Abstract
Introduction
The time required to reach clinical remission varies in patients with chronic urticaria (CU). The objective of this study is to develop a predictive model using a machine learning methodology to predict time to clinical remission for patients with CU.
Methods
Adults with ≥ 2 ICD-9/10 relevant CU diagnosis codes/CU-related treatment > 6 weeks apart were identified in the Optum deidentified electronic health record dataset (January 2007 to June 2019). Clinical remission was defined as ≥ 12 months without CU diagnosis/CU-related treatment. A random survival forest was used to predict time from diagnosis to clinical remission for each patient based on clinical and demographic features available at diagnosis. Model performance was assessed using concordance, which indicates the degree of agreement between observed and predicted time to remission. To characterize clinically relevant groups, features were summarized among cohorts that were defined based on quartiles of predicted time to remission.
Results
Among 112,443 patients, 73.5% reached clinical remission, with a median of 336 days from diagnosis. From 1876 initial features, 176 were retained in the final model, which predicted a median of 318 days to remission. The model showed good performance with a concordance of 0.62. Patients with predicted longer time to remission tended to be older with delayed CU diagnosis, and have more comorbidities, more laboratory tests, higher body mass index, and polypharmacy during the 12-month period before the first CU diagnosis.
Conclusions
Applying machine learning to real-world data enabled accurate prediction of time to clinical remission and identified multiple relevant demographic and clinical variables with predictive value. Ongoing work aims to further validate and integrate these findings into clinical applications for CU management.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Why carry out this study? | |
While chronic urticaria (CU) typically lasts for an average of 3–5 years, individual variability makes prediction of disease duration a challenge in clinical practice. | |
We developed a predictive machine learning model using real-world health records to predict time to clinical remission. | |
What was learned from the study? | |
Patients with predicted longer time to clinical remission tended to be older, and have delayed CU diagnosis and poorer health. | |
Using a predictive machine learning model to predict time to clinical remission based on readily available clinical and demographic characteristics may help individual patients and their clinicians with a clearer disease management plan. |
Introduction
Chronic urticaria (CU) is a skin condition that can be spontaneous (CSU) or inducible (CIndU), depending on the presence or absence of identifiable triggers. Although available treatments for CU can control symptoms in many patients, there is no published evidence that they influence the disease course [1,2,3]. Patients with inadequately controlled CU face a substantial burden as the disease impacts their health-related quality of life, including social and work living [4, 5].
Published evidence suggests that patients with CU experience a wide range of cumulative remission rates ranging between 10% and 38% at 1 year and 30% and 71% at 5 years [6], and reach clinical remission at varying times with average disease duration between 3 and 5 years in adults [4, 7,8,9]. Several demographic and clinical characteristics influencing the duration of active disease among patients with CU have been reported (e.g., age, gender, presence of angioedema, hypertension), although the data are limited and inconclusive for many reasons, such as variability in patient profiles, small sample sizes, and a lack of consensus on the definition of disease remission [8,9,10].
Given the large heterogeneity of published data on time to clinical remission and the variables that influence it among patients with CU, there is a current unmet need for accurate prediction of time to remission in clinical practice. Machine learning, including random survival forests, has become an increasingly useful method in healthcare research for this purpose, as it enables the inclusion of a broad set of variables (i.e., features) and excels at handling large and complex predictor spaces [11,12,13,14]. The objective of the present study is to develop a predictive model using a machine learning methodology to predict time to clinical remission with readily available patient demographic and clinical characteristics. In addition, the features identified by the model as those that may have predictive value in clinical practice were explored.
Methods
Data Source
This study used (under license) data from the Optum deidentified electronic health record (EHR) dataset (January 2007 to June 2019), which includes information on more than 102 million lives receiving care from more than 700 hospitals and 7000 clinics in the USA as part of an integrated delivery network (IDN). Clinical information, healthcare/insurance claims, and other medical administrative data were obtained from both inpatient and ambulatory EHR, practice management systems, and numerous other internal systems. The database includes information on demographics, medications prescribed and administered, immunizations, allergies, laboratory results (including microbiology), vital signs, and other observable measurements, clinical and inpatient stay administrative data, and coded diagnoses and procedures. The medical activities of patients are recorded and available between the first date of medical insurance coverage/medical activity and the last date of medical insurance coverage/medical activity or death. Data are deidentified and comply with the patient requirements of the Health Insurance Portability and Accountability Act (HIPAA).
