Abstract
Digital Microfluidic Biochips (DMFBs) will require error-free synthesis techniques which can function at much higher speed while implementing on real-time systems and capable of tackling more complex assay operations. Until now various bio-assays are successfully implemented based on different mixing modules present on such lab-on-chips. In present work, the concept of such dedicated virtual modules has been eliminated and a novel module-less-synthesis (MLS) method is proposed for accomplishing high-performance bio-protocols. Various shift-patterns (movements) of the micro-droplets are identified to accomplish entire mixing in lesser time compared to earlier module-based synthesis methods. We have also computed the percentage of mixing accomplishment for each directional-shift of the mixer-droplet. However, path congestion problem and operational errors are inevitable in MLS approach. Hence, the path congestion and washing problem in MLS is addressed by tweaking the earlier MLS approach and a new modified-MLS (MMLS) method is proposed. Finally, washing optimization technique on MMLS method is also given. Different real-life bio assays like PCR, IVD are tested with the proposed technique as well as synthetic benchmarks (hard test benches) are also incorporated in the experiments. For both kind of benchmarks synthesis performance improved with bioassay completion time (\(T_{\mathrm{max}}\)) significantly reduced compared to existing synthesis approaches on DMFB platform.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
Digital microfluidics is the new paradigm used for monitoring and prognostic applications in the field of medical, pharmaceutical and environmental sciences [4, 27]. It can manipulate bio-fluids as discrete volume (\(10^{-9}\) or \(10^{-12}\) L) of droplets on a 2-dimensional array of electrodes [24]. Initially microfluidic biochips [27] were consists of micro-pumps and micro-valves, and continuous liquid flows through fabricated micro-channels present on such chips. Presently droplet-based digital microfluidics technology [24] are merged with software-controlled, physical-aware (cyberphysical) DMFB’s which can be extensively used for point-of-care (PoC) tests and treatments especially in developing countries where adequate laboratory facilities are missing [16].
Various in vitro procedures, such as immunoassay, real-time DNA sequencing, polymerase chain reaction (PCR), protein crystallization and other complex assays have been successfully demonstrated till now on DMFB [3]. Researchers addressed these procedures with architectural-level and physical-level synthesis [18] and optimization of synthesis completion time has been targeted. Several solutions were given in this regard which may be applied at different phases of the synthesis steps. In [30], a new routing method for cross-referencing-based DMFBs was introduced where droplet transportation problem has been target using graph clique property and latest arrival times for droplet routes are minimized. Integer linear programming (ILP) was used in [31] to explore the application of routing technique in cross-referencing DMFB. The major concern in this approach was the electrode interference which actually hindered the simultaneous movement of multiple droplets. Next, another ILP-based optimization method is used to solve the droplet routing and the pin-mapping design problems concurrently [7]. Also a contamination-aware synthesis is proposed in [11], where intra- and inter-contaminations are cleaned simultaneously within a subproblem to reduce the execution time.
In all of the above discussed cases as well as in other cases also module-based synthesis techniques are used predominantly on DMFB platform [15, 17, 18, 28] till date. This, in turn, increases the completion time of the bio-assays, because a limited number of ‘mixing-modules’ are available at any point of time on the chip [4]. Moreover, module-based synthesis methodologies decrease the chip utilization factor [4] as huge numbers of guard cells are occupied during the module operations to pad the mixer droplets from cross-contamination with other existing droplets on the chip. Very few works has been presented omitting the concept of dedicated modules on DMFB’s till date [1, 4, 19, 29]. Maftei et al. [19] first proposed the concept of routing-based synthesis where mixing time was improved by avoiding flow reversibility and eliminating static pivot points present in different mixing modules. However, no appropriate computation [19] of mixing completion time was addressed for consecutive linear movements of the mixer-droplet. Thus, it cannot be suitably applied on cyberphysical DMF chips. In addition, a simplistic assumption of straight flow of the mixer droplet is assumed [19] for faster mixing completion but no discussion about the limitation of linear flow is given.
In [29], congestion avoidance and washing mechanisms are mentioned for routing-based synthesis technique. The chip operating frequency assumed to be 100 Hz, which produces a switching time of 10 ms but no references are available where mixing operations can be performed in real-life chips at such an high frequency. The mixing time computation for various shifts of the mixer droplet was empirically deduced based on 16 Hz chip from [22], which are contradictory with the chip operating frequency. Such an assumption is inappropriate to take as most of the applications on DMF chips involve fluidic operations on viscous analyte or medium, making it very difficult to apply such 100 Hz fast switching. In addition, the mentioned techniques in [29] cannot be applied to cyberphysical chips where real-time error detection and error control are two important aspects and need to be addressed separately.
Also in all the earlier works on module less synthesis for DMFB [4, 19, 29], the concept of negative mixing completion for \(180^{\circ}\) (read as ‘one-hundred eighty degree’) shift-movements of the mixer droplet was empirically deduced. Such concept of negative mixing / reverse-mixing of micro-droplets is simply incorrect and was a result of erroneous mathematical model. In 2014, it was experimentally proved at the Microfluidics Lab, Duke University, USA that even a ‘2-electrode mixing’ can also be accomplished on a microfluidic chip [9, 10]. A mixing module of size \(1\times 2\) is used at 1 Hz switching speed and the mixing operation can be successfully completed with a series of forward and backward movements of the mixer-droplets one after another [10]. Such permissible movements on a \(1\times 2\) mixing unit can be mapped with a \(0^{\circ}\)-shift at first and then all the \(180^{\circ}\)-shifts from second shift onward which is shown in Fig. 6a. Hence, it is evident that the \(180^{\circ}\)-shifts (backward movement of the mixer-droplets) also contribute to the mixing for a certain amount. If it would have done the de-mix of the droplets, the mixing would never be completed on a \(1\times 2\) mixing unit. Thus, the previous concept of negative-mixing by \(180^{\circ}\)-shift on a DMF-chip can be rejected [9].
In this work, a module-less synthesis methodology on DMFB platform is presented which assures much higher chip utilization factor by removing the virtual modules on the chip as well as removing the extra electrodes needed as guard cells. The micro-droplets are allowed to move on the 2-D chip by different types of shift-movements and mixing is done through diffusion instead occupying a dedicated region on the DMF chip. The assay completion times are reduced by our proposed methodology as well as completion time uncertainties are also decreased. Thus, it satisfies cyberphysical DMFBs operational criteria of faster synthesis time and error handling capability by adopting definite paths for each of the mixing stages. However, in MLS approach path congestion complexities and security threats are inevitable due to the integration of cyberphysical paradigm [6]. Our technique efficiently handles congestion and reduces washing overhead by tweaking earlier MLS approach [4] and a new version of MLS named as modified-MLS (MMLS) is also proposed.
