Abstract
Background
Nonsyndromic autosomal recessive hearing loss (DFNB) is an etiologically heterogeneous disorder group showing a wide spectrum of onset ages and severity. DFNB genes are very diverse in their types and functions, making molecular diagnosis difficult. DFNB is particularly frequent in Pakistan, which may be partly due to consanguinity.
Objective
This study was performed to determine the genetic causes in Pakistani DFNB families with prelingual onset and to establish genotype-phenotype correlation.
Methods
Whole exome sequencing and subsequent genetic analysis were performed for 11 Pakistani DFNB families including eight consanguineous families.
Results
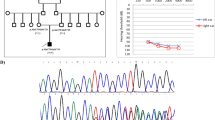
We identified eight pathogenic or likely pathogenic mutations in LOXHD1, GJB2, SLC26A4, MYO15A, and TMC1 from six families. The GJB2 mutations were identified in two families each with compound heterozygous mutations and a homozygous mutation. The compound heterozygous mutations in LOXHD1 ([p.D278Y] + [p.D1219E]) and GJB2 [p.M1?] + [p.G12Vfs*2]) were novel. The four missense or start-loss mutations were located at well conserved residues, and most in silico analysis predicted their pathogenicity. In addition to causative mutations, we found compound heterozygous mutations in PTPRQ as variants of uncertain significance.
Conclusion
This study identified biallelic mutations as the underlying cause of early onset DFNB in six Pakistani families. This study will be helpful in providing an exact molecular diagnosis and treatment of prelingual onset deafness patients.





Similar content being viewed by others
Data availability
All raw genetic data generated during this study are available upon request to the corresponding author.
References
Akil O, Dyka F, Calvet C, Emptoz A, Lahlou G, Nouaille S, Boutet de Monvel J, Hardelin JP, Hauswirth WW, Avan P et al (2019) Dual AAV-mediated gene therapy restores hearing in a DFNB9 mouse model. Proc Natl Acad Sci USA 116:4496–4501
Anwar S, Riazuddin S, Ahmed ZM, Tasneem S, Ateeq-ul-Jaleel, Khan SY, Griffith AJ, Friedman TB, Riazuddin S (2009) SLC26A4 mutation spectrum associated with DFNB4 deafness and Pendred’s syndrome in Pakistanis. J Hum Genet 54:266–270
Carrasquillo MM, Zlotogora J, Barges S, Chakravarti A (1997) Two different connexin 26 mutations in an inbred kindred segregating non-syndromic recessive deafness: implications for genetic studies in isolated populations. Hum Mol Genet 6:2163–2172
Dianatpour M, Smith E, Hashemi SB, Farazifard MA, Nezafat N, Razban V, Mani A (2021) Identification of homozygous mutations for hearing loss. Gene 778:145464
Ding Y, Leng J, Fan F, Xia B, Xu P (2013) The role of mitochondrial DNA mutations in hearing loss. Biochem Genet 51:588–602
Dror AA, Politi Y, Shahin H, Lenz DR, Dossena S, Nofziger C, Fuchs H, Hrabe de Angelis M, Paulmichl M, Weiner S et al (2010) Calcium oxalate stone formation in the inner ear as a result of an Slc26a4 mutation. J Biol Chem 285:21724–21735
Edvardson S, Jalas C, Shaag A, Zenvirt S, Landau C, Lerer I, Elpeleg O (2011) A deleterious mutation in the LOXHD1 gene causes autosomal recessive hearing loss in Ashkenazi Jews. Am J Med Genet 155A:1170–1172
Eisenberger T, Di Donato N, Decker C, Delle Vedove A, Neuhaus C, Nürnberg G, Toliat M, Nürnberg P, Mürbe D, Bolz HJ (2018) A C-terminal nonsense mutation links PTPRQ with autosomal-dominant hearing loss, DFNA73. Genet Med 20:614–621
Everett LA, Belyantseva IA, Noben-Trauth K, Cantos R, Chen A, Thakkar SI, Hoogstraten-Miller SL, Kachar B, Wu DK, Green ED (2001) Targeted disruption of mouse pds provides insight about the inner-ear defects encountered in Pendred syndrome. Hum Mol Genet 10:153–161