Study Design and Sample Selection
A retrospective study design was implemented (Fig. 1) whereby adult patients with CU were initially identified based on the presence of an International Classification of Diseases 9th/10th edition (ICD-9/10) diagnosis code for urticaria (Supplementary Table E1, excluding secondary codes). Confirmation of the disease was required based on the presence of a second diagnosis of urticaria and/or angioneurotic edema (Supplementary Table E1), or a CU-related treatment (Supplementary Table E2) > 6 weeks apart. For every patient included in the study sample, the index date was defined as the date of the first urticaria diagnosis. Patients were included in the study if they had ≥ 12 months of available data before the index date (i.e., the baseline period) and ≥ 12 months of available data after the last CU diagnosis and/or CU-related treatment. The follow-up period was defined as the time from the index date to data end. In accordance with the definition of CU in the guidelines [4], patients who had ≤ 6 weeks of disease duration (i.e., between the index date and the last CU diagnosis and/or CU-related treatment) were excluded, as they were considered to have acute urticaria (Fig. 2).
Study design. CU chronic urticaria. (1) Data beginning was defined as the latest of either the first date of medical insurance coverage or the first date of any clinical activity (including visits, encounters, diagnoses, prescriptions, observations, laboratory tests, procedures, and medication administration). (2) Data end was defined as the earliest date among medical coverage end, last clinical activity (including visits, encounters, diagnoses, prescriptions, observations, laboratory tests, procedures, and medication administration), or death. (3) Patients with ≤ 6 weeks of disease duration (i.e., between the index date and the last CU diagnosis and/or CU-related treatment) were excluded, as they were considered to have acute urticaria
Sample selection. CU chronic urticaria. (1) A list of diagnosis codes for urticaria, CSU, and CIndU is available in Supplementary Table E1. (2) A list of urticaria treatments is available in Supplementary Table E2. (3) Data beginning was defined as the latest of either the first date of medical insurance coverage date or the first date of any clinical activity (including visits, encounters, diagnoses, prescriptions, observations, laboratory tests, procedures, and medication administration). (4) Data end was defined as the earliest date among medical coverage end, last clinical activity (including visits, encounters, diagnoses, prescriptions, observations, laboratory tests, procedures, and medication administration), or death. (5) A patient was considered to reach clinical remission if the patient had a period of ≥ 12 months free of CU diagnosis (Supplementary Table E1)/CU-related treatment (Supplementary Table E2)
Outcome Variable Definition: A Proxy for Clinical Remission
The time from the index date to the first occurrence of clinical remission was identified for each patient in the full study sample. Clinical remission was defined as a period of ≥ 12 months free of CU (i.e., no CU diagnosis and/or any CU-related treatment), with the remission date being the start date of this period [15]. Time to clinical remission was right-censored at the date of 12 months prior to data end if no period ≥ 12 months free of CU was observed, since this is the last date at which remission status is known (for any given timepoint, ≥ 12 months of data after the timepoint are required to determine whether a patient reached remission).
Statistical Analyses
Kaplan–Meier (KM) analysis was used to assess the observed time to clinical remission. A random survival forest was used to predict the time to clinical remission and to estimate a survival function for each patient. Predicted clinical remission was defined as the point in time at which the probability of remission, as predicted by the model, reached ≥ 50%.
Predictive Machine Learning Model
A predictive machine learning model using a random survival forest was developed for the analysis of the right-censored survival data [16,17,18,19,20]. A random survival forest is a nonparametric machine learning method that generates a forest of decision trees using bootstrap samples and randomly selected features. By contrast to traditional time-to-event outcome analysis (e.g., Cox regression), random survival forests do not impose a limit on the number of preselected predictors, allow a nonlinear relationship between predictors and outcome, and do not rely on strong assumptions of proportional hazard function (i.e., constant hazard rate ratio between two patients over time). In addition, the importance of predictors can be calculated on the basis of the model to identify the factors that are closely related to time to clinical remission for patients with CU. Based on the majority vote of predictions across individual trees, the model predicts the survival function for each patient (i.e., the probability that a patient will continue to have CU as a function of time from the index date). The random survival forest model was constructed through prespecified options (i.e., number of trees, splitting rule), hyperparameter optimization (i.e., depth of a tree, minimum node size), and feature reduction (i.e., reducing the count of initially included variables without compromising accuracy). The number of trees typically determines the size of the random forest; as model performance tends to increase with the number of trees, its value was set to a number above which the performance of the model stabilized. The default splitting rule (log rank) was used for decision tree construction. Hyperparameters including the depth of trees (i.e., the maximum levels a tree can have) and minimum node size (i.e., the minimum number of observations in a node) were optimized through a random search [21]. Feature reduction was applied to filter out nonpredictive variables. Full details of the random search and the feature reduction process are outlined in Fig. 3. The “ranger” package (version 0.12) in R (version 3.6.2) [22] was used to conduct the random survival forest analysis.
