Abstract
The severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) infection that causes COVID-19 disease is still a major public health concern around the world. The main protease encoded by SARS-CoV-2 is a promising target for COVID-19 therapeutic development since it plays a crucial role in the virus’s life cycle. Repurposing known bioactive molecules is an effective strategy to fast-track the delivery of hits and leads in drug discovery. In this regard, this study assesses in-silico, 17 acridone-based alkaloids for their activity against SARS-CoV-2 main protease. The quantum chemical computations imply that the acridone alkaloids will interact better in the active sites of the enzymes due to their low energy gap. They also have good oral bioavailability as rationalized by “no rule of five violation" and favorable pharmacokinetics parameters. From the docking results, many of the alkaloids displayed a higher binding affinity than nirmatrelvir, an authorised protease inhibitor. Compound 3 (5-hydroxynoracronycine alcohol), with the better binding affinities (6W63, − 7.094 kcal/mol; 5R82, − 5.839 kcal/mol) and unique structural features is a viable candidate that could be investigated further for development of a novel target specific chemotherapeutic agent to stop SARS-CoV-2 invasion.
Similar content being viewed by others
1 Introduction
Coronavirus disease 2019 popularly called COVID-19 is an acute respiratory infectious disease that emerged at the end of 2019. The disease is caused by a novel coronavirus named severe acute respiratory syndrome coronavirus-2 (SARS-CoV-2) [1]. As of June 23, 2022, there were more than 539 million confirmed cases of COVID-19 and over 6.3 million deaths recorded worldwide (https://covid19.who.int/, accessed on 24 June 2022). Although COVID-19 cases are declining, the world is still grappling with the virus; the virus continues to infect new individuals consistently and causing new infection clusters [2].
There are currently several vaccines available that offer protection against the virus and help reduce the severity of COVID-19; however, the emergence of SARS-CoV-2 variants of concern threatens the effectiveness of these vaccines and could lead to a surge in cases and fatalities [3]. Additionally, there are a few approved chemotherapeutic (target-specific) drugs to treat this viral disease. Currently, only one protease inhibitor (Paxlovid, nirmatrelvir packaged with ritonavir) is given an emergency use authorization by the Food and Drug Administration for treating SARS-CoV-2 invasion [4]. These developments underscore the importance of developing antiviral drugs to fight SARS-CoV-2 infections. Therefore, it is imperative to keep searching for potent, safe and target-specific chemotherapeutic agents that could complement the vaccines and offer protection against the virus or treat the symptoms of the disease.
One of the core therapeutic strategies to target viruses is aimed at targeting proteins that are involved in viral replication and release [5]. SARS-CoV-2 genome encodes two polyproteins which are processed by the main protease (MPRO, also called 3CL protease) and a papain-like protease [6]. The main protease, a cysteine protease, mediates most maturation cleavage events [7]. MPRO of SARS-CoV is essential for viral survival in the host cells, thereby making it an excellent target for antiviral therapy [8]. Consequently, the inhibition of MPRO will cause definite damage to the virus. It is therefore crucial and a high priority to develop more MPRO inhibitors of SARS-CoV-2 infection.
Repurposing known bioactive molecules is an effective strategy to rapidly identify clinical candidates [9]. Old natural compounds are increasingly finding new applications in antiviral research [10]. For example, at the onset of the COVID-19 pandemic, chloroquine and hydroxychloroquine, the synthetic derivatives of the natural cinchona alkaloid quinine were controversially repurposed for the treatment of SARS-CoV-2 infection [11, 12]. Natural products and their derivatives remain an infinite resource for drug discovery and development. They contain a variety of pharmacophore patterns with a wide range of shapes and chemical complexity that have evolved over thousands of years through natural selection. Alkaloids are one of the most common phytochemicals found in plant families with prominent antiviral properties [13]. Alkaloids play an essential role in human medicine. Currently, no less than 57 plant-derived alkaloids are licensed as drugs or used in clinical environment throughout the world [14]. Acridone alkaloids are ubiquitous in the Rutaceae family, particularly in citrus plants, and many of them have biological properties such as antiviral, antimalarial, anticancer, antitumor, anti-inflammatory, and antiallergic activities [15, 16].
Some of these plants which include Citrus aurantifolia, Citrus aurantium, Citrus paradise, and Citrus limon were purportedly used in the management of COVID-19 [17]. Besides, antiviral efficacy of several types of acridones against various viruses has been demonstrated in multiple studies. Citrusinine I (Fig. 1, II) for instance showed potent antiherpetic activity against herpes simplex virus type 1 and type 2 [18]. Similarly, the acridone derivatives of type III are effective inhibitors of dengue virus and Junin virus infection at noncytotoxic concentrations [19, 20]. The intriguing biological activities of the acridone alkaloids encouraged our interest in their antiviral activity against SARS-COV-2 MPRO.
