Abstract
Background
Cancer is increasingly recognized as a metabolic disease, with evidence suggesting that oxidative phosphorylation (OXPHOS) plays a significant role in the progression of numerous cancer cells. OXPHOS not only provides sufficient energy for tumor tissue survival but also regulates conditions for tumor proliferation, invasion, and metastasis. Alterations in OXPHOS can also impair the immune function of immune cells in the tumor microenvironment, leading to immune evasion. Therefore, investigating the relationship between OXPHOS and immune escape is crucial in cancer-related research. This review aims to summarize the effects of transcriptional, mitochondrial genetic, metabolic regulation, and mitochondrial dynamics on OXPHOS in different cancers. Additionally, it highlights the role of OXPHOS in immune escape by affecting various immune cells. Finally, it concludes with an overview of recent advances in antitumor strategies targeting both immune and metabolic processes and proposes promising therapeutic targets by analyzing the limitations of current targeted drugs.
Conclusions
The metabolic shift towards OXPHOS contributes significantly to tumor proliferation, progression, metastasis, immune escape, and poor prognosis. A thorough investigation of concrete mechanisms of OXPHOS regulation in different types of tumors and the combination usage of OXPHOS-targeted drugs with existing immunotherapies could potentially uncover new therapeutic targets for future antitumor therapies.



Similar content being viewed by others
Data availability
Not applicable.
References
L.A. Broadfield, A.A. Pane, A. Talebi, J.V. Swinnen, S.M. Fendt, Lipid metabolism in cancer: new perspectives and emerging mechanisms. Dev. Cell. 56, 1363–1393 (2021). https://doi.org/10.1016/j.devcel.2021.04.013
U.E. Martinez-Outschoorn, M. Peiris-Pagés, R.G. Pestell, F. Sotgia, M.P. Lisanti, Cancer metabolism: a therapeutic perspective. Nat. Rev. Clin. Oncol. 14, 11–31 (2017). https://doi.org/10.1038/nrclinonc.2016.60
M. Reina-Campos, J. Moscat, M. Diaz-Meco, Metabolism shapes the tumor microenvironment. Curr. Opin. Cell. Biol. 48, 47–53 (2017). https://doi.org/10.1016/j.ceb.2017.05.006
L. Xia, L. Oyang, J. Lin, S. Tan, Y. Han, N. Wu, P. Yi, L. Tang, Q. Pan, S. Rao, J. Liang, Y. Tang, M. Su, X. Luo, Y. Yang, Y. Shi, H. Wang, Y. Zhou, Q. Liao, The cancer metabolic reprogramming and immune response. Mol. Cancer 20, 28 (2021). https://doi.org/10.1186/s12943-021-01316-8
H. Xu, S. Zhou, Q. Tang, H. Xia, F. Bi, Cholesterol metabolism: new functions and therapeutic approaches in cancer. Biochim. Biophys. Acta Rev. Cancer 1874, 188394 (2020). https://doi.org/10.1016/j.bbcan.2020.188394
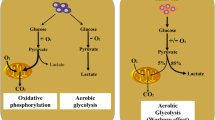
O. Warburg, On the origin of cancer cells. Science 123, 309–314 (1956). https://doi.org/10.1126/science.123.3191.309
O. Warburg, On the metabolism of cancer cells. Naturwissenschaften 12, 1131–1137 (1924). https://doi.org/10.1007/bf01504608
D. Johar, A.O. Elmehrath, R.M. Khalil, M.H. Elberry, S. Zaky, S.A. Shalabi, L.H. Bernstein, Protein networks linking Warburg and reverse Warburg effects to cancer cell metabolism. BioFactors 47, 713–728 (2021). https://doi.org/10.1002/biof.1768
S. Pavlides, D. Whitaker-Menezes, R. Castello-Cros, N. Flomenberg, A.K. Witkiewicz, P.G. Frank, M.C. Casimiro, C. Wang, P. Fortina, S. Addya, R.G. Pestell, U.E. Martinez-Outschoorn, F. Sotgia, M.P. Lisanti, The reverse Warburg effect: aerobic glycolysis in cancer associated fibroblasts and the tumor stroma. Cell. Cycle 8, 3984–4001 (2009). https://doi.org/10.4161/cc.8.23.10238
P.E. Porporato, N. Filigheddu, J.M.B. Pedro, G. Kroemer, L. Galluzzi, Mitochondrial metabolism and cancer. Cell. Res. 28, 265–280 (2018). https://doi.org/10.1038/cr.2017.155
W.T. Wang, W.L. Jin, X. Li, Intercellular communication in the tumour microecosystem: mediators and therapeutic approaches for hepatocellular carcinoma. Biochim. Biophys. Acta Mol. Basis Dis. 1868, 166528 (2022). https://doi.org/10.1016/j.bbadis.2022.166528
K.E. Allison, B.L. Coomber, B.W. Bridle, Metabolic reprogramming in the tumour microenvironment: a hallmark shared by cancer cells and T lymphocytes. Immunology 152, 175–184 (2017). https://doi.org/10.1111/imm.12777
X. Li, M. Wenes, P. Romero, S.C. Huang, S.M. Fendt, P.C. Ho, Navigating metabolic pathways to enhance antitumour immunity and immunotherapy. Nat. Rev. Clin. Oncol. 16, 425–441 (2019). https://doi.org/10.1038/s41571-019-0203-7
G. Andrejeva, J.C. Rathmell, Similarities and distinctions of Cancer and Immune metabolism in inflammation and tumors. Cell. Metab. 26, 49–70 (2017). https://doi.org/10.1016/j.cmet.2017.06.004
Z. Wu, M. Zuo, L. Zeng, K. Cui, B. Liu, C. Yan, L. Chen, J. Dong, F. Shangguan, W. Hu, H. He, B. Lu, Z. Song, OMA1 reprograms metabolism under hypoxia to promote colorectal cancer development. EMBO Rep. 22, e50827 (2021). https://doi.org/10.15252/embr.202050827
L. Liu, X. Zhang, H. Ding, X. Liu, D. Cao, Y. Liu, J. Liu, C. Lin, N. Zhang, G. Wang, J. Hou, B. Huang, Y. Zhang, J. Lu, Arginine and lysine methylation of MRPS23 promotes breast cancer metastasis through regulating OXPHOS. Oncogene 40, 3548–3563 (2021). https://doi.org/10.1038/s41388-021-01785-7
H.J. Kim, P. Maiti, A. Barrientos, Mitochondrial ribosomes in cancer. Semin. Cancer Biol 47, 67–81 (2017). https://doi.org/10.1016/j.semcancer.2017.04.004
V.R. Fantin, J. St-Pierre, P. Leder, Attenuation of LDH-A expression uncovers a link between glycolysis, mitochondrial physiology, and tumor maintenance. Cancer Cell. 9, 425–434 (2006). https://doi.org/10.1016/j.ccr.2006.04.023
R. Moreno-Sánchez, S. Rodríguez-Enríquez, A. Marín-Hernández, E. Saavedra, Energy metabolism in tumor cells. FEBS J. 274, 1393–1418 (2007). https://doi.org/10.1111/j.1742-4658.2007.05686.x
M. Wu, A. Neilson, A.L. Swift, R. Moran, J. Tamagnine, D. Parslow, S. Armistead, K. Lemire, J. Orrell, J. Teich, S. Chomicz, D.A. Ferrick, Multiparameter metabolic analysis reveals a close link between attenuated mitochondrial bioenergetic function and enhanced glycolysis dependency in human tumor cells. Am. J. Physiol. Cell. Physiol. 292, C125–C136 (2007). https://doi.org/10.1152/ajpcell.00247.2006
C.H. Chao, C.Y. Wang, C.H. Wang, T.W. Chen, H.Y. Hsu, H.W. Huang, C.W. Li, R.T. Mai, Mutant p53 attenuates oxidative phosphorylation and facilitates Cancer Stemness through Downregulating miR-200c-PCK2 Axis in basal-like breast Cancer. Mol. Cancer Res. 19, 1900–1916 (2021). https://doi.org/10.1158/1541-7786.Mcr-21-0098