Construction of the random survival forest. (1) Statistical significance of variable importance was not evaluated for three reasons. First, with a sample size of 112,443, statistical significance is likely to be found for many variables that are not clinically relevant [23]. Second, even if a substantial number of variables are completely unimportant (true zeros, in statistical terms), with so many variables included in the model (176), a number of variables will appear to be significant at conventional levels [24]. Third, statistical significance testing imposes a substantial computation burden
Model Performance Metrics
The predictive model’s performance was assessed statistically using concordance of (the degree of agreement between) the observed and predicted time to clinical remission. Concordance was measured by Harrell’s c-index calculated on the out-of-bag (OOB) sample to evaluate the performance of the model as a whole [18, 25, 26]. Unlike other measures of survival performance, the c-index was selected as it does not depend on a single fixed time for evaluation and specifically accounts for censoring. To calculate the c-index, the observed time to clinical remission and the time to clinical remission predicted by the model was selected for a pair of patients. If the patient with the shorter predicted time to clinical remission was observed to remit earlier, that patient pair was deemed concordant. If the patient with the longer predicted time to clinical remission was observed to remit earlier, that patient pair was deemed discordant. If this relationship could not be determined (due to censoring), the patient pair was ignored. This exercise was then repeated for all possible patient pairs, and the c-index represents the fraction of concordant pairs.
In biomedical applications, concordance for survival function-type outcomes typically falls between 0.6 and 0.75 [27].
Variables and Clinical Interpretation of Results
All variables available during the baseline period and 6 weeks post-index were included in the predictive model as features and comprised sociodemographic and clinical data such as age at index date, body mass index (BMI), smoking status, comorbidities, time from first recorded CU-related symptom to first CU diagnosis, specialist visits, diagnoses, laboratory tests, prescriptions for medication, and procedures.
Given that the study developed a predictive model, statistical significance of variable importance was not evaluated.
To facilitate the clinical interpretation of the features identified to have the most predictive value by the model, four cohorts of patients were created based on quartiles of predicted time to clinical remission as follows: cohort 1: < 254 days; cohort 2: 254–299 days; cohort 3: 300–398 days; cohort 4: ≥ 399 days. Summary statistics for the features were reported across these cohorts.
A partial dependence plot [28] of time to clinical remission as a function of selected continuous predictor(s) was produced whereby, in the absence of other differences in clinical and demographic variables, the effect of the selected variable as an independent predictor on the predicted time to clinical remission can be elucidated.
Results
A total of 112,443 adult patients with CU met the inclusion criteria (Fig. 2). The mean age at the index date for the full study sample was 47.7 years, and 77.0% were female (Table 1). The mean disease duration was 34.3 months. Among patients with available data on CU-related symptoms, the mean time from first recorded CU-related symptom to first CU diagnosis was 80.0 days.
Among the full study sample, 82,680 patients (73.5%) reached clinical remission during a median follow-up of 43.8 months (Table 1). The observed median time to clinical remission by KM analysis was 336 days.
Model Development and Features Identified to Have the Most Predictive Value
Out of 1876 features per patient that were initially included in the model, 176 with predictive value were retained in the final model after feature reduction (Fig. 4). The model predicted the median time to clinical remission to be 318 days with a concordance of 0.62, as measured by the OOB c-index.
Clinically relevant features identified by the model to have the most predictive value for the time to clinical remission included age at index, time from first recorded CU-related symptom to first CU diagnosis, BMI, smoking status, asthma, allergic rhinitis, chronic pulmonary disease, and prior medication use.
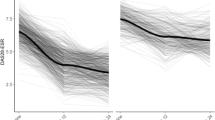
Patients in the four cohorts established by quartiles of median predicted time to clinical remission were predicted to reach remission within a median of 233 days in cohort 1 (N = 28,049), 275 days in cohort 2 (N = 28,001), 330 days in cohort 3 (N = 28,281), and 722 days in cohort 4 (N = 28,112; Table 2 and Fig. 5).
