Abstract
Background
Arginine-vasopressin (AVP) is a neuropeptide and provides learning and memory modulation. The AVP (4–5) dipeptide corresponds to the N-terminal fragment of the major vasopressin metabolite AVP (4–9), has a neuroprotective effect and used in the treatment of Alzheimer’s and Parkinson’s disease.
Methods
The main objective of the present study is to evaluate the molecular mechanism of AVP (4–5) dipeptide and to develop and synthesize chitosan nanoparticle formulation using modified version of ionic gelation method, to increase drug effectiveness. For peptide loaded chitosan nanoparticles, the synthesized experiment medium was simulated for the first time by molecular dynamics method and used to determine the stability of the peptide, and the binding mechanism to protein (HSP70) was also investigated by molecular docking calculations. A potential pharmacologically features of the peptide was also characterized by ADME (Absorption, Distribution, Metabolism and Excretion) analysis. The characterization, in vitro release study, encapsulation efficiency and loading capacity of the peptide loaded chitosan nanoparticles (CS NPs) were performed by Dynamic Light Scattering (DLS), UV–vis absorption (UV), Scanning Electron Microscopy (SEM), Fourier transform infrared (FT-IR) spectroscopy techniques. Additionally, in vitro cytotoxicity of the peptide on human neuroblastoma cells (SH-SY5Y) was examined with XTT assay and the statistical analysis was evaluated.
Results
The results showed that; hydrodynamic size, zeta potential and polydispersity index (PdI) of the peptide-loaded CS NPs were 167.6 nm, +13.2 mV, and 0.211, respectively. In vitro release study of the peptide-loaded CS NPs showed that 17.23% of the AVP (4–5)-NH2 peptide was released in the first day, while 61.13% of AVP (4–5)-NH2 peptide was released in the end of the 10th day. The encapsulation efficiency and loading capacity were 99% and 10%, respectively. According to the obtained results from XTT assay, toxicity on SHSY-5Y cells in the concentration from 0.01 μg/μL to 30 μg/μL were evaluated and no toxicity was observed. Also, neuroprotective effect was showed against H2O2 treatment.
Conclusion
The experimental medium of peptide-loaded chitosan nanoparticles was created for the first time with in silico system and the stability of the peptide in this medium was carried out by molecular dynamics studies. The binding sites of the peptide with the HSP70 protein were determined by molecular docking analysis. The size and morphology of the prepared NPs capable of crossing the blood-brain barrier (BBB) were monitored using DLS and SEM analyses, and the encapsulation efficiency and loading capacity were successfully performed with UV Analysis. In vitro release studies and in vitro cytotoxicity analysis on SHSY-5Y cell lines of the peptide were conducted for the first time.

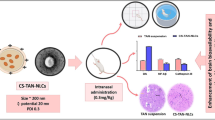
Grapical abstract










Similar content being viewed by others
Abbreviations
- AVP:
-
Arginine-vaspressin
- CS NPs:
-
Chitosan nanoparticles
- AD:
-
Alzheimer’s disease
- PD:
-
Parkinson’s disease
- NGF:
-
Nerve growth factor
- HSP:
-
Heat shock proteins
- MD:
-
Molecular Dynamics
- NVT:
-
Number of particles, Volume, and Temperature
- NPT:
-
Number of particles, Pressure, and Temperature
- Rg:
-
Gyration
- RMSD:
-
Root mean square deviation
- VMD:
-
Visual Molecular Dynamics
- ADME:
-
Absorption, Distribution, Metabolism and Excretion
- TED:
-
Total energy distribution
- PdI:
-
Polydispersity index
- FT-IR:
-
Fourier Transform Infrared
- SEM:
-
Scanning Electron Microscopy
- DLS:
-
Dynamic Light Scattering
- EE:
-
Encapsulation Efficiency
- LC:
-
Loading Capacity
- PBS:
-
Phosphate Buffered Saline
- XTT:
-
sodium 3,3′-[1(phenylamino)carbonyl]-3,4-tetrazolium]-3is(4-methoxy-6-nitro) benzene sulfonic acid hydrate
- DMSO:
-
Dimethyl Sulfoxide
References
Zenina TA, Gudasheva TA, Bukreyev YS, Seredenin SB. Neuroprotective effect of dipeptide AVP (4-5)-NH2 is associated with nerve growth factor and heat shock protein HSP70. Bull Exp Biol Med. 2007;144(4):543–5.