The organization of the paper is as follows: Sect. 2 discusses the preliminary concepts of basic construction, droplet operations and synthesis steps on a CP-DMFB, while the MLS problem formulation is shown in Sect. 3. Then, a novel module-less synthesis (MLS) approach is proposed for cyberphysical DMFB in Sect. 4. Mixing completion percentage for each directional shifts are derived in Sect. 4.1 and a new mixing architecture is presented in Sect. 4.2. The MLS synthesis algorithm as well as an application specific MLS chip architecture is presented in Sects. 4.3 and 4.4, respectively. Next, an illustrative example of PCR assay is shown on such architecture in Sect. 4.5. Congestion avoidance in MLS and wash optimization technique is also discussed in Sect. 5. Then, Sect. 6 represents the simulation results in detail for real-life assays as well as synthetic benchmarks, and finally Sect. 7 concludes the module less synthesis approach for DMFBs.
2 Preliminaries
2.1 Basic construction and operations of DMFB cell
The micro-droplets used in DMFB’s may consist of bio-medical samples like blood, serum, urine or saliva and the filler medium (usually silicone oil) are sandwiched between two parallel glass plates as shown in Fig. 1a, b.
The bottom plate consists of a series of individually controllable electrodes and a single top plate is used as ground electrode. The droplet movement is achieved by the principle of electro-wetting-on-dielectric (EWOD) [28]. The cyberphysical DMFB instruction set includes droplet transport in four directions, i.e., UP, DOWN, RIGHT and LEFT as shown in Fig. 2a. Splitting a droplet into two unit volume droplets and merging of two droplets into one is shown in Fig. 2b. Similarly, mixing, storage (incubation) and detection of the microdroplets are also inevitable operations for accomplishing any real-life assay on the cyberphysical chip [15]. Apart from that, external devices such as heaters, photo-detectors [12, 32] capacitance sensors, impedance sensors [20] are used to offer additional functionality.
2.2 Synthesis flow for DMFB
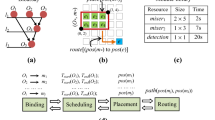
The synthesis on a CP-DMFB starts with a sequence graph (G) which is derived for a specific bioassay to be performed and it continues through several stages such as binding, scheduling, placement and routing as shown in Fig. 3.
A standardized module library is defined where various possible fluidic operations such as mixing, detection, etc., are determined along with their respective grid-size and timing requirements [23]. The modules are bind with nodes of the application graph (G) and scheduling determines the order of operations based on such binding. Finally, the placement of such modules on the chip is determined followed by routing operations.
3 Problem formulation
Design optimization of Digital Microfluidic Biochip crucially depends on the synthesis methodologies used for such chips. Until now, module-based synthesis techniques are used for cyberphysical DMFBs. A new approach is shown in [17], where smaller mixing modules are used at initial stages, merged in bigger modules, as synthesis progresses and ultimately entire chip becomes a single mixing module. However, no definite routing information is given for the intermediate stages in this approach. We have considered the module library given in [22] as a benchmark for DMFB platform, where timing requirements of different fluidic operations are provided [4]. Figure 4 illustrates different mixing modules along with their respective padding cells typically used in assay synthesis on a CP-DMFB. In addition, the mixing completion time for various modules of different sizes and the respective number of padding cells required for such modules are given in Table 1.
It is evident from Table 1, that no correlation exists between the module sizes (active mixing region) and mixing completion time. For example, a \(2\times 2\) and \(1\times 4\) module occupies same area (4 number of cells) on the chip apparently seems to take same amount of time to do the mix operations [4]. However, in actual the mixing time varies more than 100% for these two modules as shown in Table 1.
For sake of simplicity, we may assume to take the fastest mixing modules available on DMFB framework to speedup the execution time of bio-protocols. Such modules require a space of \(2\times 4\) = 8 cells and the mixing completion time is 2.9 s [4]. However, choice of such \(2\times 4\) modules actually leads to another design issue as it blocks 24 cells (electrodes) including the padding cells and utilize only the 8 electrodes which are centrally positioned during the entire mixing operation. The remaining 16 cells has to be inactive and need to work as padding cells only (Table 1) to avoid cross-contamination with other droplets as shown in Fig. 4c. Moreover, only 1 electrode is used at a particular time instant (t) where \(t = 1\) time-step and it is in the range of approximately 0.05 s \(\le t \le\) 0.125 s depending on the operating frequency (f) of the chip which generally varies between 20 and 8 Hz for a CPDMFB [4]. Thus, in module-based synthesis approach, hardware cost is very high as more than 66% module areas are unrealized during the module operations. Also, the bioassay completion time are large enough and hence not very suitable for real-time cyberphysical DMFB applications like various point-of-care (PoC) diagnostics which can be helpful for quick, error-free detection of various diseases like AIDS, malaria, dengue, and may be applicable for epidemic/pandemic diseases like ZIKA, Middle Eastern respiratory syndrome (MERS) and novel corona/COVID-19 in near future.
We have formulated the problem considering the PCR bioassay [13] and a module-based chip of size \(8\times 9\) cells shown in Fig. 5b. We have placed 4 mixing-modules on the chip to accommodate \(1{\mathrm{st}}\) layer of PCR simultaneously and omitted the \(2\times 2\) module which having the longest time requirement (Table 1). Now if \(M_{1}\), \(M_{2}\) operations are performed on \(1\times 4\) mixer-units and \(M_{3}\), \(M_{4}\) runs on \(2\times 4\) and \(2\times 3\) mixer units, respectively, then \(M_{1}\), \(M_{2}\) operations will be completed in 4.6 s and \(M_{3}\), \(M_{4}\) finishes, respectively, in 2.9 s and in 6.1 s. As \(M_{5}\) is the mixing of \(M_{1}\) and \(M_{2}\), thus \(M_{5}\) can directly start after 4.6 s, whereas \(M_{6}\) cannot be started before the completion of \(M_{4}\) (6.1 s). In next layer, if \(M_{5}\) and \(M_{6}\) run on \(2\times 4\) and \(1\times 4\) mixing modules (the fastest possible mixing units available on the chip), their completion time will be 7.5 s and 10.6 s, respectively. Thus, it is obvious that \(M_{7}\) will be completed only after 13.6 (\(\approx\) 14) s. Thus, the completion time of the PCR assay requires 14 s in module-based cyberphysical DMFB ignoring the time required for other operations like split, merge, and detection of the droplet [4]. To place the 1st layer of PCR sequence graph (G) with 4 parallel mixing operations (nodes), the optimum placement of the modules on an \(8\times 9\) (72 cells) chip size is shown in Fig. 5b, where only 9 cells are available for other operations like detection, dispensing, etc., Out of total 72 electrodes (cells). For this typical example about 88% of the chip space is occupied by the mixing modules only.