Patients with a longer predicted time to clinical remission (i.e., cohort 4) tended to be older at diagnosis relative to those with shorter predicted time to remission (i.e., cohorts 1–3) with a median age of 40 years in cohort 1 and 51 years in cohort 4 (Table 2). Relative to the other cohorts, patients in cohort 4 appeared to have longer time from first CU-related symptom to first CU diagnosis (Table 2), higher prevalence of comorbidities (e.g., asthma, allergic rhinitis; Fig. 6a ), polypharmacy [e.g., leukotriene receptor antagonists (LTRAs), antihistamines, proton pump inhibitors (PPIs); Fig. 6b], and higher prevalence of laboratory tests [e.g., creatinine, estimated glomerular filtration rate [eGFR]; Fig. 6c].
A partial dependence plot of time to clinical remission as a function of age revealed that, in the absence of other differences in clinical and demographic variables, older age at the index date, as an independent predictor, was associated with longer time to clinical remission (Supplementary Fig. E1).
Clinical characteristics identified by the model to have predictive value, reported by cohort. ** Variables with two asterisks are among the 20 most important predictors. * Variables with one asterisk are among the 50 most important predictors. † Variables with a cross were also important predictors when assessed during the 6-week period after the index date
Discussion
A predictive machine learning model was developed to predict time to clinical remission among patients with CU using real-world medical data with a high degree of agreement between the observed and predicted time to clinical remission. Older age at diagnosis, longer time from onset to CU diagnosis, increased prevalence of comorbidities, particularly asthma and allergic rhinitis, higher BMI, smoking, polypharmacy, and increased laboratory workup prior to or at the time of diagnosis appeared to relate to longer predicted time to clinical remission.
The observed median time to clinical remission was 336 days, a value that is generally consistent with prior published studies that used a similar definition of clinical remission (i.e., ≥ 12 months free of CU diagnosis/CU-related treatment). Of note, there was variation in the median time to clinical remission across published studies depending on differences in the study objectives, sample selection, methodology, and data sources. One retrospective, single-hospital chart review study of 337 patients in Thailand showed that the median disease duration for CSU was 390 days [29]. In another study using the South Korean National Health Insurance Service–National Sample Cohort, 1893 patients with incident CU experienced remission within a median of 330 days [30]. Although the estimates reported in our study generally agree with these findings despite the different methodology, the variability in the observed time to clinical remission (±309.1 days) for our large overall sample (N = 112,443) emphasizes the high variability of real-world time to CU remission in this patient population, further complicating individualized patient care in clinical practice.
Some of the clinical variables identified by our model as important in predicting time to clinical remission have been previously reported in literature (e.g., older age at diagnosis, asthma, allergic rhinitis, and high BMI) [6, 10, 29, 31,32,33,34,35]. However, there were other variables not identified by our model as important in predicting time to clinical remission but that, nonetheless, have been reported to be associated with time to clinical remission in other studies using different designs and target populations [34, 35]. For example, in two studies of Korean populations, Kim et al. found that female patients were more likely to enter remission, while Ye et al. found the inverse [34, 35]. In addition, the current study identified a partial dependence of time to clinical remission as a function of age, whereby older age predicted longer time to clinical remission. While this aligns with the findings of Kim et al., Ye found that age showed no influence on clinical remission. Nonetheless, due to the large heterogeneity of study designs, there is a lack of consensus on the association between CU and this complex set of characteristics [8, 9, 36], some of which can influence disease duration. To date, no studies have attempted to identify the predictive value of variables measured prior to CU diagnosis on time to clinical remission in the real world nor developed a model to predict time to clinical remission to support clinicians in their disease management plan. Through the use of a predictive machine learning model, the present study addresses this gap in knowledge, contributes valuable insights, and has the advantage of identifying and measuring these patient characteristics during the period before the first CU diagnosis to predict time to remission in clinical practice.
