommends that handling of clinical samples suspected of being infected with SARS-CoV-2 requires a BSL-2 or equivalent facility, whereas attempts to replicate the virus require BSL-3 facilities [9].
Several chemical and physical methods of viral inactivation have been evaluated for different pathogens, as a way to provide greater safety for professionals involved in the handling of potentially infectious samples and lower costs with laboratory infrastructure [10,11,12,13,14,15,16]. Among the chemical methods evaluated, the most commonly used products contain a chaotropic salt (guanidine), which acts as a surfactant/denaturant agent culminating in virus inactivation [12]. At the same time, guanidine is able to decrease the degradation of RNA molecules in samples, acting as a ribonuclease inhibitor, and therefore increasing the preservation of genetic material for application in molecular diagnostic methodologies in which RNA integrity is essential [13].
Here, we describe the use of a simple, virus-inactivating and denaturing solution as part of a swab collection kit, aiming to decrease the infectious potential of clinical samples and, at the same time, to preserve highly labile RNA molecules during transportation and storage before testing. This low-cost, accessible approach has made it possible to achieve high diagnostic accuracy as well as manipulation safety for those involved with diagnostic procedures.
Materials and methods
Denaturing solution
The guanidine isothiocyanate solution used in this study is based on protocols established by Zolfaghari et al. [17] and Chomczynski and Sacchi [18]. The denaturing solution was prepared using 4 M guanidine isothiocyanate, 2.5 M sodium citrate, and RNAse/DNAse free water and pH adjusted to 4.0. This solution has been used for the conditioning and transportation of clinical specimens that include oropharyngeal and nasopharyngeal swabs. The guanidine chaotropic salt was chosen as the denaturing and surfactant agent, assuming its ability to inactivate viruses [12]. In addition, guanidine is also a ribonuclease inhibitor, which allows the conservation of genetic material for downstream applications using molecular biology methodologies [19].
Molecular diagnostic
Sampling
When the SARS-CoV-2 was declared pandemic, in early March 2020, research groups from major Universities and research institutes throughout Brazil began to prepare themselves to offer diagnostic support, correctly foreseeing the collapse of public and private laboratories’ capabilities shortly after the epidemics reached the country. At the UFMG’s Vaccine Technology Center, we were particularly concerned about the impacts of the intense flow of infective samples in a research laboratory that was adapted to join the testing effort with limited resources and personnel. In order to increase personnel safety, to avoid losing collaborators due to infections by SARS-CoV-2, and at the same time to increase preservation of the RNA contained in clinical samples, we introduced the use of the guanidine-containing solution as collection and transport media instead of commonly used viral transport media (VTM). VTM is usually composed of a balanced salt buffer; sterile, heat-inactivated fetal bovine serum; and antibiotics, as suggested by the CDC [20]. As recommended in the WHO interim guidance protocol, combined oropharyngeal and nasopharyngeal swabs were collected, using sterile flexible-rod swabs, and placed in a single sterile 15 mL polypropylene tube, containing 1.0 mL of the described denaturing solution or VTM (whenever applicable). These collection kits were prepared in our laboratory and sent to hospitals according to their daily demand. Upon sample collection, the swabs remained immersed in the denaturing solution for at least 30 s, after which they were removed while being gently pressed against the tube wall to remove the excess absorbed solution. Swabs were discarded in an appropriated biological waste disposal and were not sent to the diagnostic laboratory in order to minimize risks of personnel contamination. Clinical specimens from the laboratory personnel were collected multiple times and processed the same way as specimens from patients in hospitals (see below).
RNA extraction and real-time polymerase chain reaction
Extraction of the total RNA from samples was performed using the QIAamp Viral RNA Mini Kit (Qiagen, USA), according to protocols provided by the manufacturer. From the 1.0 mL of DS on the collection tube, 140 μL were used for RNA extraction and the remaining volume was frozen at −80 °C for eventual repetitions or further testing. The real-time polymerase chain reaction (RT-PCR) was performed using primers and probes described in two different protocols: the United States Center for Disease Control and Prevention (CDC) protocol [20] and the Berlin (Charité/Berlin) protocol [21]. The viral gene coding for the N protein was targeted for the detection of SARS-CoV-2 RNA using the CDC protocol, and the viral gene E was used in the Charité/Berlin protocol. Probes and primers for the human RNAse P mRNA were used in both protocols as an endogenous reaction control. Reactions were carried out with the Promega GoTaq® Probe 1-Step RT-qPCR Kit (Promega, France) according to the manufacturer’s recommendations, using the QuantStudio™ 5 real-time PCR system (Thermo Fisher, USA).