Evans CG, Wisén S, Gestwicki JE. Heat shock proteins 70 and 90 inhibit early stages of amyloid beta (1-42) aggregation in vitro. J Biol Chem. 2006.
Kreek MJ, Zhou Y, Levran O. Functions of arginine vasopressin and its receptors: importance of human molecular genetics studies in bidirectional translational research. Biol Psychiatry. 2011;70(6):502.
Wolkowitz OM, Rothschild AJ. Psychoneuroendocrinology: the scientific basis of clinical practice. American psychiatric pub. 2008.
Batla A, Phé V, De Min L, Panicker JN. Nocturia in Parkinson's disease: why does it occur and how to manage? Movement Disorders Clinical Practice. 2016;3(5):443–51.
Mayer MP, Bukau B. Hsp70 chaperones: cellular functions and molecular mechanism. Cell Mol Life Sci. 2005;62(6):670.
Witt SN. Hsp70 molecular chaperones and Parkinson's disease. Biopolymers: Original Research on Biomolecules. 2010;93(3):218–28.
Lu RC, Tan MS, Wang H, Xie AM, Yu JT, Tan L. Heat shock protein 70 in Alzheimer’s disease. Biomed Res Int. 2014.
Calzoni E, Cesaretti A, Polchi A, Di Michele A, Tancini B, Emiliani C. Biocompatible polymer nanoparticles for drug delivery applications in cancer and neurodegenerative disorder therapies. J Funct Biomater. 2019;10(1).
Zhao LM, Shi LE, Zhang ZL, Chen JM, Shi DD, Yang J, et al. Preparation and application of chitosan nanoparticles and nanofibers. Braz J Chem Eng. 2011;28(3):353–62.
Ghadi A, Mahjoub S, Tabandeh F, Talebnia F. Synthesis and optimization of chitosan nanoparticles: potential applications in nanomedicine and biomedical engineering. Caspian journal of internal medicine. 2014;5(3):156–61.
Landriscina A, Rosen J, Friedman AJ. Biodegradable chitosan nanoparticles in drug delivery for infectious disease. Nanomedicine. 2015;10(10):1609–19.
Wang X, Chi N, Tang X. Preparation of estradiol chitosan nanoparticles for improving nasal absorption and brain targeting. Eur J Pharm Biopharm. 2008;70(3):735–40.
Sadigh-Eteghad S, Talebi M, Farhoudi M, Mahmoudi J, Reyhani B. Effects of levodopa loaded chitosan nanoparticles on cell viability and caspase-3 expression in PC12 neural like cells. Neurosciences. 2013;18(3):281–3.
Frisch MJ, Trucks GW, Schlegel HB, Scuseria GE, Robb MA, Cheeseman JR, et al. Gaussian 16. Revision A. 2016;3.
Wang J, Wang W, Kollman PA, Case DA. Automatic atom type and bond type perception in molecular mechanical calculations. J Mol Graph Model. 2006;25(2):247–60.
Wang J, Wolf RM, Caldwell JW, Kollman PA, Case DA. Development and testing of a general amber force field. J Comput Chem. 2004;25(9):1157–74.
da Silva AWS, Vranken WF. ACPYPE-Antechamber python parser interface. BMC research notes. 2012;5(1):367.
Balajee R, Rajan MD. Molecular docking and simulation studies of farnesyl trasnferase with the potential inhibitor theflavin. Journal of Applied Pharmaceutical Science. 2011;1(8):141.
Abraham MJ, Murtola T, Schulz R, Páll S, Smith JC, Hess B, et al. GROMACS: high performance molecular simulations through multi-level parallelism from laptops to supercomputers. SoftwareX. 2015;1:19–25.
Hornak V, Abel R, Okur A, Strockbine B, Roitberg A, Simmerling C. Comparison of multiple Amber force fields and development of improved protein backbone parameters. Proteins: Structure, Function, and Bioinformatics. 2006;65(3):712–25.