Obtaining a meaningful quantity of pure alkaloids for bioassay, even for a modest in vitro screening program, is a significant challenge [21]. Moreover, the costs and length of time required to develop a new drug are prohibitively high, with a high attrition rate. It is estimated that the mean cost of developing a new drug is about $1.3 billion USD [22], and the attrition rate of drug candidates could be over 90% [23, 24]. These drawbacks make repurposing and in-silico-aided drug discovery an appealing approach for drug discovery and development. Consequently, we report the in-silico investigation of acridone alkaloids against SARS-CoV-2 MPRO.
2 Methodology
2.1 Molecular Docking Study
2.1.1 Ligand Preparation
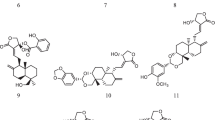
17 natural and synthetic derivatives of acridone-based alkaloids previously reported in literature were used as the ligand molecules (Fig. 2, S1) and their structure/ simplified molecular-input line-entry system (SMILES) were obtained from PubChem database (https://www.pubchem.ncbi.nlm.nih.gov/), the unavailable structures in PubChem were drawn using ChemDraw and saved in SDF format.
2.1.2 Protein Selection and Preparation for Main Protease SARS-COV-2
The structures of the target proteins for the main protease SARS-CoV-2 PDB ID: 5R82 and 6W63 in PDB format were retrieved from protein data bank (http://www.rcsb.org/pdb/home/home.do). These two were selected as a result of the presence of an inhibitor ligand at the active site of the protein. The crystallized structure was imported and viewed with Maestro 11.8 software from Schrodinger. The structure was prepared using the protein preparation wizard in the maestro interface at a pH of 7.0. During the preparation process, water molecules and other interfering ligands were removed from the protein structure.
2.1.3 Receptor Grid Generation
The receptor grid defines the interaction area between the protein and the ligand. Using the receptor grid generation tool in maestro 11.8 the binding site was identified. The coordinates of the receptor grid box was centered around the x, y and z axis for each of the protein structures.
2.1.4 Molecular Docking using Glide for SARS-CoV-2
The docking process was carried out using Glide tool on maestro 11.8. The prepared ligands were docked into the active site (x = 10.74, y = -0.67, z = 24.9) for 5R82 protein target and (x = -20.59, y = 18.08, z = -27.0) for 6W63 protein target using the standard precision algorithm (SP) and the ligand sampling option selected as flexible, then followed by extra precision with a none refine only selection for the ligand sampling, which was used to rerank the SP results to minimize false-positive [25]. At the end of the analysis, the ligand interaction tool was used to view the interaction of the ligands with the amino acid residues at the active site of the protein target.
2.2 Computational Pharmacokinetics
SwissADME (https://www.swissadme.ch), an online tool, was used to predict the pharmacokinetics and drug likeness of the acridone alkaloids. The chemical structures of each of the molecules was pasted in SMILES format to generate their pharmacokinetics properties [26].
2.3 Quantum Chemical Descriptors
The structures of the molecules were optimized using DFT at the B3LYP/6-31G* level of theory [27]. Optimization was carried out on the most stable conformers via molecular mechanics force field using spartan 14 computational chemistry software on an Intel Core i3-7020U mobile workstation (4.00 GB RAM at 2.30 GHz). The energies of the frontier molecular orbitals (FMOs) were calculated. The energy band gap, ΔE (Eq. 1), chemical hardness, η (Eq. 2), chemical softness, δ (Eq. 3) and chemical potential, CP (Eq. 4) were calculated from EHOMO and ELUMO. Lipophilicity (Log P), polar surface area (PSA), polarizability, hydrogen bond donor (HBD) and hydrogen bond acceptor (HBA) were also computed [28, 29].
3 Results and Discussion
3.1 Molecular Docking
17 acridone alkaloids were examined for their binding affinity and binding orientation against SARS-COV-2 MPRO by molecular docking. The results obtained show that the binding energy of the acridone alkaloids was between − 3.78 and − 6.09 kcal/mol against the target protein, 5R82 (Table 1 and S1) summarize the binding activity of the three best docked compounds. Compound 15 (glycocitrine I) had the highest binding energy value of − 6.09 kcal/mol. The docking results revealed that the molecules have higher binding affinity than Nirmatrelvir, a well-known SARS-COV-2 main protease inhibitor which has a binding energy value of − 4.46 kcal/mol. Compound 15 interacted with key amino acids Glu166 and Gly143 in the protein target through hydrogen bonding (Fig. 3, a). The next 2 top-hit alkaloids are compound 3 (5-hydroxynoracronycine alcohol, − 5.84 kcal/mol) and compound 14 (citpressine II, − 5.79 kcal/mol). Similar to 15, they formed strong interactions with the active sites of 5R82 via hydrogen bonding.