RNA integrity test
In order to verify the integrity of the RNA stored in the denaturing solution (DS) proposed in this work, in comparison to the commonly used VTM, oropharyngeal and nasopharyngeal swab samples were collected from four laboratory’s staff members and stored in VTM, following the protocol established by the WHO. Similarly, swab samples from the same staff were placed in the DS. After a 2-h period, RNA extraction was performed from all samples, as described above, and the RT-PCR protocol was performed to detect the RNAse P gene. To evaluate for how long the DS could maintain viral RNA preserved to be detected through RT-PCR, different DS tubes were spiked with 16,400 copies of SARS-CoV-2 genomic RNA and maintained for up to 16 days at either 4 °C or room temperature. After that, the viral RNA was extracted and evaluated by RT-PCR. Results obtained were analyzed in the QuantStudio™ Design and Analysis software (v.1.5.1), and graphs were generated using the GraphPad Prism software (v.8.4.2).
Detection of viral RNA from samples stored in the viral transport medium and the denaturing solution
To evaluate the preservation of the viral RNA in VTM versus DS, clinical samples were collected from hospitalized COVID-19 suspected patients using either DS or VTM. Sample collection was carried out according to the WHO protocol and the molecular diagnosis was processed as described above. Obtained results were analyzed in the QuantStudio™ Design and Analysis software (v.1.5.1), and graphs were generated using the GraphPad Prism software (v.8.0.2).
Inactivation of SARS-CoV-2 by DS
107 TCID50 of SARS-CoV-2 (stock at 107 TCID50/100 μL) were added to 1 mL of either DS or VTM and incubated for 1 h or 12 h at 4 °C. After incubation, samples were desalted using Amicon Ultra-4 centrifugal filter units (cutoff = 50,000 daltons) (Merck-Millipore, Germany) as previously described and according to manufacturer’s protocols [22, 23]. Retained virus particles were recovered in DMEM cell media with no fetal calf serum. Recovered viruses were serially diluted and inoculated onto Vero ccl-81 cells in 96-well plates for TCID50 titration. Experiments were conducted in triplicates. Vero cells in 6-well plates were also inoculated with retained virus particles from desalted columns and incubated for 5 days; extracts from these cells were reinoculated in blind passages through new cell monolayers for two times, and titrated as described above. Finally, cell monolayers were inoculated with desalted DS in the absence of viruses in order to check for any cellular toxicity. Cell viability was evaluated by trypan blue staining 48 h post-inoculum. Viability is expressed in TCID50/100 μL. All experiments with live viruses were conducted in a BSL-3-level facility at the Virology Research Center, Ribeirão Preto Medical School, USP, São Paulo.
Assessment of laboratory personnel safety
Having established that collection of clinical samples in DS preserves viral RNA in levels comparable to VTM and is able to inactivate the virus, we opted to routinely receive and process only DS collected clinical samples, as described above. In order to assess the safety of our laboratory staff using such routine, we tested all laboratory personnel every 20 days. RT-PCR tests were conducted as described above. Additionally, serum samples from all laboratory members were also collected approximately 45 days after the beginning of the study, to assess the presence of antibodies against SARS-CoV-2. To that end, we employed an in-house anti-IgG COVID-19 enzyme-linked immunosorbent assay. Briefly, the nucleocapsid (N) protein of SARS-CoV-2 was expressed in transformed E. coli, purified, and used to coat 96-well ELISA plates. Tested sera were diluted in PBS-Tween20 solution, added to wells, and incubated for 30 min at 37 °C. After washing, each well was added with horseradish peroxidase (HRP)–conjugated anti-human IgG goat immunoglobulin (Fapon, China). After further washing and incubation, reactions were revealed using tetramethylbenzidine (TMB), and readings were obtained in a microplate reader at an optical density (OD) of 450 nm. The in-house indirect ELISA was validated using a panel of SARS-CoV-2-positive sera previously tested by commercial anti-SARS-CoV-2 IgG lateral flow immunochromatography (various commercial brands).