Jorgensen WL, Chandrasekhar J, Madura JD, Impey RW, Klein ML. Comparison of simple potential functions for simulating liquid water. J Chem Phys. 1983;79(2):926–35.
Bussi G, Donadio D, Parrinello M. Canonical sampling through velocity rescaling. The Journal of Chemical Physics. 2007;126(1).
Parrinello M, Rahman A. Polymorphic transitions in single crystals: a new molecular dynamics method. J Appl Phys. 1981;52(12):7182–90.
Hess B, Bekker H, Berendsen HJ, Fraaije JG. LINCS: a linear constraint solver for molecular simulations. J Comput Chem. 1997;18(12):1463–72.
Turner PJ. XMGRACE, Version 5.1. 19. Center for Coastal and Land-Margin Research, Oregon Graduate Institute of Science and Technology, Beaverton, OR. 2005.
Humphrey W, Dalke A, Schulten K. VMD: visual molecular dynamics. J Mol Graph. 1996;14(1):33–8.
Sriram M, Osipiuk J, Freeman BC, Morimoto RI, Joachimiak A. Human Hsp70 molecular chaperone binds two calcium ions within the ATPase domain. Structure. 1997;5(3):403–14.
Bienert S, Waterhouse A, de Beer TA, Tauriello G, Studer G, Bordoli L, et al. The SWISS-MODEL repository—new features and functionality. Nucleic Acids Res. 2016;45(D1):D313–9.
Friesner RA, Murphy RB, Repasky MP, Frye LL, Greenwood JR, Halgren TA, et al. Extra precision glide: docking and scoring incorporating a model of hydrophobic enclosure for protein− ligand complexes. J Med Chem. 2006;49(21):6177–96.
Halgren TA, Murphy RB, Friesner RA, Beard HS, Frye LL, Pollard WT, et al. Glide: a new approach for rapid, accurate docking and scoring. 2. Enrichment factors in database screening. J Med Chem. 2004;47(7):1750–9.
Friesner RA, Banks JL, Murphy RB, Halgren TA, Klicic JJ, Mainz DT, et al. Glide: a new approach for rapid, accurate docking and scoring. 1. Method and assessment of docking accuracy. J Med Chem. 2004;47(7):1739–49.
Harder E, Damm W, Maple J, Wu C, Reboul M, Xiang JY, et al. OPLS3: a force field providing broad coverage of drug-like small molecules and proteins. J Chem Theory Comput. 2015;12(1):281–96.
Sastry GM, Adzhigirey M, Day T, Annabhimoju R, Sherman W. Protein and ligand preparation: parameters, protocols, and influence on virtual screening enrichments. J Comput Aided Mol Des. 2013;27(3):221–34.
Søndergaard CR, Olsson MH, Rostkowski M, Jensen JH. Improved treatment of ligands and coupling effects in empirical calculation and rationalization of p K a values. J Chem Theory Comput. 2011;7(7):2284–95.
Venkatesan A, Rambabu M, Jayanthi S. Febin Prabhu Dass J. Pharmacophore feature prediction and molecular docking approach to identify novel anti-HCV protease inhibitors. J Cell Biochem. 2018;119(1):960–6.
Divya K, Jisha MS. Chitosan nanoparticles preparation and applications. Environ Chem Lett. 2018;16(1):101–12.
Sullivan DJ, Cruz-Romero M, Collins T, Cummins E, Kerry JP, Morris MA. Synthesis of monodisperse chitosan nanoparticles. Food Hydrocoll. 2018;83:355–64.
Ahsan SM, Thomas M, Reddy KK, Sooraparaju SG, Asthana A, Bhatnagar I. Chitosan as biomaterial in drug delivery and tissue engineering. Int J Biol Macromol. 2018;110:97–109.
Kilinc YB, Akdeste ZM, Koc RC, Bagirova M, Allahverdiyev A. Synthesis and characterization of antigenic influenza a M2e protein peptide-poly (acrylic) acid bioconjugate and determination of toxicity in vitro. Bioengineered. 2014;5(6):357–62.
Erci F, Cakir-Koc R, Isildak I. Green synthesis of silver nanoparticles using Thymbra spicata L. var. spicata (zahter) aqueous leaf extract and evaluation of their morphology-dependent antibacterial and cytotoxic activity. Artificial Cells, Nanomedicine, and Biotechnology. 2018;46:150–8.