A. Cruz-Bermúdez, R. Laza-Briviesca, R.J. Vicente-Blanco, A. García-Grande, M.J. Coronado, S. Laine-Menéndez, C. Alfaro, J.C. Sanchez, F. Franco, V. Calvo, A. Romero, P. Martin-Acosta, C. Salas, J.M. Garcia, M. Provencio, Cancer-associated fibroblasts modify lung cancer metabolism involving ROS and TGF-β signaling. Free Radic. Biol. Med 130, 163–173 (2019). https://doi.org/10.1016/j.freeradbiomed.2018.10.450
E. Reznik, M.L. Miller, Y. Şenbabaoğlu, N. Riaz, J. Sarungbam, S.K. Tickoo, H.A. Al-Ahmadie, W. Lee, V.E. Seshan, A.A. Hakimi, C. Sander, Mitochondrial DNA copy number variation across human cancers. Elife 5, (2016). https://doi.org/10.7554/eLife.10769
E. Zacksenhaus, M. Shrestha, J.C. Liu, I. Vorobieva, P.E.D. Chung, Y. Ju, U. Nir, Z. Jiang, Mitochondrial OXPHOS Induced by RB1 Deficiency in breast Cancer: implications for anabolic metabolism, stemness, and Metastasis. Trends Cancer 3, 768–779 (2017). https://doi.org/10.1016/j.trecan.2017.09.002
M. Bajzikova, J. Kovarova, A.R. Coelho, S. Boukalova, S. Oh, K. Rohlenova, D. Svec, S. Hubackova, B. Endaya, K. Judasova, A. Bezawork-Geleta, K. Kluckova, L. Chatre, R. Zobalova, A. Novakova, K. Vanova, Z. Ezrova, G.J. Maghzal, S. Magalhaes Novais, M. Olsinova, L. Krobova, Y.J. An, E. Davidova, Z. Nahacka, M. Sobol, T. Cunha-Oliveira, C. Sandoval-Acuña, H. Strnad, T. Zhang, T. Huynh, T.L. Serafim, P. Hozak, V.A. Sardao, W.J.H. Koopman, M. Ricchetti, P.J. Oliveira, F. Kolar, M. Kubista, J. Truksa, K. Dvorakova-Hortova, K. Pacak, R. Gurlich, R. Stocker, Y. Zhou, M.V. Berridge, S. Park, L. Dong, J. Rohlena, J. Neuzil, Reactivation of Dihydroorotate dehydrogenase-driven pyrimidine biosynthesis restores Tumor Growth of respiration-deficient Cancer cells. Cell. Metab. 29, 399–416.e310 (2019). https://doi.org/10.1016/j.cmet.2018.10.014
S. Rao, L. Mondragón, B. Pranjic, T. Hanada, G. Stoll, T. Köcher, P. Zhang, A. Jais, A. Lercher, A. Bergthaler, D. Schramek, K. Haigh, V. Sica, M. Leduc, N. Modjtahedi, T.P. Pai, M. Onji, I. Uribesalgo, R. Hanada, I. Kozieradzki, R. Koglgruber, S.J. Cronin, Z. She, F. Quehenberger, H. Popper, L. Kenner, J.J. Haigh, O. Kepp, M. Rak, K. Cai, G. Kroemer, J.M. Penninger, AIF-regulated oxidative phosphorylation supports lung cancer development. Cell. Res. 29, 579–591 (2019). https://doi.org/10.1038/s41422-019-0181-4
C. Gao, Y. Shen, F. Jin, Y. Miao, X. Qiu, Cancer Stem cells in small cell Lung Cancer Cell Line H446: higher dependency on oxidative phosphorylation and mitochondrial substrate-level phosphorylation than Non-Stem Cancer cells. PLoS One 11, e0154576 (2016). https://doi.org/10.1371/journal.pone.0154576
C. Wu, Y. Liu, W. Liu, T. Zou, S. Lu, C. Zhu, L. He, J. Chen, L. Fang, L. Zou, P. Wang, L. Fan, H. Wang, H. You, J. Chen, J.Y. Fang, C. Jiang, Y. Shi, NNMT-DNMT1 Axis is Essential for Maintaining Cancer Cell Sensitivity to Oxidative Phosphorylation Inhibition. Advanced science (Weinheim, Baden-Wurttemberg, Germany) 10, e2202642 (2022). https://doi.org/10.1002/advs.202202642
K. Ishikawa, K. Takenaga, M. Akimoto, N. Koshikawa, A. Yamaguchi, H. Imanishi, K. Nakada, Y. Honma, J. Hayashi, ROS-generating mitochondrial DNA mutations can regulate tumor cell metastasis. Science 320, 661–664 (2008). https://doi.org/10.1126/science.1156906
P. Sancho, E. Burgos-Ramos, A. Tavera, T. Bou Kheir, P. Jagust, M. Schoenhals, D. Barneda, K. Sellers, R. Campos-Olivas, O. Graña, C.R. Viera, M. Yuneva, B. Sainz Jr., C. Heeschen, MYC/PGC-1α balance determines the metabolic phenotype and plasticity of pancreatic Cancer stem cells. Cell. Metab. 22, 590–605 (2015). https://doi.org/10.1016/j.cmet.2015.08.015
S. Alcalá, P. Sancho, P. Martinelli, D. Navarro, C. Pedrero, L. Martín-Hijano, S. Valle, J. Earl, M. Rodríguez-Serrano, L. Ruiz-Cañas, K. Rojas, A. Carrato, L. García-Bermejo, M. Fernández-Moreno, P.C. Hermann, B. Sainz, Jr., ISG15 and ISGylation is required for pancreatic cancer stem cell mitophagy and metabolic plasticity. Nat. Commun. 11, 2682 (2020). https://doi.org/10.1038/s41467-020-16395-2
R.K. Nimmakayala, S. Rauth, R. Chirravuri Venkata, S. Marimuthu, P. Nallasamy, R. Vengoji, S.M. Lele, S. Rachagani, K. Mallya, M.P. Malafa, M.P. Ponnusamy, S.K. Batra, PGC1α-Mediated metabolic reprogramming drives the stemness of pancreatic precursor lesions. Clin. Cancer Res. 27, 5415–5429 (2021). https://doi.org/10.1158/1078-0432.Ccr-20-5020
K.W. Evans, E. Yuca, S.S. Scott, M. Zhao, N. Paez Arango, C.X. Cruz Pico, T. Saridogan, M. Shariati, C.A. Class, C.A. Bristow, C.P. Vellano, X. Zheng, A.M. Gonzalez-Angulo, X. Su, C. Tapia, K. Chen, A. Akcakanat, B. Lim, D. Tripathy, T.A. Yap, M.E.D. Francesco, G.F. Draetta, P. Jones, T.P. Heffernan, J.R. Marszalek, and F. Meric-Bernstam, oxidative phosphorylation is a metabolic vulnerability in chemotherapy-resistant triple-negative breast Cancer. Cancer Res. 81, 5572–5581 (2021). https://doi.org/10.1158/0008-5472.Can-20-3242
Y. Hu, W. Xu, H. Zeng, Z. He, X. Lu, D. Zuo, G. Qin, W. Chen, OXPHOS-dependent metabolic reprogramming prompts metastatic potential of breast cancer cells under osteogenic differentiation. Br. J. Cancer 123, 1644–1655 (2020). https://doi.org/10.1038/s41416-020-01040-y
K.M. Lee, J.M. Giltnane, J.M. Balko, L.J. Schwarz, A.L. Guerrero-Zotano, K.E. Hutchinson, M.J. Nixon, M.V. Estrada, V. Sánchez, M.E. Sanders, T. Lee, H. Gómez, A. Lluch, J.A. Pérez-Fidalgo, M.M. Wolf, G. Andrejeva, J.C. Rathmell, S.W. Fesik, C.L. Arteaga, MYC and MCL1 cooperatively promote chemotherapy-resistant breast Cancer stem cells via regulation of mitochondrial oxidative phosphorylation. Cell. Metab. 26, 633–647.e637 (2017). https://doi.org/10.1016/j.cmet.2017.09.009
N. Schömel, L. Gruber, S.J. Alexopoulos, S. Trautmann, E.M. Olzomer, F.L. Byrne, K.L. Hoehn, R. Gurke, D. Thomas, N. Ferreirós, G. Geisslinger, M.S. Wegner, UGCG overexpression leads to increased glycolysis and increased oxidative phosphorylation of breast cancer cells. Sci. Rep. 10, 8182 (2020). https://doi.org/10.1038/s41598-020-65182-y
T. Li, J. Han, L. Jia, X. Hu, L. Chen, Y. Wang, PKM2 coordinates glycolysis with mitochondrial fusion and oxidative phosphorylation. Protein Cell. 10, 583–594 (2019). https://doi.org/10.1007/s13238-019-0618-z
C.L. Chen, S.C. Hsu, T.Y. Chung, C.Y. Chu, H.J. Wang, P.W. Hsiao, S.D. Yeh, D.K. Ann, Y. Yen, H.J. Kung, Arginine is an epigenetic regulator targeting TEAD4 to modulate OXPHOS in prostate cancer cells. Nat. Commun. 12, 2398 (2021). https://doi.org/10.1038/s41467-021-22652-9