Thus, module-based chip demands bigger chip size and ultimately results in wastage of spatiotemporal resources on the chip. It also requires a huge number of activation pins, which essentially draws more power and consequently chip degradation is inevitable. In module-based mixing, we can detect the error only after the mixing process fully completed within the virtual modules. Thus, if any error occurs the entire mixing time is wasted and we need to restart the mixing from the same initial state in the same error-prone module or may be in a different module if available at all at that point of time. Thus, module-based synthesis approach involved more synthesis time and incurred more overhead cost in the event of an error.
4 Proposed synthesis method
This work primarily focuses on the synthesis of various bio assays by adopting definite path (patterns) for the mixer droplets along with traditional routing and other necessary synthesis operations on the chip. The proposed synthesis technique avoids the usage of virtual mixing modules on the DMFB chip, and a new chip architecture is derived for accomplishment of much faster synthesis. The usual operations like mixing, splitting, merging, routing and detection is generally performed by the basic steps of droplet movement through different electrode activation sequences. It satisfies the demand of minimizing assay completion time and error-free results for a cyberphysical DMFB and diffusion-based mixing or passive mixing [22] is considered in this work. We have proposed different directional shifts (movement) of the mixer-droplet and for each shift-movement; the percentage of mixing accomplishment is computed. In addition, an efficient error detection approach is also formulated.
4.1 Computation of mixing completion for various directional-shifts
This section deals with the computation of mixing completion percentage for various directional-shifts of the mixer droplet. The required mixing time of various shifts are derived based on inductive proof by taking the experimental results performed by Paik et al. [22]. In [22], authors did the laboratory experiments on various mixing modules and a module library is prepared which is given in Table 1. An optimum aspect ratio of the micro-droplet turned out to be 0.4 with 16 Hz chip operating frequency. Hence, the standard operating frequency (f) of the chip is chosen as 16 Hz in this work and the switching time (t) of moving a droplet from one electrode to any of its four neighboring electrodes is taken as:
To compute mixing completion percentage we consider both the \(1\times N\) and \(2\times N\) mixing-frameworks shown in Figs. 6 and 7, respectively, where N is a positive integer and \(2 \le N \le \infty\). Mixing operation of droplets is carried out in both the framework by following a specific pattern of shifts (directional-movement) of the mixer-droplets. From Figs. 6 and 7, it is seen the various shifts a mixer droplet can adopt are broadly categorized into three types. These are \(0^{\circ }\)-shifts (read as zero-degree shift-movements), \(90^{\circ }\)-shifts (read as ninety-degree shift-movements) and \(180^{\circ }\)-shifts (one hundred eighty degree shifts). The \(0^{\circ }\) shift-movement can further be divided into two types. \(0^{\circ }_{1}\)-shift (read as first order zero degree shift) which is \(0^{\circ }\)-shift along one cell position only as seen in \(1\times 3\), \(2\times 3\) mixing modules and \(0^{\circ }_{2}\)-shift (second order zero shift), which is a linear shift along two consecutive cells as seen in \(1\times 4\) and \(2\times 4\) modules, respectively.
From Figs. 6 and 7, it is evident that in \(1\times N\) mixing framework, mixing is performed with \(0^{\circ }\)-shift and \(180^{\circ }\)-shifts and in \(2\times N\) mixing framework \(0^{\circ }\)-shift and \(90^{\circ }\)-shift patterns are combined to accomplish mixing. Hence, linear movement of the droplet (\(0^{\circ }\)-shifts) can be possible on both mixing frameworks, whereas \(180^{\circ }\)-shifts and \(90^{\circ }\)-shifts are only specific to \(1\times N\) and \(2\times N\) mixing frameworks, respectively (Fig. 8).
To derive the mixing completion amount for different shifts in a module, we started with \(1\times 2\) mixing module where the mixing is accomplished by \(180^{\circ }\) shifts only and mixing completion time is \(\cong\) 16.95 s [22]. Hence, total numbers of time step required \(\frac{16.95}{0.0625} = \lceil 271.2 \rceil \cong\) 272 steps. We round off to the next higher value as a pessimistic assumption of mixing time requirement.
Hence, we may conclude that a total of 272 steps of \(180^{\circ }\)-shifts can accomplish 100% mixing in a \(1\times 2\) mixing module. Then, one shift of \(180^{\circ }\) accomplished = \(\frac{100}{272} \cong\) 0.37% of mixing.
Now if we consider the next bigger module in \(1\times N\) mixing framework (Fig. 6b), where mixing is done in combination of one \(0^{\circ }\)-shift followed by \(180^{\circ }\)-shift. Such shift-pattern is repeated continuously and required total mixing time is 12.5 s. Thus, in \(1\times 3\) module, a total of \(\frac{12.5}{0.0625} = 200\) steps should be needed to accomplish the mixing operation. From Fig. 6b, it is evident that the total number of shift-movements (200) is equally divided into \(0^{\circ }\)-shifts and \(180^{\circ }\)-shifts. Hence, \(180^{\circ }\)-shifts cumulatively accomplish \(100 \times 0.37 = 37\%\) of the total mixing. Remaining 63% of mixing is accomplished by the remaining 100 number of \(0^{\circ }\)-shifts. Thus, we can find mixing completion amount for a single \(0^{\circ }\)-shift designated as \(0^{\circ }_{1} \cong\) \(\frac{63.0}{100} \cong\) 0.63%.
Similarly from \(1\times 4\) module we may find the mixing completion amount by two consecutive linear shifts designated as \(0^{\circ }_{1,2} \cong\) 4.8%. Thus, the second linear shift \((0^{\circ }_{2})\) individually accomplishes the following amount of mixing. \(0^{\circ }_{2} \approx (0^{\circ }_{1,2} - 0^{\circ }_{1})\) \(\approx\) (4.8−0.63) \(\approx\) 4.16% of mixing.
From Fig. 7a–c it can be seen that in a \(2\times 2\) module only \(90^{\circ }\) shift patterns are available, whereas in a \(2\times 3\) module one \(0^{\circ }\)-shift is followed by two \(90^{\circ }\)-shifts and in a \(2\times 4\) module two \(0^{\circ }\)-shifts are followed by two \(90^{\circ }\)-shifts and so on so forth. Hence, mixing completion percentage for various linear shift movements \((0^{\circ }_{1}\) and \(0^{\circ }_{2})\) and \(90^{\circ }\)-shifts are also computed from the \(2\times N\) mixing-framework and all the results are summarized in Table 2.