Everett LA, Glaser B, Beck JC, Idol JR, Buchs A, Heyman M, Adawi F, Hazani E, Nassir E, Baxevanis AD et al (1997) Pendred syndrome is caused by mutations in a putative sulphate transporter gene (PDS). Nat Genet 17:411–422
Gasteiger E, Gattiker A, Hoogland C, Ivanyi I, Appel RD, Bairoch A (2003) ExPASy: the proteomics server for in-depth protein knowledge and analysis. Nucleic Acids Res 31:3784–3788
Grillet N, Schwander M, Hildebrand MS, Sczaniecka A, Kolatkar A, Velasco J, Webster JA, Kahrizi K, Najmabadi H, Kimberling WJ et al (2009) Mutations in LOXHD1, an evolutionarily conserved stereociliary protein, disrupt hair cell function in mice and cause progressive hearing loss in humans. Am J Hum Genet 85:328–337
Hildebrand MS, Thorne NP, Bromhead CJ, Kahrizi K, Webster JA, Fattahi Z, Bataejad M, Kimberling WJ, Stephan D, Najmabadi H et al (2010) Variable hearing impairment in a DFNB2 family with a novel MYO7A missense mutation. Clin Genet 77:563–571
Kelsell DP, Dunlop J, Stevens HP, Lench NJ, Liang JN, Parry G, Mueller RF, Leigh IM (1997) Connexin 26 mutations in hereditary non-syndromic sensorineural deafness. Nature 387:80–83
Khan A, Han S, Wang R, Ansar M, Ahmad W, Zhang X (2019) Sequence variants in genes causing nonsyndromic hearing loss in a pakistani cohort. Mol Genet Genomic Med 7:e917
Kitajiri SI, McNamara R, Makishima T, Husnain T, Zafar AU, Kittles RA, Ahmed ZM, Friedman TB, Riazuddin S, Griffith AJ (2007) Identities, frequencies and origins of TMC1 mutations causing DFNB7/B11 deafness in Pakistan. Clin Genet 72:546–550
Kremer H (2019) Hereditary hearing loss; about the known and the unknown. Hear Res 376:58–68
Kudo T, Ikeda K, Oshima T, Kure S, Tammasaeng M, Prasansuk S, Matsubara Y (2001) GJB2 (connexin 26) mutations and childhood deafness in Thailand. Otol Neurotol 22:858–861
Kurima K, Peters LM, Yang Y, Riazuddin S, Ahmed ZM, Naz S, Arnaud D, Drury S, Mo J, Makishima T et al (2002) Dominant and recessive deafness caused by mutations of a novel gene, TMC1, required for cochlear hair-cell function. Nat Genet 30:277–284
Le Nabec A, Collobert M, Le Maréchal C, Marianowski R, Férec C, Moisan S (2021) Whole-genome sequencing improves the diagnosis of DFNB1 monoallelic patients. Genes (Basel) 12:1267
Lee AJ, Nam DE, Choi YJ, Nam SH, Choi BO, Chung KW (2020) Alanyl-tRNA synthetase 1 (AARS1) gene mutation in a family with intermediate Charcot-Marie-Tooth neuropathy. Genes Genom 42:663–672
Liburd N, Ghosh M, Riazuddin S, Naz S, Khan S, Ahmed Z, Riazuddin S, Liang Y, Menon PS, Smith T et al (2001) Novel mutations of MYO15A associated with profound deafness in consanguineous families and moderately severe hearing loss in a patient with Smith-Magenis syndrome. Hum Genet 109:535–541
Lieu JEC, Kenna M, Anne S, Davidson L (2020) Hearing loss in children: a review. JAMA 324:2195–2205
Maeda S, Nakagawa S, Suga M, Yamashita E, Oshima A, Fujiyoshi Y, Tsukihara T (2009) Structure of the connexin 26 gap junction channel at 3.5 Å resolution. Nature 458:597–602
Maekawa K, Nishio SY, Abe S, Goto SI, Honkura Y, Iwasaki S, Kanda Y, Kobayashi Y, Oka SI, Okami M et al (2019) Mutational spectrum and clinical features of patients with LOXHD1 variants identified in an 8074 hearing loss patient cohort. Genes (Basel) 10:735
Mahmood U, Bukhari SA, Ali M, Ahmed ZM, Riazuddin S (2021) Identification of hearing loss-associated variants of PTPRQ, MYO15A, and SERPINB6 in Pakistani families. Biomed Res Int 2021:5584788