Against the target protein 6W63, the binding energy of the alkaloids range from − 3.92 to − 7.09 kcal/mol (Table S2). Compound 3 (5-hydroxynoracronycine alcohol) having the best binding energy value of − 7.09 kcal/mol, also demonstrated higher binding energy compared to Nirmatrelvir with binding energy value of − 4.79 kcal/mol (Table 1). Compound 3 establishes two interactions in the protein target through conventional hydrogen bonding and pi-pi interaction, with the amino acids Hie41, Glu166 and Hie163 (Fig. 3, b). Likewise, the compounds 10 (citrusinine I) and 2 (5-hydroxynoracronycine) with high binding scores bound to Cys44, Hie41, Hie164, and Thr190, respectively, formed conventional hydrogen bonding and pi-pi interaction. It is instructive to note that a related activity profile was reported for compound 2 elsewhere [9]. Similar interactions have been reported for these amino acids and it was established that Gly143, Cys145, Glu166 and His163 of SARS-CoV-2 Mpro were the most attractive residues to form hydrogen bonds with the ligands [30].
It was observed that majority of the alkaloids had a binding affinity for both target proteins that surpassed that of the reference inhibitor. It appears that protein 6W63 is more susceptible to the acridone alkaloids than 5R82. Structurally, it can be inferred that substitution on the acridone (compound 1) correlates to better MPRO activity. For example, compound 10 (− 6.51 kcal/mol), an N-methyl acridone containing dihydroxy and dimethoxy groups on position 1 and 5 and position 3 and 4, respectively, had a higher binding affinity than compound 1 (− 4.39 kcal/mol, Table S2). On the other hand, demethylation of the 3-methoxy and/or acetylation of 5-hydroxy group of compound 10 resulted in minimal loss of activity, as shown by the decreased binding score of − 4.47 and − 5.57 kcal/mol for compound 11 and 12, respectively (Table S2). However, compound 13, with a methoxy group on position 3 of ring A and a methoxy and a hydroxy groups on ring C, possessed even weaker activity (− 3.92 kcal/mol) than compound 1.
Compound 3 (5-hydroxynoracronycine alcohol), a pyranoacridone showed the best activity for the two studied MPRO proteins. Its higher activity compared to other pyranoacridones (2, 4, 5, 6, 7, 8 & 9) and the methoxyacridones (10, 11, 12, 13, 14, 16 & 17) can be premised on additional hydrogen bonding contributed by the extra hydroxymethyl moiety on the pyran ring. Strikingly, compound 15, a 4-prenylated acridone had a slightly higher binding energy (with protein 5R82) than compound 3. The prenyl group has been known to increase affinity for biological membranes and also an improved interaction with proteins [31, 32]. It is instructive to note that the pyranoacridones listed above are also a variant of 4-prenylation; with cyclization between C-3 OH and C-4 prenyl group. All ligand-receptor interactions are presented (Figures S2a-r and S3a-r).
3.2 Pharmacokinetics
Drug-likeness is a huge factor that must be taken into consideration for any compound to be considered a good drug candidate. Lipinski and colleagues proposed the rule of five (Ro5) which are: molecular weight < 500, (octanol/water coefficient) log P ≤ 5, hydrogen bond donors ≤ 5, hydrogen bond acceptors ≤ 10 [33]. Ro5 together with polar surface area (PSA) can be associated with drugs oral bioavailability or promiscuity [34]. Orally bioavailable compounds with a PSA less than 140 A2 have been found to exhibit better intestinal absorption [35] while compounds that penetrate the blood-brain barrier typically have PSA values less than 70 A2 [34]. Thus, the physicochemical properties of the acridone alkaloids were evaluated and the results shown in Table 2. The PSA values of the compounds are within the acceptable cutoff, varying from 24.86 to 91.92 A2. This indicates their possible intestinal permeability. Furthermore, all the acridone alkaloids investigated did not violate any of the rules, supporting them as good drug candidates.