The variables identified as important by the predictive model and the general trends observed among the cohorts suggest that poor general health status may contribute to longer time to clinical remission. For example, patients with longer predicted time to clinical remission appeared to have higher prevalence of comorbidities, such as depression and diabetes, before their first CU diagnosis, as well as the presence and result of several laboratory tests, which are not typically performed without an underlying cause for concern and tend to be targeted, if necessary [37]. Additionally, the use of certain medications prior to first CU diagnosis, including LTRAs, antihistamines, and PPIs, was identified as important by the predictive model. Of note, omalizumab use during the year before or 6 weeks after first CU diagnosis was observed in less than 1% of the full study sample and was not identified by our model as important in predicting time to clinical remission. The potential for the characteristics identified as important by the predictive model, such as comorbidities, polypharmacy, and increased laboratory workup, to help predict time to clinical remission raise intriguing questions regarding the pathogenesis of CU. Autoallergy and autoimmunity have been proposed as pathogenetic mechanisms in urticaria [38], and it could be speculated that patients with asthma and seasonal and perennial allergic rhinitis may be more likely to have autoallergic disease. It is also possible that the general proinflammatory state associated with smoking and higher BMI [39,40,41,42] could be indicative of the underlying mechanisms related to urticaria.
The predictive machine learning model developed in this study is a novel statistical approach used to predict time to clinical remission and to identify features with predictive value in patients for whom remission status is already known (i.e., comparing a predicted outcome with an observed outcome in the same population). Unlike standard approaches typically used to identify the predictive value of a number of preselected features, the machine learning methodology applied in this study allowed for the evaluation of a large number of features without imposing strong assumptions on the form of the relationship between the outcome and its predictors (i.e., the relationship can be highly nonlinear), making it possible to identify features with predictive value that may not be obvious in clinical practice. Therefore, the trained model could assist clinicians in their efforts to predict time to clinical remission of CU using readily accessible patient information in real-world settings. It could also enable clinicians to gain a better understanding of the possible disease course and expected timeline for individual patients with CU who are currently in their care based on prespecified clinical and demographic characteristics. Furthermore, this information could help guide the development of an interactive tool for disease management.
Limitations
This study is subject to certain limitations, including those inherent to retrospective studies using EHR data, such as limited generalizability outside commercially insured US populations, as well as coding errors and omissions. Due to the absence of specific ICD-9/10 codes for CU, the study cohort was identified based on the presence of ≥ 2 ICD-9/10 diagnostic codes that relate to the disease, including secondary codes, such as allergic urticaria or angioneurotic edema. Therefore, it is possible that some cases might have been misclassified as cases with CU or that others could have been missed. There is no consensus on the definition of clinical remission for CU, so a period of ≥ 12 months free of CU diagnosis/CU-related treatment was adopted for this study based on input from clinical experts and some published studies [29, 34]. This definition of clinical remission also does not consider recurrences after the 12 months free of CU diagnosis/CU-related treatment, which is an avenue for further research. Measures of disease severity [e.g., Urticaria Activity Score (UAS7)] were not available in the EHR database. Therefore, future work is also warranted to investigate the relationship between CU severity and time to clinical remission.
Conclusions
This is the first study to successfully leverage a predictive machine learning methodology not only to identify important variables but also to predict time to clinical remission in CU using real-world medical data. While the model stands to be validated with additional data sources, the time to clinical remission reported was similar to other published studies conducted in other countries [16, 17]. Multiple relevant factors were identified that had predictive power for longer time to clinical remission, including older age at diagnosis, longer time to diagnosis, higher prevalence of comorbidities, particularly asthma and allergic rhinitis, higher BMI, smoking, polypharmacy, and increased laboratory tests in the year preceding or during the first 6 weeks following the first CU diagnosis. The important features identified through the predictive machine learning model (1) reveal that there might be complex interactions among variables, which are difficult to identify and interpret through clinical judgment alone, (2) allow clinicians to better understand the course and duration of CU, and support management decisions pertaining to the disease, and (3) provide insights into factors involved in the pathogenesis of the condition. Ongoing work aims to translate this predictive model into real-world application(s) that could support clinicians in better assessing patients’ time to clinical remission based on collectable information and support better disease management.
References
Vestergaard C, Deleuran M. Chronic spontaneous urticaria: latest developments in aetiology, diagnosis and therapy. Ther Adv Chronic Dis. 2015;6(6):304–13. https://doi.org/10.1177/2040622315603951.
Dressler C, Werner RN, Eisert L, et al. Chronic inducible urticaria: a systematic review of treatment options. J Allergy Clin Immunol. 2018;141(5):1726–34. https://doi.org/10.1016/j.jaci.2018.01.031 (PubMed PMID: 29438771).
Lavery WJ, Bernstein JA. Allergy and Asthma: acute and chronic urticaria. Mahmoudi M (ed). Springer, Cham. 2019.
Zuberbier T, Aberer W, Asero R, et al. The EAACI/GA(2)LEN/EDF/WAO guideline for the definition, classification, diagnosis and management of urticaria. Allergy. 2018;73(7):1393–414. https://doi.org/10.1111/all.13397 (PubMed PMID: 29336054).