Statistical analysis
Whenever applicable, data were compared using a two-way analysis of variance (ANOVA) implemented on the GraphPad Prism Software 8.0.2 (San Diego, USA).
Results
RNA integrity tests
Oropharyngeal and nasopharyngeal samples from four different laboratory members were collected in VTM or DS and processed. Integrity of the extracted RNA was analyzed by RT-PCR, looking for the detection of the RNAse P human mRNA. We observed no differences in the cycle threshold (CT) obtained for the detection of the targeted mRNA, regardless of the collection media (samples A to D), suggesting that the preservation of RNA in the two solutions is similar (Fig. 1).
Having established that DS can be reliably used to collect genetic material, we next asked for how long DS would keep viral genomic RNA preserved for detection, either at 4 °C or at room temperature. When compared to day 0 after spiking SARS-CoV-2 RNA into DS tubes, there were no differences in viral RNA detection in days 1, 2, 4, 8, and 16, stored under refrigeration or at room temperature (Fig. 2).
RNA viability in DS solution after storage at 4 °C and room temperature for 16 days. DS tubes were spiked with SARS-CoV-2 RNA and stored under refrigeration or at room temperature. The presence of SARS-CoV-2 RNA was detected by RT-PCR (N1 and N2 viral gene probes) in the samples after 1, 2, 4, 8, and 16 days of storage
Detection of viral RNA from samples stored in the viral transport medium and the denaturing solution
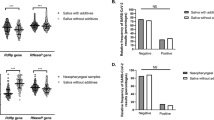
We next evaluated the detection of SARS-CoV-2 viral RNA from clinical samples collected in VTM or DS. We observed no difference in the amplification profile (determined by the cycle threshold) of the N1, N2, and N3 viral genes or of the RNAse P mRNA (endogenous control), regardless of the employed collection media (Fig. 3). Results depicted in Fig. 3 show the mean CT from six randomly selected SARS-CoV-2-positive patients compared to six randomly selected SARS-CoV-2-negative individuals. These results are representative of a much larger panel of results obtained so far. All tests were also conducted using the Charité/Berlin RT-qPCR protocol, and results were similar (not shown).
Comparative analysis of viral genes and RNAse P mRNA amplification in clinical samples extracted from VTM or DS. Mean CT of positive and negative samples for N1, N2, and N3 viral genes, extracted from VTM and DS (n = 6). VTM+: positive samples with viral transport media, DS+: positive samples with denaturing solution, VTM−: negative samples with viral transport media, DS−: negative samples with denaturing solution. Data were compared using a two-way analysis of variance (ANOVA). **P < 0.05 when comparing the groups VTM− and DS−
SARS-CoV-2 inactivation in denaturing solution
The use of the proposed DS has two main objectives: first, to ensure RNA preservation so that diagnostic qPCR can be performed unequivocally; and second, to eliminate virus infectivity so that laboratory personnel are not exposed to infectious clinical samples during processing of samples. To look into virus inactivation by DS, we conducted virus multiplication experiments after loading viable viruses in either DS or VTM and incubating for 1 h or 12 h at 4 °C. Virus-loaded transport media were filtered through Amicon columns, and the retained virus particles were eluted, serially diluted, and inoculated in fresh Vero cell monolayers. As presented in Table 1, the virus loaded in DS is inactivated after incubation for as little as 1 h, whereas virus loaded in VTM retains viability regardless of the incubation period. Viruses loaded in DS were not detected even after two rounds of blind passages in vero cells. Cells remained 96% viable after 48 h incubation with desalted DS in the absence of virus, as attested by trypan blue staining, suggesting that the lack of viral replication after viruses were loaded in DS is not due to cell toxicity.