On oral administration, it is required that the drug seamlessly transits through biological membranes, where most of the drugs are absorbed, as does the distribution of the drug across the blood–brain barrier (BBB) [35]. Sufficient intestinal absorption of a drug is crucial for its pharmacokinetic profile in the body, affecting its absorption, distribution and elimination [35], and ultimately its final actions. The results of the pharmacokinetics properties of the acridone alkaloids are shown in Table 3. All the compounds show high human intestinal absorption, this denotes that the compounds could be easily absorbed from the intestinal tract upon oral administration. It should be noted that these molecules are moderately lipophilic and should therefore theoretically be better able to penetrate across the membrane throughout the intestine [36].
The blood brain barrier (BBB) contains active efflux transporters (e.g., Permeability glycoprotein) and various enzymatic proteins which play a large contributing role in bioavailability and distribution of orally administered drugs. BBB acts as a sieve that prevents access of polar molecules to the brain. A BBB model will predict whether a compound will cross over the barrier and exert therapeutic effects on the brain [37]. From the result, only 35% of the compounds have the ability to cross the blood brain barrier. Interestingly, none of the compounds is a substrate for P-gp (Permeability glycoprotein). P-gp is an efflux transporter pump found in the cell membrane which is responsible for conveying drugs away from the cell membrane and cytoplasm which causes therapeutic failure when the concentration of the drug is reduced [38].
3.3 Quantum Chemical Descriptors
Table 4 shows the HOMO and LUMO energies as well as the chemical descriptors of the selected compounds. Ionization energy is defined as the energy required to remove an electron from a molecule’s ground state and it is related to EHOMO. Hence the higher the EHOMO of a molecule, the lower its potential to donate an electron and high value indicates chemical stability [38]. ELUMO is related to the electron affinity which is defined as the energy released when a molecule in the ground state captures an electron and the lower the ELUMO of a molecule, the more its ability to accept electrons from nearby molecules. The EHOMO for the compounds range from − 5.21 to − 5.95 eV while ELUMO values range from − 1.25 to 2.36 eV. The energy band gap explains the stabilities of the molecules. It further furnishes qualitative information on the interaction of a molecule with an enzyme [39, 40]. The energy band gap is the difference between EHOMO and ELUMO and low energy band gap indicates high chemical reactivity of the compounds. From Table 4, the acridone alkaloids with the lowest band gap are compound 2 < compound 6 < compound 3 ? compound 11. Therefore, compound 2 is expected to interact better with the enzymes than the other alkaloids.
The HOMO and LUMO of Acridone are on almost all parts of the molecule. The region most preferred for electrophilic attack is the carbonyl oxygen (red) while the –NH group is the most preferred site for nucleophilic attack (Fig. 4) [41, 42]. The HOMO of 5-Hydroxynoracronycine spreads across the entire molecule leaving out the methyl groups on the pyran ring and the carbonyl group. However, the LUMO left out both the C4, C5-double bond and the methyl groups of the heterocyclic pyran ring. The electrostatic potential shows that the hydroxyl and the carbonyl groups are the most preferred sites for electrophilic attack while the N-methyl group is the most preferred site for nucleophilic attack (Fig S4a). The HOMO of 5-Hydroxynoracronycine alcohol spreads across the entire molecule leaving the carbonyl group. Similarly, the LUMO also spreads throughout the entire molecule except for the pyran ring. The electrostatic potential shows that the hydroxyl group on the pyran ring is the most preferred site for electrophilic attack while the N-methyl group is the most preferred site for nucleophilic attack (Fig S4b). The HOMO of 9-Hydroxyacronycine spreads across the entire molecule leaving out both the dimethyl and the oxygen atom of 2 H-chromene. Also, the LUMO spreads across almost the entire molecule leaving out the dimethyl flanks of the pyran ring and the hydroxy group. The electrostatic potential shows that the carbonyl and the methoxy oxygen are the most preferred sites for electrophilic attack while the hydrogen on the hydroxy group is the most preferred site for nucleophilic attack (Fig S4c). The HOMO of Citracridone-I spreads across the entire molecule but leaving out the carbonyl, 6-hydroxy and 5-methoxy groups. Similarly, the LUMO spreads across almost the entire molecule but leaving out substantial portion of the pyran ring and the 5-methoxy group. The electrostatic potential shows that the carbonyl oxygen is the most preferred site for electrophilic attack while the hydrogen on 6-hydroxy group is the most preferred site for nucleophilic attack (Fig S4d). The HOMO of Citracridone-II spreads across almost all the entire molecule leaving out the carbonyl and methoxy groups while the LUMO