Liwo A, Tempczyk A, Oldziej S, Shenderovich MD, Hruby VJ, Talluri S, et al. Exploration of the conformational space of oxytocin and arginine-vasopressin using the electrostatically driven Monte Carlo and molecular dynamics methods. Biopolymers. 1996;38(2):157–75.
Schmidt JM, Ohlenschläger O, Rüterjans H, Grzonka Z, Kojro E, Pavo I, et al. Conformation of [8-arginine] vasopressin and V1 antagonists in dimethyl sulfoxide solution derived from two-dimensional NMR spectroscopy and molecular dynamics simulation. Eur J Biochem. 1991;201(2):355–71.
Lipinski CA, Lombardo F, Dominy BW, Feeney PJ. Experimental and computational approaches to estimate solubility and permeability in drug discovery and development settings. Adv Drug Deliv Rev. 1997;23(1–3):3–25.
Lipinski CA. Lead-and drug-like compounds: the rule-of-five revolution. Drug Discov Today Technol. 2004;1(4):337–41.
Leo A, Hansch C, Elkins D. Partition coefficients and their uses. Chem Rev. 1971;71(6):525–616.
Tihanyi K, Vastag M (Eds.). Solubility, delivery and ADME problems of drugs and drug-candidates. Bentham Science Publishers. 2011.
Veber DF, Johnson SR, Cheng HY, Smith BR, Ward KW, Kopple KD. Molecular properties that influence the oral bioavailability of drug candidates. J Med Chem. 2002;45(12):2615–23.
Johansen A, Hansen HD, Svarer C, Lehel S, Leth-Petersen S, Kristensen JL, et al. The importance of small polar radiometabolites in molecular neuroimaging: a PET study with [11C] Cimbi-36 labeled in two positions. J Cereb Blood Flow Metab. 2018;38(4):659–68.
Carpenter TS, Kirshner DA, Lau EY, Wong SE, Nilmeier JP, Lightstone FC. A method to predict blood-brain barrier permeability of drug-like compounds using molecular dynamics simulations. Biophys J. 2014;107(3):630–41.
Dressman JB, Lennernas H. Oral drug absorption: prediction and assessment: CRC Press; 2000.
Mohammadpour Dounighi N, Eskandari R, Avadi MR, Zolfagharian H. Mir Mohammad Sadeghi a, Rezayat M. preparation and in vitro characterization of chitosan nanoparticles containing Mesobuthus eupeus scorpion venom as an antigen delivery system. Journal of Venomous Animals and Toxins Including Tropical Diseases. 2012;18(1):44–52.
Anicuta SG, Dobre L, Stroescu M, Jipa I. Fourier transform infrared (FTIR) spectroscopy for characterization of antimicrobial films containing chitosan. Analele Universită Ńii din Oradea Fascicula: Ecotoxicologie, Zootehnie şi Tehnologii de Industrie Alimentară. 2010:1234–40.
Liu CG, Desai KGH, Chen XG, Park HJ. Preparation and characterization of nanoparticles containing trypsin based on hydrophobically modified chitosan. J Agric Food Chem. 2005;53(5):1728–33.
Negrea P, Caunii A, Sarac I, Butnariu M. The study of infrared spectrum of chitin and chitosan extract as potential sources of biomass. Digest Journal of Nanomaterials & Biostructures (DJNB). 2015;10(4).
Silva SM, Braga CR, Fook MV, Raposo CM, Carvalho LH, Canedo EL. Application of infrared spectroscopy to analysis of chitosan/clay nanocomposites. In Infrared Spectroscopy-Materials Science, Engineering and Technology. InTech. (2012).
Mazancová P, Némethová V, Treľová D, Kleščíková L, Lacík I, Rázga F. Dissociation of chitosan/tripolyphosphate complexes into separate components upon pH elevation. Carbohydr Polym. 2018;192:104–10.
PQS version 3.1, Parallel Quantum Solutions, 2013 Green Acres Road, Suite A Fayetteville, Arkansas, 72703 USA.