From Table 2, it is now evident that \(0^{\circ }_{1}\) and \(0^{\circ }_{2}\) shifts are available in both the mixing framework we have considered but more amount of mixing accomplishment is achieved in \(2\times N\) framework compared to \(1\times N\) mixing framework. This is due to avoidance of flow reversibility of the shift patterns as well as multiple pivot points are introduced in \(2\times N\) mixing-framework, which essentially accelerates the overall mixing process and mixing accomplished in much faster time. Hence, we have taken the mixing percentage values of linear shifts \((0^{\circ }_{1}\) and \(0^{\circ }_{2})\) from \(2\times N\) framework to derive the further linear shift movements possible and propose a new mixing module architecture along with their respective mixing patterns.
4.2 Proposed mixing architecture
If we calculate the mixing completion amount for each \(0^{\circ }_{2}\)-shift, we have found it is more than its immediate predecessor of \(0^{\circ }_{1}\)-shifts. It essentially establishes the assumption of more number of one directional straight-movement (consecutive \(0^{\circ }\)-shifts) of the mixer droplet along the same coordinate axis can accomplish faster mixing than combining horizontal and vertical movement one after another. Thus, a \(1\times 4\) module can finish much faster mixing than a \(2\times 2\) module, irrespective of occupying the same area on the chip. Thus, mixing pattern can essentially reduce the mixing time a lot as well as overall synthesis time of the assays. However, if we fix the module size (\(2\times N\) where \(N_{\mathrm{max}}=4\)) as in module-based approach [22], we essentially restrict the shift-patterns a mixer-droplet can have. So using Lagranges formula, we extrapolate the timing requirements for next bigger module where \(N\ge 5\) to derive next higher consecutive liner movements of the droplet.
From the above discussed premise, we propose a newer mixing modules under \(2\times N\) mixing-framework by extending the value of N. This is with an intention to achieve more linear shifts in the mixing patterns; as achieving such shift-patterns would minimize the mixing completion time. Lagranges Interpolation formula [21] is applied to find the probable completion time of mixing (dilution) for next bigger mixing modules, where \(N\ge 5\) and \(x_{1}\), \(x_{2}\), \(x_{3},\dots , x_{n}\) denotes module length as N value increases and \(y_{1}\), \(y_{2}\), \(y_{3},\dots , y_{n}\) denotes mixing time for each module. The above scenario is modeled as follows:
From Eq. (2) it is seen f(x) is a polynomial and it’s Minima exits at \({x}= -6.5\) i.e., \(f^{\prime }(x)={x}-6.5\) and value of vertex is \(-0.125\) and accordingly the mixing completion time for \(2\times 6\) and \(2\times 7\) modules should be zero (for \({x}= 6\) and 7), which is an infeasible condition for any practical experiments. Due to less number of experimental data available for curve fitting we have fitted the curve using (Matlab R2009b) by simulating 1000 times and a more pessimistic assumption of mixing time (\(y_{0},y_{1},y_{2,\dots })\) in various modules has been considered to fit the curve properly without any loss of generality. Such repeated measurements provide a more satisfactory estimate, as we can use the spread found in the observed values to derive the characteristics of the underlying error. It is often convenient to assume that the distribution of measurements around the true value by the Gaussian distribution [2] as below:
where \(P(x){\mathrm{d}}x\) is the probability of obtaining a value between x and \((x + dx); x_{0}\) is the true value, and \(\sigma = \sqrt{(x-x_{0})^{2}}\) is the standard deviation or root-mean-square error of x. Hence, the error of a single observation is quoted as \(\pm \,\sigma\), where percentage of error becomes insignificant (\(\sigma \le 0.02\)) and we refer to this as an experimental error [2] and arrived at Eq. (4).
From above Eq. (4); it is observed the mixing time improves till \({N}=6\) and then again it starts to increase for \(N\ge 7\) which is shown in the below graph. Hence, we propose new mixer unit architectures as \(2\times N\) where \({N}= 5\) and 6 and the shift-pattern of mixing in these newly proposed modules are shown in Figs. 9 and 10, respectively, where a mixer-droplet can take up to 3 or 4 consecutive linear moves.
For higher values of \(N \ge 7\), achieving more linear movements (patterns) doesn’t improve the mixing time rather deteriorate as shown in Fig. 8. Hence, after 4 number of consecutive linear moves \((0^{\circ }_{1,4})\), the needed turbulence is introduced in form of a mandatory \(90^{\circ }\) shift in our shift pattern which improves the mixing time. The respective mixing completion percentage for \(0^{\circ }_{3}\) and \(0^{\circ }_{4}\) are also given in Table 2 in italic.
On the contrary, such linear movements cannot be continued for as many as possible in reality and thus limitation of such linear movement is given in Lemma 1.
Lemma 1
A maximum of 4 consecutive straight/linear movement of the droplet is possible in MLS method.
Proof
Using the module library statistics for DMFB as shown in Table 1, it can be seen that \(2\times 4\) is the maximum module size available in the literature [22]. We have derived the mixing completion percentages for various shift patterns given in Table 2 and extend the mixing architecture under \(2\times N\) mixing-framework. From Eq. (2) it is found for \(N \ge 5\); the mixing time \(T_{\mathrm{mix}}\) initially decreases and for \(N \ge 6\), \(T_{\mathrm{mix}}\) starts increasing again.
Hence, maximum module size possible on a DMFB under \(2\times N\) framework is 6, where \(2\le N \le 6\) as shown in Fig. 10. Now following the mix-patterns of \(2\times N\) mixing-framework, where mixing is accomplished combining \(0^{\circ }\) and \(90^{\circ }\)-shifts only, the intended maximum number of consecutive linear moves possible in a \(2\times 6\) module is 4 and after that a mandatory \(90^{\circ }\)-shift has to be taken by the droplet. No other way it is possible to get more than 4 number of linear moves in such a mixer and for bigger dimension mixers the mixing time is increasing as already proved. Hence, MLS can have a maximum of 4 number of consecutive linear movements to attain fastest possible mixing. □
4.3 Module less synthesis process
We perform the assay-synthesis using above-derived shift-movements (\(180^{\circ }\), \(90^{\circ }\), \(0^{\circ }_{1}\), \(0^{\circ }_{2}\), \(0^{\circ }_{3}\) and \(0^{\circ }_{4}\)) rather assigning a dedicated module for the mixing operations on the chip. The proposed methodology uses the above-derived shifts and finds out paths (patterns) for each mixing operation until 100% mixing (dilution) is achieved. It mainly attacks the synthesis problem at the point of mixing which are the major time consuming re-configurable operations on the chip. Apart from that, washing optimization as well as an efficient error detection mechanism is also given for proposed module less synthesis (MLS). The other non-re-configurable operations are mentioned in passing and production of waste droplets and routing them off the chip is not addressed (Fig. 11).
Figure 12 shows the pseudocode for MLS operations where the inputs to the algorithm are taken as the DAG representation of the bio protocol to execute sequence graph (G), an array representing the DMFB chip (A), the set of permissible shifts (S), and set of wash droplets (W).