spreads across almost the entire molecule leaving out part of the pyran ring and the methoxy groups. The electrostatic potential shows that the carbonyl and hydroxyl oxygen are the preferred sites for electrophilic attack while the methoxy, pyran and N-methyl groups are the most preferred sites for nucleophilic attack (Fig S4e). The HOMO of Citracridone-III spreads across the acridone and pyran rings while the LUMO spreads across almost the entire molecule leaving out part of the pyran ring and hydroxyl groups. The electrostatic potential shows that the carbonyl and hydroxyl oxygen are the preferred sites for electrophilic attack while the hydrogen on 6-hydroxy group is the most preferred site for nucleophilic attack (Fig S4f). The HOMO of Citracridone-I Derivative spreads across almost all parts of the molecule, leaving out the acetate, methyl on the pyran ring and the carbonyl group. The LUMO also spreads across the entire molecule but leaves out the double-bond and the dimethyl flanks of the pyran ring, but it was spread to the carbonyl group. The electrostatic potential shows that the pyran, carbonyl, hydroxyl and the acetate oxygen are sites for electrophilic attack while the methoxy group is the most preferred site for nucleophilic attack (Fig S4g). The HOMO and LUMO of Citracridone-I Derivative II spread across almost all parts of the molecule, with the HOMO leaving out the acetate, carbonyl, methoxy and part of the pyran ring. The LUMO follows a similar pattern as the HOMO but spreads to the carbonyl group. The electrostatic potential shows that the carbonyl, hydroxyl and the acetate oxygen are the preferred site for electrophilic attack while the pyran ring, methyl on the acetate and N-methyl groups are the most preferred site for nucleophilic attack (Fig S4h). The HOMO and LUMO of Citrusinine-I spread across almost all parts of the molecule, with the HOMO leaving out the carbonyl and methyl on the methoxy groups while the LUMO leaves out the methoxy groups. The electrostatic potential shows that the carbonyl, hydroxyl and methoxy oxygen are the preferred site for electrophilic attack while the methyl on the N-methyl group is the most preferred site for nucleophilic attack (Fig S4i). The HOMO and LUMO of Citrusinine-II spread across almost all parts of the molecule, with the HOMO leaving out the carbonyl and methyl on the methoxy group while the LUMO leaves out the methoxy and 5-hydroxy groups. The electrostatic potential shows that the carbonyl, hydroxyl and methoxy oxygen are the preferred site for electrophilic attack while the methyl on the methoxy group is the most preferred site for nucleophilic attack (Fig S4j). The HOMO and LUMO of Citrusinine-I derivative spread across almost all parts of the molecule, with the HOMO leaving out the acetate, carbonyl and methyl on the methoxy groups while the LUMO leaves out the acetate and the 4-methoxy group. The electrostatic potential shows that the carbonyl, hydroxyl and the acetate oxygen are the most preferred sites for electrophilic attack while the methyl on the methoxy groups are most preferred sites for nucleophilic attack (Fig S4k). The HOMO and LUMO of Citpressine I spread across almost all parts of the molecule, with the HOMO leaving out the 6-hydroxy, the methoxy groups and the carbonyl carbon while the LUMO leaves out the methyl on the methoxy groups. The electrostatic potential shows that the carbonyl and hydroxyl oxygen are the most preferred sites for electrophilic attack while the methyl on the methoxy and N-methyl groups are most preferred sites for nucleophilic attack (Fig S4l). The HOMO and LUMO of Citpressine II spread across almost all parts of the molecule, with the HOMO leaving out the methoxy groups and the carbonyl carbon while the LUMO leaves out the methyl on the methoxy groups. The electrostatic potential shows that the carbonyl and hydroxyl oxygen are the most preferred sites for electrophilic attack while the methoxy and N-methyl groups are most preferred sites for nucleophilic attack (Fig S4m). The HOMO and LUMO of Glycocitrine I spread across almost all regions of the molecule but leaving out the prenyl group and the methoxy methyl. In addition, the HOMO left out the carbonyl group while the LUMO left out the 5-hydroxyl group. The highest negative (red) region is located at the carbonyl and hydroxyl oxygen while blue (positive region) is located on the methoxy group (Fig S4n). The HOMO and LUMO of Grandisine I spread across almost every part of the compound. However, the HOMO left out the methoxy groups and the carbonyl carbon while the LUMO leaves out the methyl groups. The highest negative sites (orange and red) in the electrostatic potential map are located on the carbonyl and hydroxyl oxygen indicating the preferred site for electrophilic attack while the methoxy groups are preferred sites for nucleophilic attack that is, the blue, positive regions (Fig S4o). The HOMO and LUMO of N10-Citpressine II derivative spread across it in the same manner as Grandisine I. The highest negative (orange and red) and the highest positive sites (blue) in the electrostatic potential map are as observed in Grandisine I (Fig S4p).