Balci K, Akyuz S. A vibrational spectroscopic investigation on benzocaine molecule. Vib Spectrosc. 2008;48(2):215–28.
Murdock RC, Braydich-Stolle L, Schrand AM, Schlager JJ, Hussain SM. Characterization of nanomaterial dispersion in solution prior to in vitro exposure using dynamic light scattering technique. Toxicol Sci. 2008;101(2):239–53.
Patil P, Bhoskar M. Optimization and evaluation of spray dried chitosan nanoparticles containing doxorubicin. Int J Curr Pharm Res. 2014;6(1):7–15.
Hu YL. Qi W, Han F, Shao JZ. Gao JQ Toxicity evaluation of biodegradable chitosan nanoparticles using a zebrafish embryo model International journal of nanomedicine. 2011;6:3351.
Tang ZX, Qian JQ, Shi LE. Preparation of chitosan nanoparticles as carrier for immobilized enzyme. Appl Biochem Biotechnol. 2007;136(1):77–96.
Jarudilokkul S, Tongthammachat A, Boonamnuayvittaya V. Preparation of chitosan nanoparticles for encapsulation and release of protein. Korean J Chem Eng. 2011;28(5):1247.
Wang JJ, Zeng ZW, Xiao RZ, Xie T, Zhou GL, Zhan XR, et al. Recent advances of chitosan nanoparticles as drug carriers. Int J Nanomedicine. 2011;6:765.
Dudhani AR, Kosaraju SL. Bioadhesive chitosan nanoparticles: preparation and characterization. Carbohydr Polym. 2010;81(2):243–51.
Huyck L, Ampe C, Van Troys M. The XTT cell proliferation assay applied to cell layers embedded in three-dimensional matrix. Assay and drug development technologies. 2012;10(4):382–92.
López-García J, Lehocký M, Humpolíček P, Sáha P. HaCaT keratinocytes response on antimicrobial atelocollagen substrates: extent of cytotoxicity, cell viability and proliferation. Journal of functional biomaterials. 2014;5(2):43–57.
Zhao D, Yu S, Sun B, Gao S, Guo S, Zhao K. Biomedical applications of chitosan and its derivative nanoparticles. Polymers. 2018;10(4):462.
Cho Y, Shi R, Borgens RB. Chitosan nanoparticle-based neuronal membrane sealing and neuroprotection following acrolein-induced cell injury. J Biol Eng. 2010;4(1):2.
Pangestuti R, Kim SK. Neuroprotective properties of chitosan and its derivatives. Marine Drugs. 2010;8(7):2117–28.
Acknowledgements
Authors are also very thankful to Rita Podzuna for allowing using the docking program with Schrödinger’s Small-Molecule Drug Discovery Suite. In this study, the infrastructure of Applied Nanotechnology and Antibody Production Laboratory established with TUBITAK support (project numbers: 115S132 and 117S097) was used. Authors would thank to TUBITAK for their support.
Availability of data and materials
Data sharing not applicable to this article as no datasets were generated or analysed during the current study.
Funding
This study was supported by the Research funds of Istanbul University [ONAP-2423].
Author information
Authors and Affiliations
Contributions
SG: Participated in the design of the study, carried out the FTIR, band component analysis study, molecular docking, and molecular dynamic simulation and drafted the manuscript. YBK and TZ: Participated in the design of the experimental study, (synthesize and characterize nanoparticles) drafted the manuscript. RK: Participated in the design of the experimental study (cytotoxicity studies) drafted the manuscript. BB and YK: Participated in the design of the molecular docking and molecular dynamic simulation. AO and SA: Responsible for the study design and gave final approval of the version to be published. All authors read and approved the final manuscript and provide financial and administrative support.
Corresponding author
Ethics declarations
Conflict of interests
On behalf of all authors, the corresponding author states that there is no conflict of interest.
Consent for publication
Not applicable.
Ethics approval and consent to participate
Not applicable.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Kecel-Gunduz, S., Budama-Kilinc, Y., Cakir-Koc, R. et al. In Silico design of AVP (4–5) peptide and synthesis, characterization and in vitro activity of chitosan nanoparticles. DARU J Pharm Sci 28, 139–157 (2020). https://doi.org/10.1007/s40199-019-00325-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s40199-019-00325-9