The initial routing to be conducted as per traditional routing method [18] for transportation of the droplet from its reservoir to the intended starting position. The MLS mixing approach starts with the identification of all the mixing operations \((M_{j})\) in a particular layer \((L_{i}\) and initialization of the coordinates for all mixing operations at first level (\(L_{i}\) where \({i} = 1\)) of sequence graph \(G = (V, E)\). Usually, the four corner areas of the chip are chosen to start the first stage of mixing according to the sample and reagent reservoir positions.
Next, it searches for all possible shifts from the initial coordinate position. It first checks for the availability of linear-shifts (\(0^{\circ }\)-moves). If no (\(0^{\circ }\)-shift is available, \(90^{\circ }\)-shifts are searched. The parallel movement of mixing-droplets on the cyberphysical DMFB is always done by checking the routing constraints at each time-step and the percentage of mixing completion \(({\mathrm{Mix}}_{\mathrm{percentage}}\) and present coordinate positions of the mixer droplets are updated accordingly in the control software [line no. 35].
The mixer-droplets can move only in left, right, up and down directions, while maintaining routing constraints and can be modeled as disjunctive normal form (DNF) of four functions \(F_{1}, F_{2}, F_{3}\, {\mathrm{and}}\, F_{4}\) [Eq. (6) represents F1]. As each instant of time t which droplet should move in which direction out of four possibilities on a DMF chip is a decision problem and iteration of such decision problems over many instances of time actually formed the optimization problem for finding minimal (profitable shift-patterns in case of MLS) for all the mixer droplets and thus the mixing problem essentially converted into a routing problem.
•Routing protocol: To route a \(i{\mathrm{th}}\) droplet (\(S^{t}_{i ,x,y}\)) from its current cell position to the left cell at very next time stamp \(F_{1}= S^{t+1}_{i ,x-1,y}=1\) it is necessary that no other \(j{\mathrm{th}}\) droplet (\(S^{t}_{j ,x,y}\)) come to the neighbor seven cell as shown in Fig. 11, where \(j \in i\) but \(j \ne i\). The problem can be formulated using four functions in disjunctive normal form as below
Each of the function can be represented as a conjunctive normal form and for example the function \(F_{i}\) where \({i} = 1\); is shown below:
Similarly \(F_{2}\), \(F_{3}\), \(F_{4}\) will represent right, up and down movement of the droplet, respectively. Also the sum of mixing completion after 1, 2 or more number of \(0^{\circ }\)-shifts are represented as:
where i is an positive integer and \({i}=1, 2, 3,\dots ., n\) and \(P_{\mathrm{sum}}(n)\) is equal to percentage of mixing completion after \(n{\mathrm{th}}\) consecutive \(0^{\circ }\) move. Hence, putting the value of n up to 4 we can get
The above symbols signifies (read as) following:
-
\(0^{\circ }_{1}\) signifies: Individual \(1{\mathrm{st}}\) \(0^{\circ }\)-move along one cell position.
-
\(0^{\circ }_{2}\) signifies: \(2{\mathrm{nd}}\) \(0^{\circ }\)-move only and \(0^{\circ }_{3}\) signifies: \(3{\mathrm{rd}}\) \(0^{\circ }\)-move and so on. Similarly, \(0^{\circ }_{1,2}\) signifies combination of \(1{\mathrm{st}}\) and \(2{\mathrm{nd}}\) \(0^{\circ }\)-move as a whole.
-
\(0^{\circ }_{1,3}\) signifies 3 consecutive linear move (\(1{\mathrm{st}}\), \(2{\mathrm{nd}}\), \(3{\mathrm{rd}}\) move together) and so on and so forth.
MLS mixing uses the proposed shift patterns and the precedence order of these shift-patterns are set according to their mixing completion amount from higher to lower which is as follows:
According to MLS_CP, if a \(0^{\circ }_{1}\)-shift is available, then the droplet tends to achieve more consecutive straight run movements \(0^{\circ }_{2}\), \(0^{\circ }_{3}\) or a \(0^{\circ }_{4}\)-shift in a greedy manner until a collision occurs with other droplets or path exhausted for further linear movement. If an X-shift is to be obtained, then left and right distance to the boundary from the current coordinate position is checked. If routing constraints on both sides are satisfied, then the droplet is moved in the direction of maximum length available [4]. However, if both the left and right distances are equal (line no. 20–22, Fig. 12), then the droplet will choose to move towards another droplet, which will co-parent its child in the immediate next level of sequence graph. If routing constraints are not satisfied in either direction, then the droplet takes an \(180^{\circ }\)-shift from its current position rather stalling the droplet on the same place.
As Stall the droplet will accomplish 0% mixing completion and mixing time would increase and hence an \(180^{\circ }\)-shift is always preferable over STALL operation while avoiding deadlock situation for more than one droplet on the chip. That is where MLS patterns differ compared to traditional routing as in traditional VLSI routing going backward, i.e., \(180^{\circ }\)-shift is always a loss proposition in terms of time and cost but in MLS it is profitable over STALL operation.
The pseudocode for Split_and_Merge Module is given in Fig. 13, where SPLIT_SETs are formed for all completed mixing operations. When fully mixed droplets are at corner electrode, there will be 3 coordinates at max in the SPLIT_SET.
Else, the droplets coordinate position can of two types, i.e., it can either reside on boundary electrode cell or at some intermediate cells of the chip. Hence, there will be either 2 or 4 coordinates in the Split_Set. Manhattan distances are calculated from all the coordinates (x, y) of the Spit_Sets of the co-parents and the least distance is found out [4]. An electrode at distance Min (Manhattan Distance)/2 is chosen as target cell and the corresponding droplets are routed to that target electrode (line no. 5–8, Fig. 13). The mixing operations resume again for next level of synthesis according to the application graph (G) and the process continues till the entire assay synthesis ends.
In our proposed synthesis we try to follow the most profitable mixing pattern of a \(2\times 6\) module and after each \(0^{\circ }_{4}\)-shift the droplet has to take a mandatory \(90^{\circ }\)-shift. The minimum time-steps (t) required in MLS for 100% mixing completion is 27t where \({t}= 1\) time-step and \({f}=16\) Hz.
4.4 MLS-chip architecture
From the derived shift patterns shown in Figs. 14 and 16 we propose a new application specific chip architecture for MLS approach (for \(|M_{j}| \le 4\); at any layer \(L_{i} \exists G\)). The re-configurable operations (mixing) are performed along the boundary cells of the chip (Fig. 15a, b), and the center area of the chip is left out for non-re-configurable operations like containing wash droplets (\(W_i\)), heater and detection units, etc. Such an architecture obviously reduce routing costs among various stages of a bio-protocol and allow the MLS synthesis to be done in optimum time with a symmetric pattern obtained for all the mixing operations \(M_{1},\dots , M_{4}\) (Fig. 15).