D.B. Rivadeneira, M.C. Caino, J.H. Seo, A. Angelin, D.C. Wallace, L.R. Languino, D.C. Altieri, Survivin promotes oxidative phosphorylation, subcellular mitochondrial repositioning, and tumor cell invasion. Sci. Signal. 8, ra80 (2015). https://doi.org/10.1126/scisignal.aab1624
Y. Yang, J. He, B. Zhang, Z. Zhang, G. Jia, S. Liu, T. Wu, X. He, N. Wang, SLC25A1 promotes tumor growth and survival by reprogramming energy metabolism in colorectal cancer. Cell. Death Dis. 12, 1108 (2021). https://doi.org/10.1038/s41419-021-04411-2
C. Ge, Y. Wang, Y. Feng, S. Wang, K. Zhang, X. Xu, Z. Zhang, Y. Zhao, Y. Wang, L. Gao, F. Dai, S. Xie, C. Wang, Suppression of oxidative phosphorylation and IDH2 sensitizes colorectal cancer to a naphthalimide derivative and mitoxantrone. Cancer Lett. 519, 30–45 (2021). https://doi.org/10.1016/j.canlet.2021.06.015
J. Zhao, Y. Wang, Y. Wang, J. Gao, H. Yang, X. Wu, H. Li, Transcription Factor FXR Activates DHRS9 to Inhibit the Cell Oxidative Phosphorylation and Suppress Colon Cancer Progression. Anal. Cell. Pathol. (Amst.) 2022, 8275574 (2022). https://doi.org/10.1155/2022/8275574
N.M. Anderson, X. Qin, J.M. Finan, A. Lam, J. Athoe, R. Missiaen, N. Skuli, A. Kennedy, A.S. Saini, T. Tao, S. Zhu, I. Nissim, A.T. Look, G. Qing, M.C. Simon, H. Feng, Metabolic enzyme DLST promotes Tumor Aggression and reveals a vulnerability to OXPHOS Inhibition in High-Risk Neuroblastoma. Cancer Res. 81, 4417–4430 (2021). https://doi.org/10.1158/0008-5472.Can-20-2153
C. Raggi, M.L. Taddei, E. Sacco, N. Navari, M. Correnti, B. Piombanti, M. Pastore, C. Campani, E. Pranzini, J. Iorio, G. Lori, T. Lottini, C. Peano, J. Cibella, M. Lewinska, J.B. Andersen, L. di Tommaso, L. Viganò, G. Di Maira, S. Madiai, M. Ramazzotti, I. Orlandi, A. Arcangeli, P. Chiarugi, F. Marra, Mitochondrial oxidative metabolism contributes to a cancer stem cell phenotype in cholangiocarcinoma. J. Hepatol. 74, 1373–1385 (2021). https://doi.org/10.1016/j.jhep.2020.12.031
A. Vyas, R.A. Harbison, D.L. Faden, M. Kubik, D. Palmer, Q. Zhang, H.U. Osmanbeyoglu, K. Kiselyov, E. Méndez, U. Duvvuri, Recurrent human papillomavirus-related Head and Neck Cancer undergoes metabolic reprogramming and is driven by oxidative phosphorylation. Clin. Cancer Res. 27, 6250–6264 (2021). https://doi.org/10.1158/1078-0432.Ccr-20-4789
X. Zhang, Y. Dong, M. Zhao, L. Ding, X. Yang, Y. Jing, Y. Song, S. Chen, Q. Hu, Y. Ni, ITGB2-mediated metabolic switch in CAFs promotes OSCC proliferation by oxidation of NADH in mitochondrial oxidative phosphorylation system. Theranostics 10, 12044–12059 (2020). https://doi.org/10.7150/thno.47901
G.M. Fischer, A. Jalali, D.A. Kircher, W.C. Lee, J.L. McQuade, L.E. Haydu, A.Y. Joon, A. Reuben, M.P. de Macedo, F.C.L. Carapeto, C. Yang, A. Srivastava, C.R. Ambati, A. Sreekumar, C.W. Hudgens, B. Knighton, W. Deng, S.D. Ferguson, H.A. Tawbi, I.C. Glitza, J.E. Gershenwald, Y.N. Vashisht Gopal, P. Hwu, J.T. Huse, J.A. Wargo, P.A. Futreal, N. Putluri, A.J. Lazar, R.J. DeBerardinis, J.R. Marszalek, J. Zhang, S.L. Holmen, M.T. Tetzlaff, M.A. Davies, Molecular Profiling reveals Unique Immune and metabolic features of Melanoma Brain Metastases. Cancer Discov 9, 628–645 (2019). https://doi.org/10.1158/2159-8290.Cd-18-1489
Y. Yang, G. Zhang, F. Guo, Q. Li, H. Luo, Y. Shu, Y. Shen, J. Gan, L. Xu, H. Yang, Mitochondrial UQCC3 modulates Hypoxia Adaptation by orchestrating OXPHOS and glycolysis in Hepatocellular Carcinoma. Cell. Rep. 33, 108340 (2020). https://doi.org/10.1016/j.celrep.2020.108340
J.L. Tan, F. Li, J.Z. Yeo, K.J. Yong, M.A. Bassal, G.H. Ng, M.Y. Lee, C.Y. Leong, H.K. Tan, C.S. Wu, B.H. Liu, T.H. Chan, Z.H. Tan, Y.S. Chan, S. Wang, Z.H. Lim, T.B. Toh, L. Hooi, K.N. Low, S. Ma, N.R. Kong, A.J. Stein, Y. Wu, M.T. Thangavelu, A. Suzuki, G. Periyasamy, J.M. Asara, Y.Y. Dan, G.K. Bonney, E.K. Chow, G.D. Lu, H.H. Ng, Y. Kanagasundaram, S.B. Ng, W.L. Tam, D.G. Tenen, and L. Chai, New High-Throughput Screening identifies Compounds that reduce viability specifically in Liver Cancer cells that express high levels of SALL4 by inhibiting oxidative phosphorylation. Gastroenterology 157, 1615–1629.e1617 (2019). https://doi.org/10.1053/j.gastro.2019.08.022
T. La, S. Chen, T. Guo, X.H. Zhao, L. Teng, D. Li, M. Carnell, Y.Y. Zhang, Y.C. Feng, N. Cole, A.C. Brown, D. Zhang, Q. Dong, J.Y. Wang, H. Cao, T. Liu, R.F. Thorne, F.M. Shao, X.D. Zhang, L. Jin, Visualization of endogenous p27 and Ki67 reveals the importance of a c-Myc-driven metabolic switch in promoting survival of quiescent cancer cells. Theranostics 11, 9605–9622 (2021). https://doi.org/10.7150/thno.63763
A. Salhi, A.C. Jordan, I.I. Bochaca, A. Izsak, F. Darvishian, Y. Houvras, K.M. Giles, I. Osman, Oxidative Phosphorylation Promotes Primary Melanoma Invasion. Am. J. Pathol 190, 1108–1117 (2020). https://doi.org/10.1016/j.ajpath.2020.01.012
G. Gentric, Y. Kieffer, V. Mieulet, O. Goundiam, C. Bonneau, F. Nemati, I. Hurbain, G. Raposo, T. Popova, M.H. Stern, V. Lallemand-Breitenbach, S. Müller, T. Cañeque, R. Rodriguez, A. Vincent-Salomon, H. de Thé, R. Rossignol, F. Mechta-Grigoriou, PML-Regulated mitochondrial metabolism enhances Chemosensitivity in Human ovarian cancers. Cell. Metab. 29, 156–173.e110 (2019). https://doi.org/10.1016/j.cmet.2018.09.002
S. Sriramkumar, R. Sood, T.D. Huntington, A.H. Ghobashi, T.T. Vuong, T.X. Metcalfe, W. Wang, K.P. Nephew, and H.M. O’Hagan, Platinum-induced mitochondrial OXPHOS contributes to cancer stem cell enrichment in ovarian cancer. J. Transl Med. 20, 246 (2022). https://doi.org/10.1186/s12967-022-03447-y
Y. Huang, Y. Du, Y. Zheng, C. Wen, H. Zou, J. Huang, H. Zhou, H. Zhao, L. Wu, Ct-OATP1B3 promotes high-grade serous ovarian cancer metastasis by regulation of fatty acid beta-oxidation and oxidative phosphorylation. Cell. Death Dis. 13, 556 (2022). https://doi.org/10.1038/s41419-022-05014-1
H.J. Choi, Y.L. Jhe, J. Kim, J.Y. Lim, J.E. Lee, M.K. Shin, J.H. Cheong, FoxM1-dependent and fatty acid oxidation-mediated ROS modulation is a cell-intrinsic drug resistance mechanism in cancer stem-like cells. Redox Biol. 36, 101589 (2020). https://doi.org/10.1016/j.redox.2020.101589
K. Birkenmeier, S. Dröse, I. Wittig, R. Winkelmann, V. Käfer, C. Döring, S. Hartmann, T. Wenz, A.S. Reichert, U. Brandt, M.L. Hansmann, Hodgkin and Reed-Sternberg cells of classical Hodgkin lymphoma are highly dependent on oxidative phosphorylation. Int. J. Cancer 138, 2231–2246 (2016). https://doi.org/10.1002/ijc.29934
