Abstract
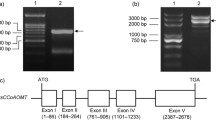
Three chalcone synthase (CHS) genes were isolated from Curcuma longa Linn. using TAIL-PCR. ClCHS1 and ClCHS2 were 1,460 and 1,407 bp in length, respectively, containing 1,191 bp open reading frame (ORF) that encodes 396 amino acids, whereas ClCHS3 was 1,394 bp in length containing 1,170 bp ORF that encodes 389 amino acids. The structure of all three genes comprise two exons and one intron which are consistent with the other CHS gene family. Southern blot analysis using a ClCHS conserved fragment revealed the ClCHS genes belong to a gene family. Phylogenetic analysis showed the three putative ClCHS proteins to be closely related to DCS (diketide-CoA synthase) protein, a product of CHS-like gene in C. longa, which condenses malonyl-CoA with feruloyl-CoA or coumaroyl-CoA as the substrate in curcuminoid synthesis. These results suggest that the interaction of substrate and enzyme between the three putative ClCHS proteins and DCS might be highly similar. Homology modeling and docking analysis were consistent, indicating that the same substrate (coumaroyl-CoA) can be used in the putative ClCHS1 and ClCHS2 proteins. However, the putative ClCHS3 protein seems to have seven amino acids deletion in a loop involved in the binding site formation, suggesting that the binding site with coumaroyl-CoA might be altered.





Similar content being viewed by others
Abbreviations
- AIC:
-
Akaike information criterion
- BAS:
-
Benzalacetone synthase
- CHS:
-
Chalcone synthase
- ClCHS:
-
Curcuma longa chalcone synthase
- CURS:
-
Curcumin synthase
- DCS:
-
Diketide-CoA synthase
- DDBJ:
-
DNA data bank of Japan
- NCBI:
-
National Center for Biotechnology Information
- PDB:
-
Protein data bank
- STS:
-
Stilbene synthase
- TAIL-PCR:
-
Thermal asymmetric interlaced polymerase chain reaction
- VPS:
-
Valerophenone synthase
- 2-PS:
-
2-pyrone synthase CHS
References
Abascal F, Zardoya R, Posada D (2005) ProtTest: selection of best-fit models of protein evolution. Bioinformatics 21:2104–2210
Agrawal GK, Pandey RN, Agrawal VP (1992) Isolation of DNA from Choerospondias asillaris leaves. Biotech Biodiv Lett 2:19–24
Arnold K, Bordoli L, Kopp J, Schwede T (2006) The SWISS-MODEL Workspace: a web-based environment for protein structure homology modeling. Bioinformatics 22:195–201
Chavalittumrong P, Jirawattanapong W (1992) Variation of active constituents of Curcuma domestica rhizomes at different ages. Thai J Pharm Sci 16:165–174
Clegg MT, Cummings MP, Durbin ML (1997) The evolution of plant nuclear genes. Proc Natl Acad Sci U S A 94:7791–7798
Davis TM, Shields ME, Zhang Q, Tombolato-Tezic D, Bennetzen JL, Pontaroli AC, Wang H, Yao Q, SanMiguel P, Folta KM (2010) An examination of targeted gene neighborhoods in strawberry. BMC Plant Biol 10:81. doi:10.1186/1471-2229-10-81
Delano WL (2008) The PyMOL Molecular Graphics System. DeLano Scientific LLC, Palo Alto
Eckermann S, Schröder G, Schmidt J, Strack D, Edrada RA, Helariutta Y, Elomaa P, Kotilainen M, Kilpeläinen I, Proksch P, Teeri TH, Schröder J (1998) New pathway to polyketide in plants. Nature 396:387–390
Ferrer JL, Jez JM, Bowman ME, Dixon RA, Noel JP (1999) Structure of chalcone synthase and the molecular basis of plant polyketide biosynthesis. Nat Struct Biol 8:775–784
Gasteiger E, Hoogland C, Gattiker A, Duvaud S, Wilkins MR, Appel RD, Bairoch A (2005) Protein identification and analysis tools on the ExPASy server. In: Walker JM (ed) The proteomics protocols handbook. Humana Press. pp 571–607
Han YY, Ming F, Wang JW, Wen JG, Ye MM, Shen DL (2006) Cloning and characterization of a novel chalcone synthase gene from Phalaenopsis hybrida orchid flowers. Russ J Plant Physiol 53:223–230
Hanhineva KJ, Kärenlampi SO (2007) Production of transgenic strawberries by temporary immersion bioreactor system and verification by TAIL-PCR. BMC Biotechnol 7:11–23
Helariutta Y, Kotilainen M, Elomaa P, Kalkkinen N, Bremer K, Teeri TH, Albert VA (1996) Duplication and functional divergence in the chalcone synthase gene family of Asteraceae: evolution with substrate change and catalytic simplification. Proc Natl Acad Sci U S A 93:9033–9038
Hiserodt R, Hartman TG, Ho CT, Rosen RT (1996) Characterization of powdered turmeric by liquid chromatography-mass spectrometry and gas chromatography-mass spectrometry. J Chromatogr A 740:51–63
Huelsenbeck JP, Ronquist F (2001) MRBAYES: Bayesian inference of phylogeny. Bioinformatics 17:754–755