More number of parallel mix. operations (\(>4\)) can be mapped with bigger chip sizes in a similar manner and the symmetric mix-pattern can be achieved. A typical MLS-architecture of a \(8\times 8\) chip for PCR bioassay is shown below (Fig. 15) for example purpose.
4.5 Illustrative example: mixing stages for PCR
The mixing stages of PCR shown in Fig. 5a consist of seven mixing operations denoted by \(M_{1}\) to \(M_{7}\). The path obtained by MLS method for simultaneous mixing operations (\(M_{1}\), \(M_{2}\), \(M_{3}\) and \(M_{4}\)) in the first layer of PCR are shown by red, green, violet and blue color, respectively, in Fig. 14 for an \(8\times 8\) chip size. Figure 16a, b shows the MLS mixing-patterns (path) according to our proposed shifts-movements obtained till \(17{\mathrm{th}}\) and \(31{\mathrm{st}}\) time-steps, respectively. The corresponding mixing completion percentages are computed and shown below. The associated coordinate positions written in subscript are the final placement of the droplet on the chip and RHS value signifies respective mixing completion percentage till 17t where t = one time-step.
The example Fig. 14 represents the initial arrangement for a PCR bio-assay synthesis and Fig. 15a, b shows the respective patterns for each of the Mixing operations of layer 1 which are symmetric to each other. The obtained shifts with respect to time-steps (t) are given below.
The symmetric pattern mentioned above is valid for all the mixing operations as shown in Fig. 15a, b and it essentially optimize the mixing procedure with minimum area and lesser time to finish the mixing. Table 3 shows mixing completion percentage and elapsed time with respect to each shift-pattern for a PCR assay.
MLS pattern also decreases the overhead cost for washing the common cells which are typically used by two or more types of fluids. Hence, in between two fluids flow through the same cell, wash operation must be performed. From Fig. 15a , it is evident that with the MLS pattern the cell with 2, 7 time stamps (the cell number is designated by the time-stamps at which two different fluid flows on the cell, respectively, at 2t and 7t). With careful observation we found, only the following cells needs to be washed.
Cell no: (2–7) which having a time gap of 5t and cell no. (8–15), (9–16) and (10–17) having a time gap of 7t such that the wash droplet \(W_{i}\) can easily be passed in that time gap. For example in first 17 time-steps, out of 17 cells traversed by a mixer droplet, only 4 cells required to be washed. The washing algorithm for MLS is discussed in detail in next section.
The corresponding mixing completion (100%) and their respective coordinate positions at \(31t\) are shown in Fig. 15b which are \(M_{1(1,3)}\), \(M_{2(1,6)}\), \(M_{3(8,3)}\), and \(M_{4(6,8)}\). Next at \(32{\mathrm{nd}}\) time-step, \(M_{1}\), \(M_{2}\), \(M_{3}\), \(M_{4}\) split according to the Split_and_Merge algorithm which requires 4 time-steps (Fig. 16).
On 36t, next level of mixing operations \(M_{5}\) and \(M_{6}\) can be started as per the sequence graph of PCR and completed at 62t, then again the Split_and_Merge of 2nd level of PCR will be done. Similarly, the merging of \(M_{5}\) and \(M_{6}\) to form \(M_{7}\) is done (ended at 65t) followed by the final mixing operation \(M_{7}\) which needs further 27 steps. Thus, \(M_{5}\), \(M_{6}\) and \(M_{7}\) able to finish the mixing in optimum number of time steps shown in Fig. 17 and the PCR is completed at \(92{\mathrm{nd}}\) time-steps. Thus, total time required to finish the PCR assay is \((92 \times 0.0625) = 5.75\,{\mathrm{s}}\) which is approximately 40% improvement compared to earlier module-based methods. Similarly, we have applied MLS methods on other available benchmark assays [5] including some hard test benches [5] and the obtained result is quite significant.
5 Congestion avoidance in MLS method
In MLS-CP method, entire mixing operation has been converted into routing steps of various shift patterns where the mixer-droplets traverse on the chip according to the precedence . One shortfall of this method is it will essentially cause higher congestion on the chip as time progresses. Also for more number of parallel mixing operations (\(\ge 4\)) incurred more congestion at the later stage of the synthesis process and the available route path decreases with time. Accordingly the washing overhead also increases with more congestion on the chip. To avoid such congestion and to reduce the overhead washing cost we have introduced a modified model of module-less synthesis.
In this Modified-Module-Less-Synthesis (MMLS) approach, we have introduced the concept of chain formation by various shift-movements as shown in Fig. 18. We run (\(\ge 4\)) number of mixing operations simultaneously on a minimum \(8\times 8\) chip size and more flexibility is given with the starting locations, i.e., mixing can now start from the corner area as well as any middle area of the chip. To tackle these extra flexibility in the new modified model, we restrict the consecutive linear movements up to \(0^{\circ }_{1,3}\) and discarded the next higher \(0^{\circ }_{4}\)-shift from the list of available shifts (S). As forming a chain with \(0^{\circ }_{1,4}\) on the chip would take more space and due to that other non-reconfigurable operations may be compromised and on the other side just discarding \(0^{\circ }_{4}\) deteriorate the mixing performance to a negligible amount as \(0^{\circ }_{1,3}\) and \(0^{\circ }_{1,4}\) almost accomplishes similar amount of mixing individually (Table 2).
MMLS first attempts to form the chain mixing patterns if possible, else traditional MLS shifts are followed [4]. Now, to form the chain the droplet need to take two consecutive (\(90^{\circ }\)-shift) on \(4{\mathrm{th}}\) and \(5{\mathrm{th}}\) time-step even though it might have the scope of taking a better shift-movement (\(0_{3}/0_{2}/0_{1}\)), which would have accomplished more mixing percentage at the end of respective time steps. However, taking 2 consecutive \(90^{\circ }\)-shifts provide the opportunity to get one more \(Z_{3}\)-shifts on \(6{\mathrm{th}}\), \(7{\mathrm{th}}\), \(8{\mathrm{th}}\) time steps which ultimately form the chain and accomplish much faster mixing. Therefore, forming a chain might seem to be disadvantageous but over the long run, it becomes beneficial compared to traditional MLS process, as the chip-space used for that particular mixing operation is much lesser (\(5\times 2 = 10\) cells only).
As in Fig. 18, all the 5 mixer droplets \(M_{1}, M_{2},\dots , M_{5}\) starts simultaneously on the \(8\times 8\) chip and out of these 5-mixer droplets \(M_{1}, M_{2}\) and \(M_{3}\) are able to form a chain on the chip shown by red, blue and green color. Droplet \(M_{4}\) and \(M_{5}\) were not able to form the chain as there was not enough space on the chip to take two consecutive \(90^{\circ }\)-shifts on \(4{\mathrm{th}}\) and \(5{\mathrm{th}}\) time-steps and hence they follow earlier MLS shift movements.