Assessment of laboratory personnel safety
All 19 laboratory personnel working at the UFMG’s Vaccine Technology Center were evaluated during the current COVID-19 epidemic in Brazil. Laboratory members were tested at least 4 times (with one exception), in 20-day interval, and also by an in-house anti-IgG COVID-19 ELISA. Notably, none of the laboratory members tested positive in any of the tests (Table 2). PCR tests were also conducted using the Charité/Berlin protocol, and results were similar (not shown). These findings indicate that there was no SARS-CoV-2 contamination of any of the professionals involved in the diagnostic process.
Discussion
As the COVID-19 pandemic keep on spreading and pressing health policies all over the world, the potential for laboratory-acquired SARS-CoV-2 infection from accidental exposure during transportation of clinical samples and laboratory diagnosis cannot be overstated. Indeed, there has been criticism towards the CDC recommendation to collect and transport clinical specimens in VTM that preserves virus’ infectivity [24]. Therefore, inactivation of potentially infectious clinical samples is a critically important process during all stages of sample manipulation, from the initial sample collection, to processing and the final discard of the residual material [12]. Previous studies with other coronaviruses have established that treatment with heat, ultraviolet light, inactivating chemicals, and a variety of detergents are effective in inactivating beta-coronavirus [11, 25,26,27,28] Leclercq et al. [29] showed that heat treatment effectively inactivates MERS-CoV, and Kumar et al. [13] demonstrated that the use of solutions containing guanidine isothiocyanate is capable of inactivating a strain of the MERS-CoV, closely related to SARS-CoV and SARS-CoV-2. At the same time, the use of denaturing agents such as guanidine isothiocyanate provides stability to the genetic material and has been largely used in protocols designed to purify nucleic acids.
In the present study, we have replaced the virus transport media (VTM) which is commonly used in diagnostic protocols [30] by a denaturing solution (DS) containing guanidine isothiocyanate, to improve conditions of transportation and storage of clinical samples suspected of containing SARS-CoV-2. Our data demonstrated that the use of the denaturing solution in the pre-analytical process does not interfere with the detection of the presence of viral genes or the endogenous human mRNA (RNAse P), and results obtaining from samples collected in VTM or DS were identical in terms of diagnostic accuracy. Of particular importance is the fact that the DS is able to keep RNA samples preserved and able to be detected by qPCR for up to 16 days, either at 4 °C or at room temperature. This is an essential feature considering the logistics to take collected samples to diagnostic laboratories in countries with limited numbers of RT-PCR-capacitated facilities. Equally important is the fact that viable SARS-CoV-2 is inactivated upon loading on DS, as demonstrated by virus multiplication attempts in cell culture after live SARS-CoV-2 was kept in DS. Contrarily, viruses loaded in traditional VTM were readily able to replicate in cell monolayers.
The use of VTM is particularly indicated when virus viability is important, especially when SARS-CoV-2 isolation is to be attempted. However, only BSL-3 laboratories should be used to perform experiments involving replicative viruses, whereas diagnostic procedures that do not involve virus replication can be conducted in BSL-2 laboratories [10]. Media containing live viruses undoubtedly bring risks to laboratory personnel that manipulate clinical samples under lower biosafety standards [31], and accidental exposure to SARS-CoV-2 in medical and laboratory personnel has been reported [32, 33]. Indeed, in a provocative letter, Dewar and others have stated that the requirement to use virus-preserving VTM imposes unnecessary risks on transportation and laboratory professionals without any epidemiologic benefits [24]. This is especially true when we consider that most of the approved COVID-19 diagnostic tests used all over the world are nucleic acid–based molecular tests in which preservation of SARS-CoV-2 viability is not necessary. Therefore, the use of collection and transport media able to inactivate SARS-CoV-2 with no loss of the diagnostic analytical power can be critical to avoid accidental infections among diagnostic laboratories’ personnel and transport crews.
One final aspect that may be relevant to professionals involved with SARS-CoV-2 diagnostics, is that we performed multiple evaluation of our laboratory personnel and detected no accidental exposure to SARS-CoV-2 during more than 3 months working with an average of 80 samples/day. During this period, more than 8000 tests were conducted, including samples from more than 4100 patients and a SARS-CoV-2 positivity rate of approximately 25%. These data are obviously circumstantial, and cannot be used as an indication of virus inactivation when the denaturing solution is employed as collection and transport media. Moreover, we are not able to quantify to which extent the good laboratory practices adopted here could also be responsible for the absence of accidental SARS-CoV-2 exposition in our laboratory personnel. Nonetheless, the use of the denaturing transport media is a critical part of such practices, and the fact that none of our laboratory members was infected—or even seroconverted—is an important indication that DS has been helpful in avoiding accidental exposition to the virus.