E.D. Lagadinou, A. Sach, K. Callahan, R.M. Rossi, S.J. Neering, M. Minhajuddin, J.M. Ashton, S. Pei, V. Grose, K.M. O’Dwyer, J.L. Liesveld, P.S. Brookes, M.W. Becker, C.T. Jordan, BCL-2 inhibition targets oxidative phosphorylation and selectively eradicates quiescent human leukemia stem cells. Cell. Stem Cell. 12, 329–341 (2013). https://doi.org/10.1016/j.stem.2012.12.013
G. Petrella, G. Ciufolini, R. Vago, D.O. Cicero, The interplay between oxidative phosphorylation and glycolysis as a potential marker of bladder Cancer progression. Int. J. Mol. Sci. 21, (2020). https://doi.org/10.3390/ijms21218107
L. Huang, Y. Xie, W. Han, S. Jiang, L. Zeng, Oxidative Phosphorylation-Related Signature Participates in Cancer Development, and PTPRG Overexpression Suppresses the Cancer Progression in Clear Cell Renal Cell Carcinoma. J Immunol Res 2022, 8300187 (2022). https://doi.org/10.1155/2022/8300187
J. van den Ameele, A.H. Brand, Neural stem cell temporal patterning and brain tumour growth rely on oxidative phosphorylation. Elife 8, (2019). https://doi.org/10.7554/eLife.47887
C.L. Chen, C.Y. Lin, H.J. Kung, Targeting mitochondrial OXPHOS and their Regulatory signals in prostate cancers. Int. J. Mol. Sci. 22, (2021). https://doi.org/10.3390/ijms222413435
P.J. Fernandez-Marcos, J. Auwerx, Regulation of PGC-1α, a nodal regulator of mitochondrial biogenesis. Am. J. Clin. Nutr. 93, 884s–890 (2011). https://doi.org/10.3945/ajcn.110.001917
R.P. Kumar, S. Ray, P. Home, B. Saha, B. Bhattacharya, H.M. Wilkins, H. Chavan, A. Ganguly, J. Milano-Foster, A. Paul, P. Krishnamurthy, R.H. Swerdlow, S. Paul, Regulation of energy metabolism during early mammalian development: TEAD4 controls mitochondrial transcription. Development 145, (2018). https://doi.org/10.1242/dev.162644
J. Liang, R. Cao, X. Wang, Y. Zhang, P. Wang, H. Gao, C. Li, F. Yang, R. Zeng, P. Wei, D. Li, W. Li, W. Yang, Mitochondrial PKM2 regulates oxidative stress-induced apoptosis by stabilizing Bcl2. Cell. Res. 27, 329–351 (2017). https://doi.org/10.1038/cr.2016.159
C.L. Jones, B.M. Stevens, A. D’Alessandro, J.A. Reisz, R. Culp-Hill, T. Nemkov, S. Pei, N. Khan, B. Adane, H. Ye, A. Krug, D. Reinhold, C. Smith, J. DeGregori, D.A. Pollyea, C.T. Jordan, Inhibition of amino acid metabolism selectively targets human leukemia stem cells. Cancer Cell. 34, 724–740.e724 (2018). https://doi.org/10.1016/j.ccell.2018.10.005
P. Kaur, S. Nagar, M. Bhagwat, M. Uddin, Y. Zhu, I. Vancurova, A. Vancura, Activated heme synthesis regulates glycolysis and oxidative metabolism in breast and ovarian cancer cells. PLoS One 16, e0260400 (2021). https://doi.org/10.1371/journal.pone.0260400
Y. Sugiyama, Y. Hagiya, M. Nakajima, M. Ishizuka, T. Tanaka, S. Ogura, The heme precursor 5-aminolevulinic acid disrupts the Warburg effect in tumor cells and induces caspase-dependent apoptosis. Oncol. Rep. 31, 1282–1286 (2014). https://doi.org/10.3892/or.2013.2945
R. Filograna, M. Mennuni, D. Alsina, N.G. Larsson, Mitochondrial DNA copy number in human disease: the more the better? FEBS Lett. 595, 976–1002 (2021). https://doi.org/10.1002/1873-3468.14021
M. Brandon, P. Baldi, D.C. Wallace, Mitochondrial mutations in cancer. Oncogene 25, 4647–4662 (2006). https://doi.org/10.1038/sj.onc.1209607
J.B. Nunes, J. Peixoto, P. Soares, V. Maximo, S. Carvalho, S.S. Pinho, A.F. Vieira, J. Paredes, A.C. Rego, I.L. Ferreira, M. Gomez-Lazaro, M. Sobrinho-Simoes, K.K. Singh, J. Lima, OXPHOS dysfunction regulates integrin-β1 modifications and enhances cell motility and migration. Hum. Mol. Genet. 24, 1977–1990 (2015). https://doi.org/10.1093/hmg/ddu612
D. Grasso, L.X. Zampieri, T. Capelôa, J.A. Van de Velde, P. Sonveaux, Mitochondria in cancer. Cell Stress 4, 114–146 (2020). https://doi.org/10.15698/cst2020.06.221
A.S. Tan, J.W. Baty, L.F. Dong, A. Bezawork-Geleta, B. Endaya, J. Goodwin, M. Bajzikova, J. Kovarova, M. Peterka, B. Yan, E.A. Pesdar, M. Sobol, A. Filimonenko, S. Stuart, M. Vondrusova, K. Kluckova, K. Sachaphibulkij, J. Rohlena, P. Hozak, J. Truksa, D. Eccles, L.M. Haupt, L.R. Griffiths, J. Neuzil, M.V. Berridge, Mitochondrial genome acquisition restores respiratory function and tumorigenic potential of cancer cells without mitochondrial DNA. Cell. Metab. 21, 81–94 (2015). https://doi.org/10.1016/j.cmet.2014.12.003
L. Ippolito, A. Morandi, M.L. Taddei, M. Parri, G. Comito, A. Iscaro, M.R. Raspollini, F. Magherini, E. Rapizzi, J. Masquelier, G.G. Muccioli, P. Sonveaux, P. Chiarugi, E. Giannoni, Cancer-associated fibroblasts promote prostate cancer malignancy via metabolic rewiring and mitochondrial transfer. Oncogene 38, 5339–5355 (2019). https://doi.org/10.1038/s41388-019-0805-7
J.L. Spees, S.D. Olson, M.J. Whitney, D.J. Prockop, Mitochondrial transfer between cells can rescue aerobic respiration. Proc. Natl. Acad. Sci. U. S. A. 103, 1283–1288 (2006). https://doi.org/10.1073/pnas.0510511103
L.F. Dong, J. Kovarova, M. Bajzikova, A. Bezawork-Geleta, D. Svec, B. Endaya, K. Sachaphibulkij, A.R. Coelho, N. Sebkova, A. Ruzickova, A.S. Tan, K. Kluckova, K. Judasova, K. Zamecnikova, Z. Rychtarcikova, V. Gopalan, L. Andera, M. Sobol, B. Yan, B. Pattnaik, N. Bhatraju, J. Truksa, P. Stopka, P. Hozak, A.K. Lam, R. Sedlacek, P.J. Oliveira, M. Kubista, A. Agrawal, K. Dvorakova-Hortova, J. Rohlena, M.V. Berridge, J. Neuzil, Horizontal transfer of whole mitochondria restores tumorigenic potential in mitochondrial DNA-deficient cancer cells. Elife 6, (2017). https://doi.org/10.7554/eLife.22187
D. Torralba, F. Baixauli, F. Sánchez-Madrid, Mitochondria Know No Boundaries: mechanisms and functions of intercellular mitochondrial transfer. Front. Cell. Dev. Biol. 4, 107 (2016). https://doi.org/10.3389/fcell.2016.00107
D.C. Wallace, Mitochondria and cancer. Nat. Rev. Cancer 12, 685–698 (2012). https://doi.org/10.1038/nrc3365
P. Dey, A. Kundu, R. Sachan, J.H. Park, M.Y. Ahn, K. Yoon, J. Lee, N.D. Kim, I.S. Kim, B.M. Lee, H.S. Kim, PKM2 Knockdown induces autophagic cell death via AKT/mTOR pathway in human prostate Cancer cells. Cell. Physiol. Biochem. 52, 1535–1552 (2019). https://doi.org/10.33594/000000107
L. de Bari, L. Moro, S. Passarella, Prostate cancer cells metabolize d-lactate inside mitochondria via a D-lactate dehydrogenase which is more active and highly expressed than in normal cells. FEBS Lett. 587, 467–473 (2013). https://doi.org/10.1016/j.febslet.2013.01.011