Fricke J, Avila G, Keller T, et al. Prevalence of chronic urticaria in children and adults across the globe: systematic review with meta-analysis. Allergy. 2020;75(2):423–32. https://doi.org/10.1111/all.14037 (PubMed PMID: 31494963).
Balp MM, Halliday AC, Severin T, et al. Clinical remission of chronic spontaneous urticaria (CSU): a targeted literature review. Dermatol Ther (Heidelb). 2021. https://doi.org/10.1007/s13555-021-00641-6 (PubMed PMID: 34807372).
Maurer M, Weller K, Bindslev-Jensen C, et al. Unmet clinical needs in chronic spontaneous urticaria. A GA(2)LEN task force report. Allergy. 2011;66(3):317–30. https://doi.org/10.1111/j.1398-9995.2010.02496.x. (PubMed PMID: 21083565).
Toubi E, Kessel A, Avshovich N, et al. Clinical and laboratory parameters in predicting chronic urticaria duration: a prospective study of 139 patients. Allergy. 2004;59(8):869–73. https://doi.org/10.1111/j.1398-9995.2004.00473.x (PubMed PMID: 15230821).
Nebiolo F, Bergia R, Bommarito L, et al. Effect of arterial hypertension on chronic urticaria duration. Ann Allergy Asthma Immunol. 2009;103(5):407–10. https://doi.org/10.1016/S1081-1206(10)60360-2 (PubMed PMID: 19927539).
Stepaniuk P, Kan M, Kanani A. Natural history, prognostic factors and patient perceived response to treatment in chronic spontaneous urticaria. Allergy Asthma Clin Immunol. 2020;16:63. https://doi.org/10.1186/s13223-020-00459-5.PubMedPMID:32834828;PubMedCentralPMCID:PMCPMC7371813.
Kononenko I. Machine learning for medical diagnosis: history, state of the art and perspective. Artif Intell Med. 2001;23(1):89–109. https://doi.org/10.1016/s0933-3657(01)00077-x (PubMed PMID: 11470218).
Beam AL, Kohane IS. Big data and machine learning in health care. JAMA. 2018;319(13):1317–8. https://doi.org/10.1001/jama.2017.18391 (PubMed PMID: 29532063).
Ghassemi M, Naumann T, Schulam P, et al. A review of challenges and opportunities in machine learning for health. AMIA Jt Summits Transl Sci Proc. 2020;2020:191–200. (PubMed PMID: 32477638; PubMed Central PMCID: PMCPMC7233077).
Padula WV, Kreif N, Vanness DJ, et al. Machine Learning Methods in Health Economics and Outcomes Research-The PALISADE Checklist: a good practices report of an ISPOR Task Force. Value Health. 2022;25(7):1063–80. https://doi.org/10.1016/j.jval.2022.03.022 (PubMed PMID: 35779937).
Bernstein JA, Lang DM, Khan DA, et al. The diagnosis and management of acute and chronic urticaria: 2014 update. J Allergy Clin Immunol. 2014;133(5):1270–7. https://doi.org/10.1016/j.jaci.2014.02.036 (PubMed PMID: 24766875).
Adham D, Abbasgholizadeh N, Abazari M. Prognostic factors for survival in patients with gastric cancer using a random survival forest. Asian Pac J Cancer Prev. 2017;18(1):129–34. https://doi.org/10.22034/APJCP.2017.18.1.129.PubMedPMID:28240020;eng.
Hsich E, Gorodeski EZ, Blackstone EH, et al. Identifying important risk factors for survival in patient with systolic heart failure using random survival forests. Circ Cardiovasc Qual Outcomes. 2011;4(1):39–45. https://doi.org/10.1161/CIRCOUTCOMES.110.939371.PubMedPMID:21098782;PubMedCentralPMCID:PMCPMC3991475.
Ishwaran H, Kogalur UB, Blackstone EH, et al. Random survival forests. Ann Appl Stat. 2008;2(3):841–60. https://doi.org/10.1214/08-AOAS169.
Scheffner I, Gietzelt M, Abeling T, et al. Patient survival after kidney transplantation: Important role of graft-sustaining factors as determined by predictive modeling using random survival forest analysis. Transplantation. 2020;104(5):1095–107. https://doi.org/10.1097/TP.0000000000002922 (PubMed PMID: 31403555).