References
Gorbalenya AE, Baker SC, Baric RS et al (2020) The species Severe acute respiratory syndrome-related coronavirus: classifying 2019-nCoV and naming it SARS-CoV-2. Nat Microbiol 5:536–544. https://doi.org/10.1038/s41564-020-0695-z
Wu Z, McGoogan JM (2020) Characteristics of and important lessons from the Coronavirus Disease 2019 (COVID-19) outbreak in China: summary of a report of 72,314 cases from the Chinese Center for Disease Control and Prevention. JAMA 323:1239–1242. https://doi.org/10.1001/jama.2020.2648
Coronavirus (COVID-19) - Events as they happen. World Health Organization, Geneva, 2020. Available online: https://www.who.int/emergencies/diseases/novel-coronavirus-2019/events-as-they-happen (Accessed 14 April 2020)
Worldometers. Confirmed cases and deaths by country, territory, or conveyance. Available online: https://www.worldometers.info/coronavirus/#countries (accessed on 07 January, 2021)
Peto J (2020) Covid-19 mass testing facilities could end the epidemic rapidly. BMJ 368:1163. https://doi.org/10.1136/bmj.m1163
Wang CJ, Ng CY, Brook RH (2020) Response to COVID-19 in Taiwan: big data analytics, new technology, and proactive testing. JAMA 323:1341–1342. https://doi.org/10.1001/jama.2020.3151
Sung H, Yoo CK, Han MG, Lee SW, Lee H, Chun S, Lee WG, Min WK (2020) Preparedness and rapid implementation of external quality assessment helped quickly increase COVID-19 testing capacity in the Republic of Korea. Clin Chem 66:979–981. https://doi.org/10.1093/clinchem/hvaa097
Thomson GA (2020) COVID-19: Leaving lockdown-of Schrodinger, cats, testing and masks. Int J Clin Pract 74:e13519. https://doi.org/10.1111/ijcp.13519
World Health Organization et al. (2020b) Laboratory testing for coronavirus disease (COVID-19) in suspected human cases: interim guidance, 19 March 2020. World Health Organization
Sagripanti JL, Hülseweh B, Grote G, Voss L, Böhling K, Marschall HJ (2011) Microbial inactivation for safe and rapid diagnostics of infectious samples. Appl Environ Microbiol 77:7289–7295. https://doi.org/10.1128/AEM.05553-11
Darnell ME, Subbarao K, Feinstone SM, Taylor DR (2004) Inactivation of the coronavirus that induces severe acute respiratory syndrome, SARS-CoV. J Virol Methods 121:85–91
Blow JA, Dohm DJ, Negley DL, Mores CN (2004) Virus inactivation by nucleic acid extraction reagents. J Virol Methods 119:195–198. https://doi.org/10.1016/j.jviromet.2004.03.015
Kumar M, Mazur S, Ork BL, Postnikova E, Hensley LE, Jahrling PB, Johnson R, Holbrook MR (2015) Inactivation and safety testing of Middle East Respiratory Syndrome Coronavirus. J Virol Methods 223:13–18. https://doi.org/10.1016/j.jviromet.2015.07.002
Hartman AL, Cole KS, Homer LC (2012) Verification of inactivation methods for removal of biological materials from a biosafety level 3 select agent facility. Appl Biosaf 17:70–75
Wigginton KR, Kohn T (2012) Virus disinfection mechanisms: the role of virus composition, structure, and function. Curr Opin Virol 2:84–89. https://doi.org/10.1016/j.coviro.2011.11.003
Wood BA, Mioulet V, Henry E, Gray A, Azhar M, Thapa B, Diederich S, Hoffmann B, Beer M, King DP, Eschbaumer M (2020) Inactivation of foot-and-mouth disease virus A/IRN/8/2015 with commercially available lysis buffers. J Virol Methods 278:113835. https://doi.org/10.1016/j.jviromet.2020.113835
Zolfaghari R, Chen X, Fisher EA (1993) Simple method for extracting RNA from cultured cells and tissue with guanidine salts. Clin Chem 39:1408–1411