Q. Liu, Y. Sun, Z. Fei, Z. Yang, K. Duan, J. Zi, Q. Cui, M. Yu, W. Xiong, Leptin promotes fatty acid oxidation and OXPHOS via the c-Myc/PGC-1 pathway in cancer cells. Acta Biochim. Biophys. Sin (Shanghai) 51, 707–714 (2019). https://doi.org/10.1093/abbs/gmz058
B.N. Whitley, E.A. Engelhart, S. Hoppins, Mitochondrial dynamics and their potential as a therapeutic target. Mitochondrion 49, 269–283 (2019). https://doi.org/10.1016/j.mito.2019.06.002
F. Bonnay, A. Veloso, V. Steinmann, T. Köcher, M.D. Abdusselamoglu, S. Bajaj, E. Rivelles, L. Landskron, H. Esterbauer, R.P. Zinzen, J.A. Knoblich, Oxidative metabolism drives immortalization of neural stem cells during tumorigenesis. Cell 182, 1490–1507.e1419 (2020). https://doi.org/10.1016/j.cell.2020.07.039
K. Saito, Q. Zhang, H. Yang, K. Yamatani, T. Ai, V. Ruvolo, N. Baran, T. Cai, H. Ma, R. Jacamo, V. Kuruvilla, J. Imoto, S. Kinjo, K. Ikeo, K. Moriya, K. Suzuki, T. Miida, Y.M. Kim, C.P. Vellano, M. Andreeff, J.R. Marszalek, Y. Tabe, M. Konopleva, Exogenous mitochondrial transfer and endogenous mitochondrial fission facilitate AML resistance to OxPhos inhibition. Blood Adv. 5, 4233–4255 (2021). https://doi.org/10.1182/bloodadvances.2020003661
S. Maertin, J.M. Elperin, E. Lotshaw, M. Sendler, S.D. Speakman, K. Takakura, B.M. Reicher, O.A. Mareninova, P.J. Grippo, J. Mayerle, M.M. Lerch, A.S. Gukovskaya, Roles of autophagy and metabolism in pancreatic cancer cell adaptation to environmental challenges. Am. J. Physiol. Gastrointest. Liver Physiol. 313, G524–Gg536 (2017). https://doi.org/10.1152/ajpgi.00138.2017
J. Franco, U. Balaji, E. Freinkman, A.K. Witkiewicz, E.S. Knudsen, Metabolic reprogramming of pancreatic Cancer mediated by CDK4/6 inhibition elicits unique vulnerabilities. Cell. Rep. 32, 107793 (2020). https://doi.org/10.1016/j.celrep.2020.107793
S. Kitajima, E. Ivanova, S. Guo, R. Yoshida, M. Campisi, S.K. Sundararaman, S. Tange, Y. Mitsuishi, T.C. Thai, S. Masuda, B.P. Piel, L.M. Sholl, P.T. Kirschmeier, C.P. Paweletz, H. Watanabe, M. Yajima, D.A. Barbie, Suppression of STING Associated with LKB1 loss in KRAS-Driven Lung Cancer. Cancer Discov 9, 34–45 (2019). https://doi.org/10.1158/2159-8290.Cd-18-0689
G. Wang, J. Xu, J. Zhao, W. Yin, D. Liu, W. Chen, S.X. Hou, Arf1-mediated lipid metabolism sustains cancer cells and its ablation induces anti-tumor immune responses in mice. Nat. Commun. 11, 220 (2020). https://doi.org/10.1038/s41467-019-14046-9
N. Jiang, B. Xie, W. Xiao, M. Fan, S. Xu, Y. Duan, Y. Hamsafar, A.C. Evans, J. Huang, W. Zhou, X. Lin, N. Ye, S. Wanggou, W. Chen, D. Jing, R.C. Fragoso, B.N. Dugger, P.F. Wilson, M.A. Coleman, S. Xia, X. Li, L.Q. Sun, A.M. Monjazeb, A. Wang, W.J. Murphy, H.J. Kung, K.S. Lam, H.W. Chen, J.J. Li, Fatty acid oxidation fuels glioblastoma radioresistance with CD47-mediated immune evasion. Nat. Commun. 13, 1511 (2022). https://doi.org/10.1038/s41467-022-29137-3
J. Li, Y. Ye, Z. Liu, G. Zhang, H. Dai, J. Li, B. Zhou, Y. Li, Q. Zhao, J. Huang, J. Feng, S. Liu, P. Ruan, J. Wang, J. Liu, M. Huang, X. Liu, S. Yu, Z. Liang, L. Ma, X. Gou, G. Zhang, N. Chen, Y. Lu, C. Di, Q. Xia, J. Pan, R. Feng, Q. Cai, S. Su, Macrophage mitochondrial fission improves cancer cell phagocytosis induced by therapeutic antibodies and is impaired by glutamine competition. Nat. cancer 3, 453–470 (2022). https://doi.org/10.1038/s43018-022-00354-5
P.S. Minhas, L. Liu, P.K. Moon, A.U. Joshi, C. Dove, S. Mhatre, K. Contrepois, Q. Wang, B.A. Lee, M. Coronado, D. Bernstein, M.P. Snyder, M. Migaud, R. Majeti, D. Mochly-Rosen, J.D. Rabinowitz, K.I. Andreasson, Macrophage de novo NAD(+) synthesis specifies immune function in aging and inflammation. Nat. Immunol. 20, 50–63 (2019). https://doi.org/10.1038/s41590-018-0255-3
J.M. Weiss, L.C. Davies, M. Karwan, L. Ileva, M.K. Ozaki, R.Y. Cheng, L.A. Ridnour, C.M. Annunziata, D.A. Wink, D.W. McVicar, Itaconic acid mediates crosstalk between macrophage metabolism and peritoneal tumors. J. Clin. Invest. 128, 3794–3805 (2018). https://doi.org/10.1172/jci99169
C. Liu, M. Chikina, R. Deshpande, A.V. Menk, T. Wang, T. Tabib, E.A. Brunazzi, K.M. Vignali, M. Sun, D.B. Stolz, R.A. Lafyatis, W. Chen, G.M. Delgoffe, C.J. Workman, S.G. Wendell, D.A.A. Vignali, Treg cells promote the SREBP1-Dependent metabolic fitness of Tumor-Promoting macrophages via repression of CD8(+) T cell-derived Interferon-γ. Immunity 51, 381–397.e386 (2019). https://doi.org/10.1016/j.immuni.2019.06.017
Y. Wang, F. Wang, L. Wang, S. Qiu, Y. Yao, C. Yan, X. Xiong, X. Chen, Q. Ji, J. Cao, G. Gao, D. Li, L. Zhang, Z. Guo, R. Wang, H. Wang, G. Fan, NAD(+) supplement potentiates tumor-killing function by rescuing defective TUB-mediated NAMPT transcription in tumor-infiltrated T cells. Cell. Rep. 36, 109516 (2021). https://doi.org/10.1016/j.celrep.2021.109516
B. Bengsch, A.L. Johnson, M. Kurachi, P.M. Odorizzi, K.E. Pauken, J. Attanasio, E. Stelekati, L.M. McLane, M.A. Paley, G.M. Delgoffe, E.J. Wherry, Bioenergetic insufficiencies due to metabolic alterations regulated by the inhibitory receptor PD-1 are an early driver of CD8(+) T cell exhaustion. Immunity 45, 358–373 (2016). https://doi.org/10.1016/j.immuni.2016.07.008
X. Zheng, Y. Qian, B. Fu, D. Jiao, Y. Jiang, P. Chen, Y. Shen, H. Zhang, R. Sun, Z. Tian, H. Wei, Mitochondrial fragmentation limits NK cell-based tumor immunosurveillance. Nat. Immunol. 20, 1656–1667 (2019). https://doi.org/10.1038/s41590-019-0511-1
X. Michelet, L. Dyck, A. Hogan, R.M. Loftus, D. Duquette, K. Wei, S. Beyaz, A. Tavakkoli, C. Foley, R. Donnelly, C. O’Farrelly, M. Raverdeau, A. Vernon, W. Pettee, D. O’Shea, B.S. Nikolajczyk, K.H.G. Mills, M.B. Brenner, D. Finlay, L. Lynch, Metabolic reprogramming of natural killer cells in obesity limits antitumor responses. Nat. Immunol. 19, 1330–1340 (2018). https://doi.org/10.1038/s41590-018-0251-7
H. Dong, N.M. Adams, Y. Xu, J. Cao, D.S.J. Allan, J.R. Carlyle, X. Chen, J.C. Sun, L.H. Glimcher, The IRE1 endoplasmic reticulum stress sensor activates natural killer cell immunity in part by regulating c-Myc. Nat. Immunol. 20, 865–878. https://doi.org/10.1038/s41590-019-0388-z
F. Hossain, A.A. Al-Khami, D. Wyczechowska, C. Hernandez, L. Zheng, K. Reiss, L.D. Valle, J. Trillo-Tinoco, T. Maj, W. Zou, P.C. Rodriguez, A.C. Ochoa, Inhibition of fatty acid oxidation modulates immunosuppressive functions of myeloid-derived suppressor cells and enhances Cancer Therapies. Cancer Immunol. Res. 3, 1236–1247 (2015). https://doi.org/10.1158/2326-6066.Cir-15-0036