Steele AJ, Denaxas SC, Shah AD, et al. Machine learning models in electronic health records can outperform conventional survival models for predicting patient mortality in coronary artery disease. PLoS One. 2018;13(8): e0202344. https://doi.org/10.1371/journal.pone.0202344.
Bergstra J, Bengio Y. Random search for hyper-parameter optimization. J Mach Learn Res. 2012;13(2).
Wright MN, Ziegler A. ranger: a fast implementation of random forests for high dimensional data in C++ and R. J Stat Softw. 2017;77(1):1–17.
Lantz B. The large sample size fallacy. Scand J Caring Sci. 2013;27(2):487–92. https://doi.org/10.1111/j.1471-6712.2012.01052.x (PubMed PMID: 22862286).
Bland JM, Altman DG. Multiple significance tests: the Bonferroni method. BMJ. 1995;310(6973):170. https://doi.org/10.1136/bmj.310.6973.170.PubMedPMID:7833759;PubMedCentralPMCID:PMCPMC2548561.
Harrell FE Jr, Califf RM, Pryor DB, et al. Evaluating the yield of medical tests. JAMA. 1982;247(18):2543–6 (PubMed PMID: 7069920).
Alabdallah A, Ohlsson M, Pashami S, et al. The Concordance Index decomposition—a measure for a deeper understanding of survival prediction models. arXiv preprint arXiv:220300144. 2022.
Schmid M, Wright MN, Ziegler A. On the use of Harrell's C for clinical risk prediction via random survival forests. Expert Syst Appl. 2016;63(C):450–459. https://doi.org/10.1016/j.eswa.2016.07.018.
Friedman JH. Greedy function approximation: a gradient boosting machine. Ann Stat. 2001:1189–1232.
Kulthanan K, Jiamton S, Thumpimukvatana N, et al. Chronic idiopathic urticaria: prevalence and clinical course. J Dermatol. 2007;34(5):294–301. https://doi.org/10.1111/j.1346-8138.2007.00276.x (PubMed PMID: 17408437).
Eun SJ, Lee JY, Kim DY, et al. Natural course of new-onset urticaria: results of a 10-year follow-up, nationwide, population-based study. Allergol Int. 2019;68(1):52–8. https://doi.org/10.1016/j.alit.2018.05.011 (PubMed PMID: 29945815).
Olze H, Zuberbier T. Comorbidities between nose and skin allergy. Curr Opin Allergy Clin Immunol. 2011;11(5):457–63. https://doi.org/10.1097/ACI.0b013e32834a9764 (PubMed PMID: 21822129).
Vadasz Z, Kessel A, Hershko AY, et al. Seasonal exacerbation of asthma is frequently associated with recurrent episodes of acute urticaria. Int Arch Allergy Immunol. 2016;169(4):263–6. https://doi.org/10.1159/000446183 (PubMed PMID: 27240837).
Magen E, Chikovani T, Waitman DA, et al. Factors related to omalizumab resistance in chronic spontaneous urticaria. Allergy Asthma Proc. 2019;40(4):273–8. https://doi.org/10.2500/aap.2019.40.4230 (PubMed PMID: 31262381).
Kim YS, Park SH, Han K, et al. Clinical course of chronic spontaneous urticaria in the korean adult population. Allergy Asthma Immunol Res. 2018;10(1):83–7. https://doi.org/10.4168/aair.2018.10.1.83.PubMedPMID:29178681;PubMedCentralPMCID:PMCPMC5705488.
Ye YM, Yoon J, Woo SD, et al. Clustering the clinical course of chronic urticaria using a longitudinal database: Effects on urticaria remission. Allergy Asthma Immunol Res. 2021;13(3):390–403. https://doi.org/10.4168/aair.2021.13.3.390.PubMedPMID:33733635;PubMedCentralPMCID:PMCPMC7984955.
Fok JS, Kolkhir P, Church MK, et al. Predictors of treatment response in chronic spontaneous urticaria. Allergy. 2021;76(10):2965–81. https://doi.org/10.1111/all.14757 (PubMed PMID: 33539587).
Schaefer P. Acute and chronic urticaria: Evaluation and treatment. Am Fam Physician. 2017;95(11):717–24 (PubMed PMID: 28671445).
Asero R, Tedeschi A, Marzano AV, et al. Chronic urticaria: a focus on pathogenesis. F1000Res. 2017;6:1095. https://doi.org/10.12688/f1000research.11546.1. (PubMed PMID: 28751972; PubMed Central PMCID: PMCPMC5506533).