Chomczynski P, Sacchi N (2006) The single-step method of RNA isolation by acid guanidinium thiocyanate-phenol-chloroform extraction: twenty-something years on. Nat Protoc 1:581–585. https://doi.org/10.1038/nprot.2006.83
Chomczynski P, Sacchi N (1987) Single-step method of RNA isolation by acid guanidinium thiocyanate-phenol-chloroform extraction. Anal Biochem 162:156–159. https://doi.org/10.1006/abio.1987.9999
Research use only 2019-novel Coronavirus (2019-nCoV) real-time RT-PCR primer and probe information. CDC - Centers for disease control and prevention, Atlanta. 2020. Available online: https://www.cdc.gov/coronavirus/2019-ncov/lab/rt-pcr-panel-primer-probes.html (Accessed 15 May 2020)
Corman VM, Landt O, Kaiser M, Molenkamp R, Meijer A, Chu DKW, Bleicker T, Brünink S, Schneider J, Schmidt ML, Mulders DGJC, Haagmans BL, van der Veer B, van den Brink S, Wijsman L, Goderski G, Romette JL, Ellis J, Zambon M, Peiris M, Goossens H, Reusken C, Koopmans MPG, Drosten C (2020) Detection of 2019 novel coronavirus (2019-nCoV) by real-time RT-PCR. Euro Surveill 25:2000045. https://doi.org/10.2807/1560-7917.ES.2020.25.3.2000045
Burton JE, Easterbrook L, Pitman J, Anderson D, Roddy S, Bailey D, Vipond R, Bruce CB, Roberts AD (2017) The effect of a non-denaturing detergent and a guanidinium-based inactivation agent on the viability of Ebola virus in mock clinical serum samples. J Virol Methods 250:34–40. https://doi.org/10.1016/j.jviromet.2017.09.020
Pastorino B, Touret F, Gilles M, Luciani L, de Lamballerie X, Charrel RN (2020) Evaluation of chemical protocols for inactivating SARS-CoV-2 infectious samples. Viruses 12:624. https://doi.org/10.3390/v12060624
Dewar R, Baunoch D, Wojno K, Parkash V, Khosravi-Far R (2020) Viral transportation in covid-19 pandemic: inactivated virus transportation should be implemented for safe transportation and handling at diagnostics laboratories. Arch Pathol Lab Med 10:5858–5917. https://doi.org/10.5858/arpa.2020-0175-LE
Darnell ME, Taylor DR (2006) Evaluation of inactivation methods for severe acute respiratory syndrome coronavirus in noncellular blood products. Transfusion 46:1770–1777. https://doi.org/10.1111/j.1537-2995.2006.00976.x
Kariwa H, Fujii N, Takashima I (2006) Inactivation of SARS coronavirus by means of povidone-iodine, physical conditions and chemical reagents. Dermatology 212:119–123. https://doi.org/10.1159/000089211
Rabenau HF, Biesert L, Schmidt T, Bauer G, Cinatl J, Doerr HW (2005) SARS-coronavirus (SARS-CoV) and the safety of a solvent/detergent (S/D) treated immunoglobulin preparation. Biologicals 33:95–99. https://doi.org/10.1016/j.biologicals.2005.01.003
Kochel TJ, Kocher GA, Ksiazek TG, Burans JP (2017) Evaluation of TRIzol LS inactivation of viruses. Appl Biosaf 22:52–55. https://doi.org/10.1177/1535676017713739
Leclercq I, Batéjat C, Burguière AM, Manuguerra JC (2014) Heat inactivation of the Middle East respiratory syndrome coronavirus. Influenza Other Respir Viruses 8:585–586. https://doi.org/10.1111/irv.12261
Lab Alert: CDC posts new standard operating procedure for creating viral transport media. CDC - Centers for disease control and prevention (CDC), Atlanta. 2020. Available online: https://www.cdc.gov/csels/dls/locs/2020/new_sop_for_creating_vtm.html (Accessed 15 May 2020)
Orellana C (2004) Laboratory-acquired SARS raises worries on biosafety. Lancet Infect Dis 4:64. https://doi.org/10.1016/s1473-3099(04)00911-9
Wang D, Hu B, Hu C, Zhu F, Liu X, Zhang J, Wang B, Xiang H, Cheng Z, Xiong Y, Zhao Y, Li Y, Wang X, Peng Z (2020) Clinical characteristics of 138 hospitalized patients with 2019 novel coronavirus-infected pneumonia in Wuhan, China. JAMA 323:1061–1069. https://doi.org/10.1001/jama.2020.1585