C. Xu, S. Sun, T. Johnson, R. Qi, S. Zhang, J. Zhang, K. Yang, The glutathione peroxidase Gpx4 prevents lipid peroxidation and ferroptosis to sustain Treg cell activation and suppression of antitumor immunity. Cell. Rep. 35, 109235 (2021). https://doi.org/10.1016/j.celrep.2021.109235
G. Ercolano, A. Gomez-Cadena, N. Dumauthioz, G. Vanoni, M. Kreutzfeldt, T. Wyss, L. Michalik, R. Loyon, A. Ianaro, P.C. Ho, C. Borg, M. Kopf, D. Merkler, P. Krebs, P. Romero, S. Trabanelli, and C. Jandus, PPARɣ drives IL-33-dependent ILC2 pro-tumoral functions. Nat. Commun. 12, 2538 (2021). https://doi.org/10.1038/s41467-021-22764-2
T. Saha, C. Dash, R. Jayabalan, S. Khiste, A. Kulkarni, K. Kurmi, J. Mondal, P.K. Majumder, A. Bardia, H.L. Jang, S. Sengupta, Intercellular nanotubes mediate mitochondrial trafficking between cancer and immune cells. Nat. Nanotechnol 17, 98–106 (2022). https://doi.org/10.1038/s41565-021-01000-4
N.A. Bonekamp, B. Peter, H.S. Hillen, A. Felser, T. Bergbrede, A. Choidas, M. Horn, A. Unger, R. Di Lucrezia, I. Atanassov, X. Li, U. Koch, S. Menninger, J. Boros, P. Habenberger, P. Giavalisco, P. Cramer, M.S. Denzel, P. Nussbaumer, B. Klebl, M. Falkenberg, C.M. Gustafsson, N.G. Larsson, Small-molecule inhibitors of human mitochondrial DNA transcription. Nature 588, 712–716 (2020). https://doi.org/10.1038/s41586-020-03048-z
L. Testai, A. Martelli, L. Flori, A.F.G. Cicero, A. Colletti, Coenzyme Q(10): clinical applications beyond Cardiovascular Diseases. Nutrients 13, (2021). https://doi.org/10.3390/nu13051697
J.J. Shen, Y.C. Zhan, H.Y. Li, Z. Wang, Ouabain impairs cancer metabolism and activates AMPK-Src signaling pathway in human cancer cell lines. Acta Pharmacol. Sin 41, 110–118 (2020). https://doi.org/10.1038/s41401-019-0290-0
Q. Zhou, H. Li, Y. Li, M. Tan, S. Fan, C. Cao, F. Meng, L. Zhu, L. Zhao, M.X. Guan, H. Jin, Y. Sun, Inhibiting neddylation modification alters mitochondrial morphology and reprograms energy metabolism in cancer cells. JCI Insight 4, (2019). https://doi.org/10.1172/jci.insight.121582
S. Rodríguez-Enríquez, S.C. Pacheco-Velázquez, Á Marín-Hernández, J.C. Gallardo-Pérez, D.X. Robledo-Cadena, I. Hernández-Reséndiz, J.D. García-García, J. Belmont-Díaz, R. López-Marure, L. Hernández-Esquivel, R. Sánchez-Thomas, and R. Moreno-Sánchez, Resveratrol inhibits cancer cell proliferation by impairing oxidative phosphorylation and inducing oxidative stress. Toxicol. Appl. Pharmacol. 370, 65–77 (2019). https://doi.org/10.1016/j.taap.2019.03.008
V. Chen, R.E. Staub, S. Fong, M. Tagliaferri, I. Cohen, E. Shtivelman, Bezielle selectively targets mitochondria of cancer cells to inhibit glycolysis and OXPHOS. PLoS One 7, e30300 (2012). https://doi.org/10.1371/journal.pone.0030300
M.H. Cheng, H.L. Huang, Y.Y. Lin, K.H. Tsui, P.C. Chen, S.Y. Cheng, I.W. Chong, P.J. Sung, M.H. Tai, Z.H. Wen, N.F. Chen, H.M. Kuo, BA6 Induces Apoptosis via Stimulation of Reactive Oxygen Species and Inhibition of Oxidative Phosphorylation in Human Lung Cancer Cells. Oxid. Med. Cell. Longev. 2019, 6342104 (2019). https://doi.org/10.1155/2019/6342104
A.M. Stevens, M. Xiang, L.N. Heppler, I. Tošić, K. Jiang, J.O. Munoz, A.S. Gaikwad, T.M. Horton, X. Long, P. Narayanan, E.L. Seashore, M.C. Terrell, R. Rashid, M.J. Krueger, A.E. Mangubat-Medina, Z.T. Ball, P. Sumazin, S.R. Walker, Y. Hamada, S. Oyadomari, M.S. Redell, D.A. Frank, Atovaquone is active against AML by upregulating the integrated stress pathway and suppressing oxidative phosphorylation. Blood Adv. 3, 4215–4227 (2019). https://doi.org/10.1182/bloodadvances.2019000499
D. Xue, Y. Xu, A. Kyani, J. Roy, L. Dai, D. Sun, N. Neamati, Discovery and lead optimization of Benzene-1,4-disulfonamides as oxidative phosphorylation inhibitors. J. Med. Chem. 65, 343–368 (2022). https://doi.org/10.1021/acs.jmedchem.1c01509
J.S. Lee, H. Lee, H. Jang, S.M. Woo, J.B. Park, S.H. Lee, J.H. Kang, H.Y. Kim, J. Song, S.Y. Kim, Targeting oxidative phosphorylation reverses Drug Resistance in Cancer cells by blocking Autophagy Recycling. Cells 9, (2020). https://doi.org/10.3390/cells9092013
K. Kuramoto, M. Yamamoto, S. Suzuki, T. Sanomachi, K. Togashi, S. Seino, C. Kitanaka, M. Okada, Verteporfin inhibits oxidative phosphorylation and induces cell death specifically in glioma stem cells. FEBS J. 287, 2023–2036 (2020). https://doi.org/10.1111/febs.15187
S. Thakur, B. Daley, K. Gaskins, V.V. Vasko, M. Boufraqech, D. Patel, C. Sourbier, J. Reece, S.Y. Cheng, E. Kebebew, S. Agarwal, J. Klubo-Gwiezdzinska, Metformin Targets Mitochondrial Glycerophosphate Dehydrogenase to Control Rate of Oxidative Phosphorylation and Growth of Thyroid Cancer In Vitro and In Vivo. Clin. Cancer. Res. 24, 4030–4043 (2018). https://doi.org/10.1158/1078-0432.Ccr-17-3167
V. Pasquale, G. Ducci, G. Campioni, A. Ventrici, C. Assalini, S. Busti, M. Vanoni, R. Vago, E. Sacco, Profiling and targeting of Energy and Redox Metabolism in Grade 2 bladder Cancer cells with different Invasiveness Properties. Cells 9, (2020). https://doi.org/10.3390/cells9122669
Y. Liu, Y. Sun, Y. Guo, X. Shi, X. Chen, W. Feng, L.L. Wu, J. Zhang, S. Yu, Y. Wang and Y. Shi, An Overview: The Diversified Role of Mitochondria in Cancer Metabolism. Int. J. Biol. Sci. 19, 897-915 (2023). https://doi.org/10.7150/ijbs.81609
P.V. Raninga, A. Lee, D. Sinha, L.F. Dong, K.K. Datta, X. Lu, P. Kalita-de Croft, M. Dutt, M. Hill, N. Pouliot, H. Gowda, M. Kalimutho, J. Neuzil, K.K. Khanna, Marizomib suppresses triple-negative breast cancer via proteasome and oxidative phosphorylation inhibition. Theranostics 10, 5259–5275 (2020). https://doi.org/10.7150/thno.42705
C. Zhang, T. Liu, P. Luo, L. Gao, X. Liao, L. Ma, Z. Jiang, D. Liu, Z. Yang, Q. Jiang, Y. Wang, X. Tan, S. Luo, Y. Wang, C. Shi, Near-infrared oxidative phosphorylation inhibitor integrates acute myeloid leukemia-targeted imaging and therapy. Sci. Adv. 7, (2021). https://doi.org/10.1126/sciadv.abb6104
Y. Shi, S.K. Lim, Q. Liang, S.V. Iyer, H.Y. Wang, Z. Wang, X. Xie, D. Sun, Y.J. Chen, V. Tabar, P. Gutin, N. Williams, J.K. De Brabander, L.F. Parada, Gboxin is an oxidative phosphorylation inhibitor that targets glioblastoma. Nature 567, 341–346 (2019). https://doi.org/10.1038/s41586-019-0993-x