Morton CC, Nance WE (2006) Newborn hearing screening: a silent revolution. N Engl J Med 354:2151–2164
Nal N, Ahmed ZM, Erkal E, Alper OM, Luleci G, Dinc O, Waryah AM et al (2007) Mutational spectrum of MYO15A: the large N-terminal extension of myosin XVA is required for hearing. Hum Mutat 28:1014–1019
Naz S, Imtiaz A, Mujtaba G, Maqsood A, Bashir R, Bukhari I, Khan MR, Ramzan M, Fatima A, Rehman AU et al (2017) Genetic causes of moderate to severe hearing loss point to modifiers. Clin Genet 91:589–598
Omichi R, Shibata SB, Morton CC, Smith RJH (2019) Gene therapy for hearing loss. Hum Mol Genet 28:R65–R79
Park HR, Kanwal S, Lim SO, Nam DE, Choi YJ, Chung KW (2020) Homozygous mutations in pakistani consanguineous families with prelingual nonsyndromic hearing loss. Mol Biol Rep 47:9979–9985
Petersen MB, Willems PJ (2006) Non-syndromic, autosomal-recessive deafness. Clin Genet 69:371–392
Piazza V, Beltramello M, Menniti M, Colao E, Malatesta P, Argento R, Chiarella G, Gallo LV, Catalano M, Perrotti N et al (2005) Functional analysis of R75Q mutation in the gene coding for connexin 26 identified in a family with nonsyndromic hearing loss. Clin Genet 68:161–166
Reardon W, OMahoney CF, Trembath R, Jan H, Phelps PD (2000) Enlarged vestibular aqueduct: a radiological marker of pendred syndrome, and mutation of the PDS gene. QJM 93:99–104
Richard EM, Santos-Cortez RLP, Faridi R, Rehman AU, Lee K, Shahzad M, Acharya A, Khan AA, Imtiaz A, Chakchouk I et al (2019) Global genetic insight contributed by consanguineous pakistani families segregating hearing loss. Hum Mutat 40:53–72
Santos RL, Wajid M, Khan MN, McArthur N, Pham TL, Bhatti A, Lee K, Irshad S, Mir A, Yan K et al (2005) Novel sequence variants in the TMC1 gene in pakistani families with autosomal recessive hearing impairment. Hum Mutat 26:396
Schraders M, Oostrik J, Huygen PL, Strom TM, van Wijk E, Kunst HP, Hoefsloot LH, Cremers CW, Admiraal RJ, Kremer H (2010) Mutations in PTPRQ are a cause of autosomal-recessive nonsyndromic hearing impairment DFNB84 and associated with vestibular dysfunction. Am J Hum Genet 86:604–610
Schrauwen I, Helfmann S, Inagaki A, Predoehl F, Tabatabaiefar MA, Picher MM, Sommen M, Zazo Seco C, Oostrik J, Kremer H et al (2012) A mutation in CABP2, expressed in cochlear hair cells, causes autosomal-recessive hearing impairment. Am J Hum Genet 91:636–645
Sirmaci A, Duman D, Oztürkmen-Akay H, Erbek S, Incesulu A, Oztürk-Hişmi B, Arici ZS, Yüksel-Konuk EB, Taşir-Yilmaz S, Tokgöz-Yilmaz S et al (2009) Mutations in TMC1 contribute significantly to nonsyndromic autosomal recessive sensorineural hearing loss: a report of five novel mutations. Int J Pediatr Otorhinolaryngol 73:699–705
Sloan-Heggen CM, Bierer AO, Shearer AE, Kolbe DL, Nishimura CJ, Frees KL, Ephraim SS, Shibata SB, Booth KT, Campbell CA et al (2016) Comprehensive genetic testing in the clinical evaluation of 1119 patients with hearing loss. Hum Genet 135:441–450
Snoeckx RL, Huygen PL, Feldmann D, Marlin S, Denoyelle F, Waligora J, Mueller-Malesinska M, Pollak A, Ploski R, Murgia A et al (2005) GJB2 mutations and degree of hearing loss: a multicenter study. Am J Hum Genet 77:945–957
Tang H-Y, Fang P, Ward PA, Schmitt E, Darilek S, Manolidis S, Oghalai JS, Roa BB, Alford RL (2006) DNA sequence analysis of GJB2, encoding connexin 26: observations from a population of hearing impaired cases and variable carrier rates, complex genotypes, and ethnic stratification of alleles among controls. Am J Med Genet 140A:2401–2415