4 Conclusion
This study investigated the potential of 17 acridone-based alkaloids to inhibit SARS-CoV-2 MPRO proteins. As evidenced from the molecular docking studies, the SARS-CoV-2 MPRO proteins, which are crucial to viral replication, could be effectively inhibited by these alkaloids. The majority of these alkaloids displayed binding affinities that were greater than that of nirmatrelvir, a well-known protease inhibitor. Furthermore, these alkaloids showed favourable drug-like qualities, and also had good ADMET profiles as predicted by DFT calculations and SwissADME program. The remarkable anti-protease activity of compound 3, as indicated by its binding energies (6W63, − 7.094 kcal/mol; 5R82, − 5.839 kcal/mol), its favourable druglikeness, and its unique structural characteristics, makes it a promising candidate that could be further explored for development of a novel target specific chemotherapeutical agent to stop SARS-CoV-2 infection.
References
Adem S, VEyupoglu V, Ibrahim IM, Sarfraz I, Rasul A, Ali M, Elfiky AA (2022) Multidimensional in silico strategy for identification of natural polyphenols-based SARS-CoV-2 main protease (Mpro) inhibitors to unveil a hope against COVID-19. Comput Biol Med 145:105452. https://doi.org/10.1016/j.compbiomed.2022.105452
Dutta D, Naiyer S, Sabanaz M, Soni N, Singh V, Bhat KH, Singh N, Arora G, Mansuri MS (2022) COVID-19 Diagnosis: A Comprehensive Review of the RT-qPCR Method for Detection of SARS-CoV-2. Diagnos 12:1503. https://doi.org/10.3390/diagnostics12061503
Aleem A, Akbar Samad AB, Slenker AK (2022) Emerging Variants of SARS-CoV-2 and Novel Therapeutics Against Coronavirus (COVID-19). 2022 May 12. In: StatPearls [Internet]. Treasure Island (FL): StatPearls Publishing
Narayanan A, Narwal M, Majowicz SA, Varricchio C, Toner SA, Ballatore C, Brancale A, Murakami KS, Jose J (2022) Identification of SARS-CoV-2 inhibitors targeting Mpro and PLpro using in-cell-protease assay. Commun Biol 5:169. https://doi.org/10.1038/s42003-022-03090-9
Jain M, Anand A, Shah A (2022) Exploring the Potential Role of Theaflavin-3,3′-Digallate in Inhibiting Various Stages of SARS-CoV-2 Life Cycle: An In-Silico Approach. Chem Afr. https://doi.org/10.1007/s42250-022-00376-7
Dampalla CS, Zheng J, Perera KD, Wong LYR, Meyerholz DK, Nguyen HN, Kashipathy MM, Battaile KP, Lovell S, Kim Y, Perlman S, Groutas WC, Chang KO (2021) Postinfection treatment with a protease inhibitor increases survival of mice with a fatal SARS-CoV-2 infection. Proc Nat Acad Sci 118:e2101555118. https://doi.org/10.1073/pnas.2101555118
Suárez D, Díaz N (2020) SARS-CoV-2 Main Protease: A Molecular Dynamics Study. J Chem Info Model 60:5815–5831. https://doi.org/10.1021/acs.jcim.0c00575
Pillaiyar T, Manickam M, Namasivayam V, Hayashi Y, Jung SH (2016) An Overview of Severe Acute Respiratory Syndrome–Coronavirus (SARS-CoV) 3CL Protease Inhibitors: Peptidomimetics and Small Molecule Chemotherapy. J Med Chem 59:6595–6628. https://doi.org/10.1021/acs.jmedchem.5b01461
Hasan A, Jannat K, Bondhon TA, Jahan R, Hossan MS, de Lourdes Pereira M, Nissapatorn V, Wiart C, Rahmatullah M (2022) Can Antimalarial Phytochemicals be a Possible Cure for COVID-19? Molecular Docking Studies of Some Phytochemicals to SARS-CoV-2 3 C-like Protease. Infect Disord Drug Targets 22:e290721195143. https://doi.org/10.2174/1871526521666210729164054
Benhander GM, Abdusalam AAA (2022) Identification of Potential Inhibitors of SARS-CoV-2 Main Protease from Allium roseum L. Molecular Docking Study. Chem Afr 5:57–67. https://doi.org/10.1007/s42250-021-00296-y
Juurlink DN (2020) Safety considerations with chloroquine, hydroxychloroquine and azithromycin in the management of SARS-CoV-2 infection. Canad Med Ass J 192:E450–E453