J.R. Molina, Y. Sun, M. Protopopova, S. Gera, M. Bandi, C. Bristow, T. McAfoos, P. Morlacchi, J. Ackroyd, A.A. Agip, G. Al-Atrash, J. Asara, J. Bardenhagen, C.C. Carrillo, C. Carroll, E. Chang, S. Ciurea, J.B. Cross, B. Czako, A. Deem, N. Daver, J.F. de Groot, J.W. Dong, N. Feng, G. Gao, J. Gay, M.G. Do, J. Greer, V. Giuliani, J. Han, L. Han, V.K. Henry, J. Hirst, S. Huang, Y. Jiang, Z. Kang, T. Khor, S. Konoplev, Y.H. Lin, G. Liu, A. Lodi, T. Lofton, H. Ma, M. Mahendra, P. Matre, R. Mullinax, M. Peoples, A. Petrocchi, J. Rodriguez-Canale, R. Serreli, T. Shi, M. Smith, Y. Tabe, J. Theroff, S. Tiziani, Q. Xu, Q. Zhang, F. Muller, R.A. DePinho, C. Toniatti, G.F. Draetta, T.P. Heffernan, M. Konopleva, P. Jones, M.E. Di Francesco and J.R. Marszalek, an inhibitor of oxidative phosphorylation exploits cancer vulnerability. Nat. Med. 24, 1036–1046 (2018). https://doi.org/10.1038/s41591-018-0052-4
M. Quintela-Fandino, S. Morales, A. Cortés-Salgado, L. Manso, J.V. Apala, M. Muñoz, A. Gasol Cudos, J. Salla Fortuny, M. Gion, A. Lopez-Alonso, J. Cortés, J. Guerra, D. Malón, E. Caleiras, F. Mulero, S. Mouron, Randomized phase 0/I trial of the mitochondrial inhibitor ME-344 or placebo added to Bevacizumab in Early HER2-Negative breast Cancer. Clinical cancer research: an official journal of the American Association for Cancer Research 26, 35–45. https://doi.org/10.1158/1078-0432.CCR-19-2023
L. Zhang, J. Zhang, Z. Ye, Y. Manevich, L.E. Ball, J.R. Bethard, Y.-L. Jiang, A.-M. Broome, A.C. Dalton, G.Y. Wang, D.M. Townsend, K.D. Tew, Isoflavone ME-344 disrupts Redox Homeostasis and mitochondrial Function by Targeting Heme Oxygenase 1. Cancer Res. 79, 4072–4085. https://doi.org/10.1158/0008-5472.CAN-18-3503
J.L. Carter, K. Hege, H.A. Kalpage, H. Edwards, M. Hüttemann, J.W. Taub, Y. Ge, Targeting mitochondrial respiration for the treatment of acute myeloid leukemia. Biochem. Pharmacol. 182, 114253. https://doi.org/10.1016/j.bcp.2020.114253
J. Zhang, L. Yan, P. Wei, R. Zhou, C. Hua, M. Xiao, Y. Tu, Z. Gu, T. Wei, PEG-GO@XN nanocomposite suppresses breast cancer metastasis via inhibition of mitochondrial oxidative phosphorylation and blockade of epithelial-to-mesenchymal transition. Eur. J. Pharmacol. 895, 173866 (2021). https://doi.org/10.1016/j.ejphar.2021.173866
G.A. Vitiello, B.D. Medina, S. Zeng, T.G. Bowler, J.Q. Zhang, J.K. Loo, N.J. Param, M. Liu, A.J. Moral, J.N. Zhao, F. Rossi, C.R. Antonescu, V.P. Balachandran, J.R. Cross, R.P. DeMatteo, Mitochondrial inhibition augments the efficacy of Imatinib by resetting the metabolic phenotype of gastrointestinal stromal tumor. Clin. Cancer Res. 24, 972–984 (2018). https://doi.org/10.1158/1078-0432.Ccr-17-2697
A.W. Jakobsson, S. Kundu, J. Guo, A. Chowdhury, M. Zhao, E. Lindell, P. Bergsten, F.J. Swartling, T. Sjöblom, X. Zhang, Iron Chelator VLX600 inhibits mitochondrial respiration and promotes sensitization of Neuroblastoma cells in Nutrition-Restricted conditions. Cancers (Basel) 14, (2022). https://doi.org/10.3390/cancers14133225
X. Zhang, M. Fryknäs, E. Hernlund, W. Fayad, A. De Milito, M.H. Olofsson, V. Gogvadze, L. Dang, S. Påhlman, L.A. Schughart, L. Rickardson, P. D’Arcy, J. Gullbo, P. Nygren, R. Larsson, S. Linder, Induction of mitochondrial dysfunction as a strategy for targeting tumour cells in metabolically compromised microenvironments. Nat. Commun. 5, 3295 (2014). https://doi.org/10.1038/ncomms4295
F. Basit, L.M. van Oppen, L. Schöckel, H.M. Bossenbroek, S.E. van Emst-de, J.C. Vries, S. Hermeling, C. Grefte, M. Kopitz, P. Heroult, Hgm Willems, W.J. Koopman, Mitochondrial complex I inhibition triggers a mitophagy-dependent ROS increase leading to necroptosis and ferroptosis in melanoma cells. Cell. Death Dis. 8, e2716 (2017). https://doi.org/10.1038/cddis.2017.133
P. Ellinghaus, I. Heisler, K. Unterschemmann, M. Haerter, H. Beck, S. Greschat, A. Ehrmann, H. Summer, I. Flamme, F. Oehme, K. Thierauch, M. Michels, H. Hess-Stumpp, K. Ziegelbauer, BAY 87-2243, a highly potent and selective inhibitor of hypoxia-induced gene activation has antitumor activities by inhibition of mitochondrial complex I. Cancer Med. 2, 611–624 (2013). https://doi.org/10.1002/cam4.112
R.J. Kishton, M. Sukumar, N.P. Restifo, Metabolic regulation of T cell longevity and function in Tumor Immunotherapy. Cell. Metab. 26, 94–109 (2017). https://doi.org/10.1016/j.cmet.2017.06.016
I. Hamaidi, L. Zhang, N. Kim, M.H. Wang, C. Iclozan, B. Fang, M. Liu, J.M. Koomen, A.E. Berglund, S.J. Yoder, J. Yao, R.W. Engelman, B.C. Creelan, J.R. Conejo-Garcia, S.J. Antonia, J.J. Mulé, S. Kim, Sirt2 inhibition enhances metabolic fitness and effector functions of Tumor-Reactive T cells. Cell. Metab. 32, 420–436.e412 (2020). https://doi.org/10.1016/j.cmet.2020.07.008
M. Luo, H. Wang, Z. Wang, H. Cai, Z. Lu, Y. Li, M. Du, G. Huang, C. Wang, X. Chen, M.R. Porembka, J. Lea, A.E. Frankel, Y.X. Fu, Z.J. Chen, J. Gao, A STING-activating nanovaccine for cancer immunotherapy. Nat. Nanotechnol 12, 648–654 (2017). https://doi.org/10.1038/nnano.2017.52
L. Guerra, L. Bonetti, D. Brenner, Metabolic modulation of immunity: a New Concept in Cancer Immunotherapy. Cell. Rep. 32, 107848 (2020). https://doi.org/10.1016/j.celrep.2020.107848
R.I. Klein Geltink, J. Edwards-Hicks, P. Apostolova, D. O’Sullivan, D.E. Sanin, A.E. Patterson, D.J. Puleston, N.A.M. Ligthart, J.M. Buescher, K.M. Grzes, A.M. Kabat, M. Stanczak, J.D. Curtis, F. Hässler, F.M. Uhl, M. Fabri, R. Zeiser, E.J. Pearce, E.L. Pearce, Metabolic conditioning of CD8(+) effector T cells for adoptive cell therapy. Nat. Metab. 2, 703–716 (2020). https://doi.org/10.1038/s42255-020-0256-z
O.U. Kawalekar, R.S. O’Connor, J.A. Fraietta, L. Guo, S.E. McGettigan, A.D. Posey Jr., P.R. Patel, S. Guedan, J. Scholler, B. Keith, N.W. Snyder, I.A. Blair, M.C. Milone, C.H. June, Distinct signaling of Coreceptors regulates specific metabolism pathways and impacts Memory Development in CAR T cells. Immunity 44, 380–390 (2016). https://doi.org/10.1016/j.immuni.2016.01.021
W. Li, S. Qiu, J. Chen, S. Jiang, W. Chen, J. Jiang, F. Wang, W. Si, Y. Shu, P. Wei, G. Fan, R. Tian, H. Wu, C. Xu, H. Wang, Chimeric Antigen receptor designed to prevent ubiquitination and downregulation showed durable Antitumor Efficacy. Immunity 53, 456–470.e456 (2020). https://doi.org/10.1016/j.immuni.2020.07.011
D. Alizadeh, R.A. Wong, X. Yang, D. Wang, J.R. Pecoraro, C.F. Kuo, B. Aguilar, Y. Qi, D.K. Ann, R. Starr, R. Urak, X. Wang, S.J. Forman, C.E. Brown, IL15 enhances CAR-T cell antitumor activity by reducing mTORC1 activity and preserving their stem cell memory phenotype. Cancer Immunol. Res. 7, 759–772 (2019). https://doi.org/10.1158/2326-6066.Cir-18-0466
C.R. Funk, S. Wang, K.Z. Chen, A. Waller, A. Sharma, C.L. Edgar, V.A. Gupta, S. Chandrakasan, J.T. Zoine, A. Fedanov, S.S. Raikar, J.L. Koff, C.R. Flowers, S. Coma, J.A. Pachter, S. Ravindranathan, H.T. Spencer, M. Shanmugam, E.K. Waller, PI3Kδ/γ inhibition promotes human CART cell epigenetic and metabolic reprogramming to enhance antitumor cytotoxicity. Blood 139, 523–537 (2022). https://doi.org/10.1182/blood.2021011597