Taylor JP, Metcalfe RA, Watson PF, Weetman AP, Trembath RC (2002) Mutations of the PDS gene, encoding pendrin, are associated with protein mislocalization and loss of iodide efflux: implications for thyroid dysfunction in Pendred syndrome. J Clin Endocrinol Metab 87:1778–1784
Thönnissen E, Rabionet R, Arbonès ML, Estivill X, Willecke K, Ott T (2002) Human connexin26 (GJB2) deafness mutations affect the function of gap junction channels at different levels of protein expression. Hum Genet 111:190–197
Trouillet A, Miller KK, George SS, Wang P, Ali NE, Ricci A, Grillet N (2021) Loxhd1 mutations cause mechanotransduction defects in cochlear hair cells. J Neurosci 41:3331–3343
Tucci D, Merson MH, Wilson BS (2010) A summary of the literature on global hearing impairment: current status and priorities for action. Otol Neurotol 31:31–41
Ullah S, Aslamkhan M, Rasheed A (2015) Molecular distribution of deafness loci in various ethnic groups of the Punjab, Pakistan. J Coll Physicians Surg Pak 25:573–578
Usami S, Abe S, Weston MD, Shinkawa H, Van Camp G, Kimberling WJ (1999) Non-syndromic hearing loss associated with enlarged vestibular aqueduct is caused by PDS mutations. Hum Genet 104:188–192
Wang A, Liang Y, Fridell RA, Probst FJ, Wilcox ER, Touchman JW, Morton CC, Morell RJ, Noben-Trauth K, Camper SA et al (1998) Association of unconventional myosin MYO15 mutations with human nonsyndromic deafness DFNB3. Science 280:1447–1451
Wells HRR, Freidin MB, Zainul Abidin FN, Payton A, Dawes P, Munro KJ, Morton CC, Moore DR, Dawson SJ, Williams FMK (2019) GWAS identifies 44 independent associated genomic loci for self-reported adult hearing difficulty in UK biobank. Am J Hum Genet 105:788–802
Zelante L, Gasparini P, Estivill X, Melchionda S, D’Agruma L, Govea N, Milá M, Monica MD, Lutfi J, Shohat M et al (1997) Connexin26 mutations associated with the most common form of non-syndromic neurosensory autosomal recessive deafness (DFNB1) in Mediterraneans. Hum Mol Genet 6:1605–1609
Acknowledgements
This research was supported by grants from the National Research Foundation (2019R1A2C1087547 and 2021R1A4A2001389), Republic of Korea.
Author information
Authors and Affiliations
Contributions
Chung KW planned and supervised this study. Kanwal S, Hameed R, Perveen S, and Mahreen H collected HL family samples and clinical information. Choi HJ, Kanwal S, and Tamanna N performed the molecular genetic work. Choi HJ, Son W, Lee KS, and Chung KW interpreted the genetic data and conducted the statistical analyses. Son W, and Chung KW wrote the manuscript. All the co-authors read and approved the final version of the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
Choi HJ, Kanwal S, Hameed R, Tamanna N, Perveen S, Mahreen H, Son W, Lee KS, and Chung KW declare that they have no conflict of interest.
Ethics approval and consent to participate
All participants provided written informed consent according to the Declaration of Helsinki. For minors under the age of 18 years, written consent was provided by their parents. This study was approved by the Institutional Review Board of Kongju National University (KNU_IRB_2018-62).
Patient consent for publication
Not applicable
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
HJ Choi and S Kanwal contributed equally to this work.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Choi, H.J., Kanwal, S., Hameed, R. et al. Biallelic mutations in pakistani families with autosomal recessive prelingual nonsyndromic hearing loss. Genes Genom 45, 145–156 (2023). https://doi.org/10.1007/s13258-022-01349-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13258-022-01349-3