Ito M, Ichinose Y, Kato H, Shiraishi T, Yamada T (1997) Molecular evolution and functional relevance of the chalcone synthase genes of pea. Mol Gen Genet 255:28–37
Jane SK, Rains J, Jones K (2006) Effect of curcumin on protein glycosylation, lipid peroxidation and oxygen radical generation in human red blood cells exposed to high glucose levels. Free Radic Biol Med 41:92–96
Jiang C, Schommer CK, Kim SY, Suh DY (2006) Cloning and characterization of chalcone synthase from the moss, Physcomitrella patens. Phytochemistry 67:2531–2540
Jiang C, Kim SY, Suh DY (2008) Divergence evolution of the thiolase superfamily and chalcone synthase family. Mol Phylogenet Evol 49:691–701
Katsuyama Y, Kita T, Horinouchi S (2009a) Identification and characterization of multiple curcumin synthases from the herb Curcuma longa. FEBS Lett 583:2799–2803
Katsuyama Y, Kita T, Funa N, Horinouchi S (2009b) Curcuminoid biosynthesis by two type III polyketide synthases in the herb Curcuma longa. J Biol Chem 284:11160–11170
Koduri PK, Gordon GS, Barker EI, Colpitts CC, Ashton NW, Suh DY (2010) Genome-wide analysis of the chalcone synthase superfamily genes of Physcomitrella patens. Plant Mol Biol 72:247–263
Kumar S, Tamura K, Nei M (2004) MEGA3: integrated software for molecular evolutionary genetics analysis and sequence alignment. Brief Bioinform 5:150–163
Laskowski R, Thornton J (1993) PROCHECK: a program to check the stereochemical quality of protein structures. J Appl Crystallogr 26:283–291
Liew CF, Goh CJ, Loh CS, Lim SH (1998) Cloning and characterization of full-length cDNA clones encoding chalcone synthase from the orchid Bromheadia finlaysoniana. Plant Physiol Biochem 36:647–6 56
Liu YG, Whittier RF (1995) Thermal asymmetric interlaced PCR: automatable amplification and sequencing of insert end fragments from PI and YAC clones for chromosome walking. Genomics 25:674–681
Morris GM, Huey R, Lindstrom W, Sanner MF, Belew RK, Goodsell DS, Olson AJ (2009) AutoDock4 and AutoDockTools4: automated docking with selective receptor flexibility. J Comput Chem 30:2785–2791
Okada Y, Ito K (2001) Cloning and analysis of valerophenone synthase gene expressed specifically in lupulin gland of hop (Humulus lupulus L.). Biosci Biotechnol Biochem 65:150–155
Pang Y, Shen G, Wu W, Liu X, Lin J, Tan F, Sun X, Tang K (2005) Characterization and expression of chalcone synthase gene from Ginkgo biloba. Plant Sci 168:1525–1531
Schöppner A, Kindl H (1984) Purification and properties of a stilbene synthase from induced cell suspension cultures of peanut. J Biol Chem 259:6806–6811
Shi M, Cai Q, Yao L, Mao Y, Ming Y, Ouyang G (2006) Antiproliferation and apoptosis induced by curcumin in human ovarian cancer cells. Cell Biol Int 30:221–226
Sommer H, Saedler H (1986) Structure of the chalcone synthase gene of Antirrhinum majus. Mol Gen Genet 202:429–434
Thaikert R, Paisooksantivatana Y (2009) Variation of total curcuminoids content, antioxidant activity and genetic diversity in turmeric (Curcuma Longa L.) collections. Kasetsart J (Nat Sci) 43:507–518
Trognitz F, Manosalva P, Gysin R, Niñio-Liu D, Simon R, del Herrera MR, Trognitz B, Ghislain M, Nelson R (2002) Plant defense genes associated with quantitative resistance to potato late blight in Solanum phureja x dihaploid S. tuberosum hybrids. Mol Plant Microbe Interact 15:587–597
Wu WW, Welsh MJ, Kaufman PB, Zhang HH (1997) Method in gene biotechnology. CRC Press, New York
Yang J, Huang J, Gu H, Zhong Y, Yang Z (2002) Duplication and adaptive evolution of the chalcone synthase genes of Dendranthema (Asteraceae). Mol Biol Evol 19:1752–1759
Acknowledgments
This work was financially supported by Kasetsart University Research and Development Institute (KURDI), Kasetsart University, Thailand. We also would like to thank Vichien Keeratinijakal, Kasetsart University, Thailand for providing plant materials.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Supplementary Table 1
Gene name and protein accession numbers of species used in the sequence alignment and phylogenetic tree reconstruction (DOC 83 kb)
Rights and permissions
About this article
Cite this article
Wannapinpong, S., Srikulnath, K., Thongpan, A. et al. Molecular cloning and characterization of the CHS gene family in turmeric (Curcuma longa Linn.). J. Plant Biochem. Biotechnol. 24, 25–33 (2015). https://doi.org/10.1007/s13562-013-0232-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13562-013-0232-8