V. Verma, N. Jafarzadeh, S. Boi, S. Kundu, Z. Jiang, Y. Fan, J. Lopez, R. Nandre, P. Zeng, F. Alolaqi, S. Ahmad, P. Gaur, S.T. Barry, V.E. Valge-Archer, P.D. Smith, J. Banchereau, M. Mkrtichyan, B. Youngblood, P.C. Rodriguez, S. Gupta, S.N. Khleif, MEK inhibition reprograms CD8(+) T lymphocytes into memory stem cells with potent antitumor effects. Nat. Immunol. 22, 53–66 (2021). https://doi.org/10.1038/s41590-020-00818-9
D. Chen, H.B. Barsoumian, G. Fischer, L. Yang, V. Verma, A.I. Younes, Y. Hu, F. Masropour, K. Klein, C. Vellano, J. Marszalek, M. Davies, M.A. Cortez, J. Welsh, Combination treatment with radiotherapy and a novel oxidative phosphorylation inhibitor overcomes PD-1 resistance and enhances antitumor immunity. J. Immunother Cancer 8, (2020). https://doi.org/10.1136/jitc-2019-000289
D.S. Vinay, E.P. Ryan, G. Pawelec, W.H. Talib, J. Stagg, E. Elkord, T. Lichtor, W.K. Decker, R.L. Whelan, H. Kumara, E. Signori, K. Honoki, A.G. Georgakilas, A. Amin, W.G. Helferich, C.S. Boosani, G. Guha, M.R. Ciriolo, S. Chen, S.I. Mohammed, A.S. Azmi, W.N. Keith, A. Bilsland, D. Bhakta, D. Halicka, H. Fujii, K. Aquilano, S.S. Ashraf, S. Nowsheen, X. Yang, B.K. Choi, B.S. Kwon, Immune evasion in cancer: mechanistic basis and therapeutic strategies. Semin. Cancer Biol 35(Suppl), 185–s198 (2015). https://doi.org/10.1016/j.semcancer.2015.03.004
P. Jin, J. Jiang, L. Zhou, Z. Huang, E.C. Nice, C. Huang, L. Fu, Mitochondrial adaptation in cancer drug resistance: prevalence, mechanisms, and management. J. Hematol. Oncol. 15, 97 (2022). https://doi.org/10.1186/s13045-022-01313-4
H. Tian, B. Zhang, L. Li, G. Wang, H. Li, J. Zheng, Manipulation of mitochondrial plasticity changes the metabolic competition between “Foe” and “Friend” during Tumor Malignant Transformation. Front. Oncol. 10, 1692 (2020). https://doi.org/10.3389/fonc.2020.01692
J. Han, M. Won, J.H. Kim, E. Jung, K. Min, P. Jangili, J.S. Kim, Cancer stem cell-targeted bio-imaging and chemotherapeutic perspective. Chem. Soc. Rev. 49, 7856–7878 (2020). https://doi.org/10.1039/d0cs00379d
R.A. Burrell, N. McGranahan, J. Bartek, C. Swanton, The causes and consequences of genetic heterogeneity in cancer evolution. Nature 501, 338–345 (2013). https://doi.org/10.1038/nature12625
S. Turajlic, A. Sottoriva, T. Graham, C. Swanton, Resolving genetic heterogeneity in cancer. Nat. Rev. Genet. 20, 404–416 (2019). https://doi.org/10.1038/s41576-019-0114-6
I. Vitale, E. Shema, S. Loi, L. Galluzzi, Intratumoral heterogeneity in cancer progression and response to immunotherapy. Nat. Med. 27, 212–224 (2021). https://doi.org/10.1038/s41591-021-01233-9
V. Sica, J.M. Bravo-San Pedro, G. Stoll, G. Kroemer, Oxidative phosphorylation as a potential therapeutic target for cancer therapy. Int. J. Cancer 146, 10–17 (2020). https://doi.org/10.1002/ijc.32616
T.A. Yap, N. Daver, M. Mahendra, J. Zhang, C. Kamiya-Matsuoka, F. Meric-Bernstam, H.M. Kantarjian, F. Ravandi, M.E. Collins, M.E.D. Francesco, E.E. Dumbrava, S. Fu, S. Gao, J.P. Gay, S. Gera, J. Han, D.S. Hong, E.J. Jabbour, Z. Ju, D.D. Karp, A. Lodi, J.R. Molina, N. Baran, A. Naing, M. Ohanian, S. Pant, N. Pemmaraju, P. Bose, S.A. Piha-Paul, J. Rodon, C. Salguero, K. Sasaki, A.K. Singh, V. Subbiah, A.M. Tsimberidou, Q.A. Xu, M. Yilmaz, Q. Zhang, Y. Li, C.A. Bristow, M.B. Bhattacharjee, S. Tiziani, T.P. Heffernan, C.P. Vellano, P. Jones, C.J. Heijnen, A. Kavelaars, J.R. Marszalek, M. Konopleva, Complex I inhibitor of oxidative phosphorylation in advanced solid tumors and acute myeloid leukemia: phase I trials. Nat. Med. 29, 115–126 (2023). https://doi.org/10.1038/s41591-022-02103-8
X. Zhang, C.V. Dang, Time to hit pause on mitochondria-targeting cancer therapies. Nat. Med. 29, 29–30 (2023). https://doi.org/10.1038/s41591-022-02129-y
F. Janku, P. LoRusso, A.S. Mansfield, R. Nanda, A. Spira, T. Wang, A. Melhem-Bertrandt, J. Sugg, H.A. Ball, First-in-human evaluation of the novel mitochondrial complex I inhibitor ASP4132 for treatment of cancer. Invest. New. Drugs 39, 1348–1356 (2021). https://doi.org/10.1007/s10637-021-01112-7
Y. Xu, D. Xue, A. Bankhead, 3rd and N. Neamati, Why all the fuss about oxidative phosphorylation (OXPHOS)? J. Med. Chem. 63, 14276–14307 (2020). https://doi.org/10.1021/acs.jmedchem.0c01013
K. DePeaux, G.M. Delgoffe, Metabolic barriers to cancer immunotherapy. Nat. Rev. Immunol. 21, 785–797 (2021). https://doi.org/10.1038/s41577-021-00541-y
K.C. Kao, S. Vilbois, C.H. Tsai, P.C. Ho, Metabolic communication in the tumour-immune microenvironment. Nat. Cell. Biol. (2022). https://doi.org/10.1038/s41556-022-01002-x
P. Ghosh, C. Vidal, S. Dey, L. Zhang, Mitochondria Targeting as an effective strategy for Cancer Therapy. Int. J. Mol. Sci. 21, (2020). https://doi.org/10.3390/ijms21093363
X.T. Le, J. Lee, N.T. Nguyen, W.T. Lee, E.S. Lee, K.T. Oh, H.G. Choi, B.S. Shin, Y.S. Youn, Combined phototherapy with metabolic reprogramming-targeted albumin nanoparticles for treating breast cancer. Biomaterials Sci. 10, 7117–7132 (2022). https://doi.org/10.1039/d2bm01281b
X. Lei, K. Li, Y. Liu, Z.Y. Wang, B.J. Ruan, L. Wang, A. Xiang, D. Wu, Z. Lu, Co-delivery nanocarriers targeting folate receptor and encapsulating 2-deoxyglucose and α-tocopheryl succinate enhance anti-tumor effect in vivo. Int. J. Nanomed 12, 5701–5715 (2017). https://doi.org/10.2147/ijn.S135849
Funding
Open Access funding enabled and organized by Projekt DEAL. This work was financially supported by the National Natural Science Foundation of China (Grant No. 82103460 and No. 81972546), the Fundamental Research Funds for Central Universities.
Author information
Authors and Affiliations
Contributions
X.T.Q: Writing - Original Draft; Y.L: Writing - Reviewing & Editing; Z.Y.Z: Conceptualization, Writing - Reviewing & Editing.
Corresponding author
Ethics declarations
Ethics approval
No potential ethical issues were disclosed.
Conflict of interest
No potential conflicts of interest were disclosed.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Qiu, X., Li, Y. & Zhang, Z. Crosstalk between oxidative phosphorylation and immune escape in cancer: a new concept of therapeutic targets selection. Cell Oncol. 46, 847–865 (2023). https://doi.org/10.1007/s13402-023-00801-0
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13402-023-00801-0