As soon as we are able to form the chain, all cells consisting the chain will be blocked (\(5\times 2 = 10\) cells only) for the entire duration of the mixing, which is 36t in this case. Thus, for up to 32 time-steps, no other malicious droplet can intrude the chain and there is no scope of cross-contamination with other droplets. Also shift movement complexity of the mixer droplet reduced within the chain as the sequence of shift-movements gets fixed within a chain, which is as follows:
The chains (Fig. 18) function similar to a \(2\times 5\) module and we can omit the washing process within the chain as the mixer droplet is homogeneous in nature and there is no need to wash until the entire mixing gets finished within a chain. Only for the non-chain mixing operations (\(M_{4}\) and \(M_{5}\) in Fig. 19), we need to consider the washing overhead at each time-steps. The MLS_washing algorithm is shown in Fig. 19.
\(R_{1}\) , \(R_{2}\) are washing ratio for chain and non-chain mixing operations and \(\gamma _{1}=0.6\) and \(\gamma _{2}=0.4\) chosen as weights for chain and non-chain mixing. The wash ratio \(R_{1}\) is designated as the number of cells washed in a \(2\times 5\) chain in respect of total time steps needed for the mixing operation. For non-chain mixing \(R_{2}\) is designated as the ratio of total number of common cells utilized by two or more droplets and total number of used cells shown in Algorithm 3 [line no. 10–11]. The overall wash ratio (R) turns out to be around 0.38, i.e., only 38% cells needs to be washed. Therefore, the modified-MLS (MMLS) process is somewhat a combined approach of module-based mixing and module-less mixing strategies, incorporating advantages from both. From Fig. 18, it is seen for \(M_{1}\), \(M_{2}\) and \(M_{3}\) mixing operations, chain can be formed which is somewhat similar to module-based mixing but without the requirement of padding cells all around the chain. Thus, a lot of chip space can be saved as most of the mixing operation usually starts from the boundary cells closer to the chip reservoirs. \(M_{4}\) and \(M_{5}\) droplets are unable to form a chain and thus they follow earlier MLS shift-patterns as shown by orange and black color, respectively, as shown in Fig. 18.
6 Experimental results
In order to evaluate the proposed MLS/MMLS algorithm in Cyberphysical DMFB, two types of simulations have been performed. Primary focus has been given on determining the completion time improvements by MLS compared to the existing module-based synthesis techniques [26] using real-life bio-assays. Secondarily MLS method is applied on different synthetic benchmarks to verify successful synthesis completion of the test cases in reasonable time. For experiments we have used five real-life applications: (1) the mixing stage of PCR (7 operations) as already discussed in Sect. 4.5; (2) In_vitro Diagnostics on human physiological fluids (IVD, 28 operations); (3) the colorimetric protein assay (CPA, 103 operations); (4) the interpolation dilution of a protein (IDP, 71 operations) [32]; and (5) the sample preparation for plasmid DNA (PDNA, 19 operations) [14]. The sequence graphs for PCR, IVD, and CPA protocols can be found in [25]. Six synthetic benchmarks (hard test 1–6) are also used [5]. The simulation program is implemented in C++ language on a 2.4 GHz Intel core i3 (M370) machine with 8 GB of memory. The findings are quite encouraging and discussed below (Fig. 20).
6.1 Results on real-life assays
The MLS synthesis results on IVD of physiological fluids and PCR benchmarks are compared with other module-based synthesis technique which are based on ILP and Tabu-Search [18]. For various chip sizes (48 cells to 144 cells), the assay synthesis time is promisingly improved around 45% by MLS approach as shown in Table 4.
6.2 Chip size versus completion time in MLS
The scalability of the proposed method has also been tested by varying the chip size from \(7\times 7\) to \(16\times 16\) (49 to 256 cells) for PCR bioassay and it is observed that the completion time varies with chip size. Table 5 shows mixing completion times for different chip sizes. We run 4 parallel mixing operations of PCR (stage I) on different chip sizes starting from \(7\times 7\) to \(16\times 16\). For 4 number of parallel mixing operations, completion-time decreases till the chip size reaches \(12\times 12\) but after that increment of chip size doesn’t effect in completion time. So we can conclude that for 4 number of simultaneous mixing requirements, a maximum of \(12\times 12\) chip size is sufficient to get optimum completion time by MLS patterns. Hence, For CPA , IDP and PDNA real-life assays, the time improvement is tested on a \(12\times 12\) chip only and it is found to be around 35% as given in Table 6.
6.3 Benchmark suites (BS I and BS III) and hard test benches
The MLS algorithm is tested on Benchmark Suite-I (BS-I), which comprises of 30 hard test benches and on most commonly used Benchmark Suite III (BS-III) [8]. We have also randomly taken 6 harder test cases [5] with more than 8 number of parallel mixing operations in a single stage. For such cases, it is difficult for the MLS to find all mixing patterns and thus increases completion time as well as length of Wash path (W_path) which ultimately increases wash operation complexity as well as number of wash droplet requirement. Modified MLS (MMLS) perform complete synthesis for all the 30 different test cases in BS-I well below 7 s. For PCR and BS-III, the detailed results are given in Table 7 which shows at max. 38% cells needed to be washed out of all used cells the mixer droplets traversed. From Table 7 , it is evident as more number of chain mixing can be formed, the overall wash requirement decreases.
Finally , for hard test cases (test 1–6) and for Protein-I , Protein II assays, where number of parallel mixing is \(\ge 8\) the algorithm is tested with disabling the wash operation as well as enabling the wash operation. The average overhead time taken for wash operation (wash operation enabled) by MMLS is well below 8% which is quite promising and shown in Table 8.
7 Conclusion
The proposed synthesis method (MLS) for cyberphysical DMFBs provide faster bio-assay results by eliminating the concept of dedicated virtual modules. Mixing operations are performed through any path, available on the microfluidic chip and in accordance with the proposed shift movements (patterns). According to the experimental results, module-less synthesis produces significant improvement in terms of assay synthesis time and hence modified-MLS particularly important and beneficial for more complex, high-performance synthesis problems (bio-protocols) with higher number of (\(\ge 8\)) parallel mixing operations. Thus, it may be also helpful in detecting various epidemic or pandemic diseases like malaria, dengue, AIDS, novel corona (COVID-19), etc., and also in eliminating futuristic bio-terror attacks with the help of appropriate cyberphysical inclusions to the DMF chips.