Lentini G, Cavalluzzi MM, Habtemariam S (2020) COVID-19, Chloroquine Repurposing, and Cardiac Safety Concern: Chirality Might Help. Molec 25:1834. https://doi.org/10.3390/molecules25081834
Majnooni MB, Fakhri S, Bahrami G, Naseri M, Farzaei MH, Echeverría J (2021) Alkaloids as Potential Phytochemicals against SARS-CoV-2: Approaches to the Associated Pivotal Mechanisms. Evidence-Based Compl Alt Med 2021:6632623. https://doi.org/10.1155/2021/6632623
Heinrich M, Mah J, Amirkia V (2021) Alkaloids Used as Medicines: Structural Phytochemistry Meets Biodiversity-An Update and Forward Look. Molec 26:1836. https://doi.org/10.3390/molecules26071836
Xu L, Li S, Liang Z, Lin H, Fu R (2018) Acridone suppresses the proliferation of human breast cancer cells in vitro via ATP-binding cassette subfamily G member 2. Oncoy Lett 15:2651–2654. https://doi.org/10.3892/ol.2017.7583
Wang C, Wan J, Mei Z, Yang X (2014) Acridone alkaloids with cytotoxic and antimalarial activities from Zanthoxylum simullans Hance. Pharmacog Mag 10:73–76. https://doi.org/10.4103/0973-1296.126669
Oderinlo OO, Adenekan OA, Alawode TT, Osamudiamen PM, Oluremi BB, Oyeneyin OE, Ngoepe MP (2021) Ethnobotanical Appraisal and In-silico Investigation of Plants Used for the Management of COVID-19 in Southwestern Nigeria. Arab J Med Arom Plants 7:151–174. https://doi.org/10.48347/IMIST.PRSM/ajmap-v7i1.23748
Yamamoto N, Furukawa H, Ito Y, Yoshida S, Maeno K, Nishiyama Y (1989) Anti-herpesvirus activity of citrusinine-I, a new acridone alkaloid, and related compounds. Antiv Res 12:21–36
Mazzucco MB, Talarico LB, Vatansever S, Carro AC, Fascio ML, D’Accorso NB, García CC, Damonte EB (2015) Antiviral activity of an N-allyl acridone against dengue virus. J Biomedic Sci 22:29. https://doi.org/10.1186/s12929-015-0134-2
Sepúlveda CS, Fascio ML, Mazzucco MB, Palacios ML, Pellón RF, García CC, D’Accorso NB, Damonte EB (2008) Synthesis and evaluation of N-substituted acridones as antiviral agents against haemorrhagic fever viruses. Antivir Chem Chemother 19:41–47. https://doi.org/10.1177/095632020801900106
Daley SK, Cordell GA (2021) Alkaloids in Contemporary Drug Discovery to Meet Global Disease Needs. Molec 26:3800. https://doi.org/10.3390/molecules26133800
Wouters OJ, McKee M, Luyten J (2009) Estimated Research and Development Investment Needed to Bring a New Medicine to Market, 2009–2018. JAMA 323:844–853
Sun D, Gao W, Hu H, Zhou S (2022) Why 90% of clinical drug development fails and how to improve it? Acta Pharm Sinica B. https://doi.org/10.1016/j.apsb.2022.02.002
Paul SM, Mytelka DS, Dunwiddie CT, Persinger CC, Munos BH, Lindborg SR, Schacht AL (2010) How to improve R&D productivity: the pharmaceutical industry’s grand challenge. Nat RevDrug Discov 9:203–214. https://doi.org/10.1038/nrd3078
Tripathi SK, Muttineni R, Singh SK (2013) Extra precision docking, free energy calculation and molecular dynamics simulation studies of CDK2 inhibitors. J Theor Biol 334:87–100. https://doi.org/10.1016/j.jtbi.2013.05.014
Daina A, Michielin O, Zoete V (2017) SwissADME: a free web tool to evaluate pharmacokinetics, drug-likeness and medicinal chemistry friendliness of small molecules. Scient Rep 7:42717. https://doi.org/10.1038/srep42717
Becke AD (1993) Density-functional thermochemistry. III. The role of exact exchange. J Chem Phys 98:5648–5652. https://doi.org/10.1063/1.464913
Oyeneyin O, Ipinloju N, Ojo N, Akerele D (2021) Structural Modification of Ibuprofen as new NSAIDs via DFT, Molecular Docking and Pharmacokinetics Studies. Int J Adv Engin Pure Sci 33:614–626. https://doi.org/10.7240/jeps.928422
Omoboyowa DA, Iqbal MN, Balogun TA, Bodun DS, Fatoki JO, Oyeneyin OE (2022) Inhibitory potential of phytochemicals from Chromolaena odorata L. against apoptosis signal regulatory kinase 1: A computational model against colorectal cancer. Comput Toxic 23:100235. https://doi.org/10.1016/j.comtox.2022.100235