Elisia I, Lam V, Cho B, et al. The effect of smoking on chronic inflammation, immune function and blood cell composition. Sci Rep. 2020;10(1):19480. https://doi.org/10.1038/s41598-020-76556-7.PubMedPMID:33173057;PubMedCentralPMCID:PMCPMC7655856.
Lee J, Taneja V, Vassallo R. Cigarette smoking and inflammation: cellular and molecular mechanisms. J Dent Res. 2012;91(2):142–9. https://doi.org/10.1177/0022034511421200.PubMedPMID:21876032;PubMedCentralPMCID:PMCPMC3261116.
Shalom G, Magen E, Babaev M, et al. Chronic urticaria and the metabolic syndrome: a cross-sectional community-based study of 11 261 patients. J Eur Acad Dermatol Venereol. 2018;32(2):276–81. https://doi.org/10.1111/jdv.14548 (PubMed PMID: 28846167).
Bansal CJ, Bansal AS. Stress, pseudoallergens, autoimmunity, infection and inflammation in chronic spontaneous urticaria. Allergy Asthma Clin Immunol. 2019;15:56. https://doi.org/10.1186/s13223-019-0372-z.PubMedPMID:31528163;PubMedCentralPMCID:PMCPMC6737621.
Acknowledgements
Funding
This study and the journal’s Rapid Service Fee were funded by Novartis Pharma AG, Switzerland.
Author Contributions
Irina Pivneva, James Signorovitch, Jimmy Royer, Yawen Liang, Tom Cornwall, and Jutong Pan made substantial contributions to study conception and design, and data collection, analysis, and interpretation. Maria-Magdalena Balp, Yvonne Geissbühler, Thomas Severin, Serge Smeets, Andrii Danyliv, and Sarah Jane McKenna contributed to study conception and design, and data analysis and interpretation. Alexander M. Marsland and Weily Soong contributed to study conception and design, and data interpretation. All authors revised the manuscript critically for important intellectual content, provided final approval of the version to be published, and agree to be accountable for all aspects of the work in ensuring that questions related to the accuracy or integrity of any part of the work are appropriately investigated and resolved.
Disclosures
Irina Pivneva, James Signorovitch, Jimmy Royer, Tom Cornwall, and Jutong Pan are employees of Analysis Group, Inc., a consulting company that has provided paid consulting services to Novartis Pharma AG, Switzerland. At the time of the study, Yawen Liang was an employee of Analysis Group, Inc., which received payment from Novartis Pharma AG, Switzerland, for the conduct of this study; Yawen Liang's contributions to this study reflect work done prior to her joining Amazon. Maria-Magdalena Balp, Yvonne Geissbühler, Thomas Severin, Serge Smeets, and Andrii Danyliv are Novartis Pharma AG, Switzerland, employees and Sarah Jane McKenna is a Novartis Ireland employee and have no further conflicts of interest to disclose. Alexander M. Marsland has received grants/ educational grants/ consulting fees from Almirall, Galderma, Lilly, Novartis, Roche and UCB Pharma. Weily Soong has been an advisor and/or clinical investigator and/or received speaker’s honoraria and/or received consulting fee and/or grants and/or participated as a clinical investigator for/from the following companies: AbbVie, Allakos, Amgen, AstraZeneca, Genentech, GlaxoSmithKline, LEO, Novartis, Optinose, Pfizer, Regeneron, Sanofi and Teva.
Compliance with Ethics Guidelines
This study used data (used under license) from the Optum® de-identified Electronic Health Record (EHR) dataset (01/2007–06/2019). Data used in this study are de-identified and comply with the patient requirements of the Health Insurance Portability and Accountability Act (HIPAA); therefore, no institutional review board approval was required.
Data Availability
The data that support the findings of this study are available from Optum® de-identified Electronic Health Record. Restrictions apply to the availability of these data, which were used under license for this study.
Author information
Authors and Affiliations
Corresponding author
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial 4.0 International License, which permits any non-commercial use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc/4.0/.
About this article
Cite this article
Pivneva, I., Balp, MM., Geissbühler, Y. et al. Predicting Clinical Remission of Chronic Urticaria Using Random Survival Forests: Machine Learning Applied to Real-World Data. Dermatol Ther (Heidelb) 12, 2747–2763 (2022). https://doi.org/10.1007/s13555-022-00827-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13555-022-00827-6