Ong SWX, Tan YK, Chia PY, Lee TH, Ng OT, Wong MSY, Marimuthu K (2020) Air, surface environmental, and personal protective equipment contamination by Severe Acute Respiratory Syndrome Coronavirus 2 (SARS-CoV-2) from a symptomatic patient. JAMA 323:1610–1612. https://doi.org/10.1001/jama.2020.3227
Acknowledgements
We thank all personnel at UFMG’s Vaccine Technology Center (CTVacinas) for their dedication to the COVID-19 diagnostic support in Belo Horizonte, Brazil. Members of the CTVacinas are composed of undergraduate and graduate students, post-doc fellows, professors, and researchers at the Federal University of Minas Gerais and the René Rachou Institute (FIOCRUZ), who interrupted their research activities to dedicate themselves to aid people during such difficult times. We thank Anna Raquel dos Santos, Júlia Reis, Lisiane Gomes, Natalia Salazar, and Ricardo Gazzinelli for their help and expertise. We thank Professor Édison L. Durigon, at Laboratório de Virologia Clínica e Molecular, ICB/USP, São Paulo, Brazil, who kindly provided inactivated samples of the virus isolate HIAE-02: SARS-CoV-2/SP02/human/2020/BRA (GenBank accession number MT126808.1). We also thank the Laboratório de Virus’ team, at ICB/UFMG, for valuable technical help.
Availability of data and material
Not applicable.
Code availability
Not applicable.
Funding
The research leading to these results has, in part, received funds from CAPES, CNPq, FAPEMIG, and FIOCRUZ, and from the MCTIC through the “Rede Virus” initiative. Funds came also from UK Research and Innovation via the Global Challenges Research Fund under grant agreement “A Global Network for Neglected Tropical Diseases” (grant number MR/P027989/1). Further funding included institutional financial resources of the René Rachou Institute - Oswaldo Cruz Foundation (FIOCRUZ) and the Instituto Nacional de Ciência e Tecnologia de Vacinas (INCTV).
Author information
Authors and Affiliations
Contributions
Conceptualization, AFC, PAA, SMRT, and FGF; methodology AFC, SMRT, PAA, and FGF; validation AFC, APG, TBSS, HIS, LV, FFB, SARS, MMF, RPR; formal analysis, FGF and SMRT; data curation AFC, HIS, FFB, RPR, PAA; writing—original draft preparation, FGF, PAA, APG, and TBSS; writing—review and editing, FGF, FFB, and RPR; supervision, FGF and SMRT; project administration, FGF, SMRT, PAA, and APSMF; funding acquisition FGF; SMRT and APSMF. All authors have read and agreed to the published version of the manuscript.
Corresponding author
Ethics declarations
Ethics approval
This work was approved by the UFMG’s Ethics Committee and by the National Research Ethics Committee, under number CAAE 31686320.0.000.5149.
Consent to participate
All laboratory personnel signed consents to participate in this study.
Consent for publication
Not applicable.
Conflict of interest
The authors declare no conflict of interest. The funders had no role in the design of the study; in the collection, analyses, or interpretation of data; in the writing of the manuscript, or in the decision to publish the results.
Additional information
Responsible Editor: Giliane Souza Trindade
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Carvalho, A.F., Rocha, R.P., Gonçalves, A.P. et al. The use of denaturing solution as collection and transport media to improve SARS-CoV-2 RNA detection and reduce infection of laboratory personnel. Braz J Microbiol 52, 531–539 (2021). https://doi.org/10.1007/s42770-021-00469-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s42770-021-00469-4