Nguyen DD, Gao K, Chen J, Wang R, Wei GW (2020) Unveiling the molecular mechanism of SARS-CoV-2 main protease inhibition from 137 crystal structures using algebraic topology and deep learning. Chem Sci 11:12036–12046. https://doi.org/10.1039/D0SC04641H
Shi S, Li J, Zhao X, Liu Q, Song SJ (2021) A comprehensive review: Biological activity, modification and synthetic methodologies of prenylated flavonoids. Phytochem 191:112895. https://doi.org/10.1016/j.phytochem.2021.112895
Botta B, Vitali A, Menendez P, Misiti D, Monache DG (2005) Prenylated Flavonoids: Pharmacology and Biotechnology. Curr Medic Chem 12:713–739. https://doi.org/10.2174/0929867053202241
Lipinski CA, Lombardo F, Dominy BW, Feeney PJ (2001) Experimental and computational approaches to estimate solubility and permeability in drug discovery and development settings. Adv Drug Deliv Rev 46:3–26. https://doi.org/10.1016/s0169-409x(00)00129-0
Muchmore SW, Edmunds JJ, Stewart KD, Hajduk PJ (2010) Cheminformatic Tools for Medicinal Chemists. J Medic Chem 53:4830–4841. https://doi.org/10.1021/jm100164z
Mälkiä A, Murtomäki L, Urtti A, Kontturi K (2004) Drug permeation in biomembranes: In vitro and in silico prediction and influence of physicochemical properties. Europ J Pharm Sci 23:13–47. https://doi.org/10.1016/j.ejps.2004.05.009
Stillhart C, Vučićević K, Augustijns P, Basit AW, Batchelor H, Flanagan TR, Gesquiere I, Greupink R, Keszthelyi D, Koskinen M, Madla CM, Matthys C, Miljuš G, Mooij MG, Parrott N, Ungell AL, de Wildt SN, Orlu M, Klein S, Müllertz A (2020) Impact of gastrointestinal physiology on drug absorption in special populations––An UNGAP review. Europ J Pharm Sci 147:105280. https://doi.org/10.1016/j.ejps.2020.105280
Geldenhuys WJ, Mohammad AS, Adkins CE, Lockman PR (2015) Molecular determinants of blood–brain barrier permeation. Th Deliv 6:961–971. https://doi.org/10.4155/tde.15.32
El-Shamy NT, Alkaoud AM, Hussein RK, Ibrahim MA, Alhamzani AG, Abou-Krisha MM (2022) DFT, ADMET and Molecular Docking Investigations for the Antimicrobial Activity of 6,6’-Diamino-1,1’,3,3’-tetramethyl-5,5’-(4-chlorobenzylidene)bis[pyrimidine-2,4(1H,3H)-dione]. Molec 27:620. https://doi.org/10.3390/molecules27030620. )
Ogunyemi BT, Oderinlo OO (2022) In-silico investigation of oxoaporphine alkaloids of Xylopia aethiopica against SARS-COV-2 main protease. AROC in Nat Prod Res 2:1–12. https://doi.org/10.53858/arocnpr02010112
Arivazhagan R, Sridevi C, Prakasam A (2021) Exploring molecular structure, spectral features, electronic properties and molecular docking of a novel biologically active heterocyclic compound 4-phenylthiosemicarbazide. J Molec Struct 1232:129956. https://doi.org/10.1016/j.molstruc.2021.129956
Oyeneyin OE, Ojo ND, Ipinloju N, James AC, Agbaffa EB (2022) Investigation of Corrosion Inhibition Potentials of Some Aminopyridine Schiff Bases Using Density Functional Theory and Monte Carlo Simulation. Chem Afr 5:319–332. https://doi.org/10.1007/s42250-021-00304-1
Kumar A, Sambandam S, Ramalingam A, Krishnamoorthy R, Arumugam D, Oyeneyin OE (2022) Synthesis, molecular docking of 3-(2-chloroethyl)-2,6-diphenylpiperidin-4-one: Hirshfeld surface, spectroscopic and DFT based analyses. J Mol Struct 1262:132993. https://doi.org/10.1016/j.molstruc.2022.132993
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
On behalf of all authors, the corresponding author states that there is no conflict of interest.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic Supplementary Material
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Oderinlo, O.O., Iwegbulam, C.G., Ekweli, O.A. et al. Acridone Alkaloids: In-Silico Investigation Against SARS-CoV-2 Main Protease. Chemistry Africa 5, 1441–1450 (2022). https://doi.org/10.1007/s42250-022-00440-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s42250-022-00440-2