References
Alistar M, Pop P, Madsen J (2016) Synthesis of application-specific fault-tolerant digital microfluidic biochip architectures. IEEE Trans Comput Aided Des Integr Circuits Syst 35(5):764–777
Cartwright S (2018) Introduction to experimental error. https://www.sheffield.ac.uk/polopoly_fs/1.14221!/file/IntroToExperimentalErrors_y2. Accessed 23 July 2019
Chakrabarty K, Fair RB, Zeng J (2010) Design tools for digital microfluidic biochips: toward functional diversification and more than moore. IEEE Trans Comput Aided Des Integr Circuits Syst 29(7):1001–1017
Chakraborty S, Chakraborty S (2017) A novel approach towards biochemical synthesis on cyberphysical digital microfluidic biochip. In: 2017 30th International conference on VLSI design and 2017 16th international conference on embedded systems (VLSID). IEEE, pp 355–360
Chakraborty S, Chakraborty S, Das C, Dasgupta P (2016) Efficient two phase heuristic routing technique for digital microfluidic biochip. IET Comput Digit Tech 10(5):233–242
Chakraborty S, Das C, Chakraborty S (2018) Securing module-less synthesis on cyberphysical digital microfluidic biochips from malicious intrusions. In: 2018 31st International conference on VLSI design and 2018 17th international conference on embedded systems (VLSID), pp 467–468. https://doi.org/10.1109/VLSID.2018.116
Chang JW, Yeh SH, Huang TW, Ho TY (2013) An ILP-based routing algorithm for pin-constrained EWOD chips with obstacle avoidance. IEEE Trans Comput Aided Des Integr Circuits Syst 32(11):1655–1667
Cho M, Pan DZ (2008) A high-performance droplet routing algorithm for digital microfluidic biochips. IEEE Trans Comput Aided Des Integr Circuits Syst 27(10):1714–1724
Demonstration of 1 × 2 Cell Mixing-Module on DMFB, by Microfluidics Lab U Duke University (2018) https://www.youtube.com/watch?v=8qrksV8AadA&feature=youtu.be. Accessed 20 Feb 2020
Fair RB, DUKE Microfluidics Laboratory (2014) Department of Electrical and Computer Engineering, Duke University, USA. http://microfluidics.ee.duke.edu/. Accessed 20 Feb 2020
Huang TW, Lin CH, Ho TY (2010) A contamination aware droplet routing algorithm for the synthesis of digital microfluidic biochips. IEEE Trans Comput Aided Des Integr Circuits Syst 29(11):1682–1695
Ibrahim M, Chakrabarty K (2015) Efficient error recovery in cyberphysical digital-microfluidic biochips. IEEE Trans Multi Scale Comput Syst 1(1):46–58
Kramer MF, Coen DM (2001) Enzymatic amplification of DNA by PCR: standard procedures and optimization. Curr Protoc Mol Biol 56(1):1–15
Luo Y, Chakrabarty K, Ho TY (2012) A cyberphysical synthesis approach for error recovery in digital microfluidic biochips. In: 2012 Design, automation and test in Europe conference and exhibition (DATE). IEEE, pp 1239–1244
Luo Y, Bhattacharya BB, Ho TY, Chakrabarty K (2013) Optimization of polymerase chain reaction on a cyberphysical digital microfluidic biochip. In: Proceedings of the international conference on computer-aided design. IEEE Press, pp 622–629
Luo Y, Chakrabarty K, Ho TY (2013) Error recovery in cyberphysical digital microfluidic biochips. IEEE Trans Comput Aided Des Integr Circuits Syst 32(1):59–72
Luo Y, Chakrabarty K, Ho TY (2014) Biochemistry synthesis on a cyberphysical digital microfluidics platform under completion-time uncertainties in fluidic operations. IEEE Trans Comput Aided Des Integr Circuits Syst 33(6):903–916
Maftei E, Pop P, Madsen J (2009) Tabu search-based synthesis of dynamically reconfigurable digital microfluidic biochips. In: Proceedings of the 2009 international conference on compilers, architecture, and synthesis for embedded systems. ACM, pp 195–204
Maftei E, Pop P, Madsen J (2012) Routing-based synthesis of digital microfluidic biochips. Des Autom Embed Syst 16(1):19–44
Murran MA, Najjaran H (2012) Capacitance-based droplet position estimator for digital microfluidic devices. Lab Chip 12(11):2053–2059
Olivier PD (1992) Approximating irrational transfer functions using lagrange interpolation formula. IEE Proc D Control Theory Appl 139(1):9–12
Paik P, Pamula VK, Fair RB (2003) Rapid droplet mixers for digital microfluidic systems. Lab Chip 3(4):253–259
Pollack MG, Shenderov AD, Fair R (2002) Electrowetting-based actuation of droplets for integrated microfluidics. Lab Chip 2(2):96–101
Srinivasan V, Pamula VK, Fair RB (2004) An integrated digital microfluidic lab-on-a-chip for clinical diagnostics on human physiological fluids. Lab Chip 4(4):310–315
Su F, Chakrabarty K (2006) Benchmarks for digital microfluidic biochip design and synthesis. Duke University, Department ECE
Su F, Chakrabarty K (2008) High-level synthesis of digital microfluidic biochips. ACM J Emerg Technol Comput Syst (JETC) 3(4):1
Verpoorte E, De Rooij NF (2003) Microfluidics meets mems. Proc IEEE 91(6):930–953
Welch ERF, Lin YY, Madison A, Fair RB (2011) Picoliter DNA sequencing chemistry on an electrowetting-based digital microfluidic platform. Biotechnol J 6(2):165–176. https://doi.org/10.1002/biot.201000324
Windh S, Phung C, Grissom DT, Pop P, Brisk P (2017) Performance improvements and congestion reduction for routing-based synthesis for digital microfluidic biochips. IEEE Trans CAD Integr Circuits Syst 36(1):41–54
Xu T, Chakrabarty K (2007) A cross-referencing-based droplet manipulation method for high-throughput and pin-constrained digital microfluidic arrays. In: 2007 design, automation & test in Europe conference & exhibition, Nice, pp 1–6. https://doi.org/10.1109/DATE.2007.364651
Yuh P-H, Sapatnekar S, Yang C, Chang Y (2008) A progressive-ILP based routing algorithm for cross-referencing biochips. In: 2008 45th ACM/IEEE design automation conference, Anaheim, CA, pp 284–289
Zhao Y, Xu T, Chakrabarty K (2010) Integrated control-path design and error recovery in the synthesis of digital microfluidic lab-on-chip. ACM J Emerg Technol Comput Syst (JETC) 6(3):11
Acknowledgements
The authors would like to thank SPARC, MHRD, Govt. of India for providing grant under Project Code P1037.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Chakraborty, S., Chakraborty, S. An efficient module-less synthesis approach for Digital Microfluidic Biochip. SN Appl. Sci. 2, 1442 (2020). https://doi.org/10.1007/s42452-020-3173-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s42452-020-3173-6