Abstract
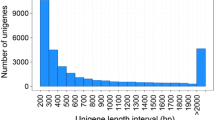
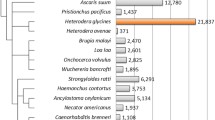
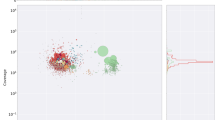
Root-knot nematodes are obligatory sedentary endoparasites that require a plant host to complete their life cycle. To understand the functions of Meloidogyne incognita nematode genes transcribed from eggs and second-stage juveniles (J2), we have constructed a normalized full-length M. incognita cDNA library and analyzed the ESTs using Pendant-Pro Sequence Analysis Suite. The 5,832 M. incognita ESTs formed 3,263 clusters and 2,241 singletons. The sequences ranged from 51 to 1,740 base pairs, and their average size was 699 base pairs. The protein length of M. incognita ESTs ranged from 150 to 299 amino acids. Forty contigs of predicted proteins that were grouped by BLASTP identity values had significant homology to the genes expressed in their organelle structures (cuticle, epidermis, extracellular matrix and muscle). Using the gomerger method of contigs, we could functionally assign GO terms to 2,147 (53.4%) of 4,024 contigs. Following the E.C. numbers method using UniProt database hits, we could functionally classify E.C. numbers to 288 (7.2%) of 4024 contigs. Also, the taxonomy was classified to 2,329 (57.9%) of 4,024 contigs. We could predict transmembrane regions of 4,024 clusters using the TMpred algorithm. Of the 4,024 contigs with transmembrane regions, 1,457 (36.2%) were assigned more than one domain, and 2,567 (63.8%) could not be assigned a transmembrane domain. The M. incognita ESTs will provide a foundation for developing novel target genes for parasite control and contribute to accelerating the research of biologically-related species.
Similar content being viewed by others
References
Abad P, Gouzy J, Aury JM, Castagnone-Sereno P, Danchin EG, et al. (2008) Genome sequence of the metazoan plant-parasitic nematode Meloidogyne incognita. Nat. Biotechnol. 26: 909–915.
Alkharouf NW, Klink VP and Matthews BF (2007) Identification of Heterodera glycines (soybean cyst nematode [SCN]) cDNA sequences with high identity to those of Caenorhabditis elegans having lethal mutant or RNAi phenotypes. Exp. Parasitol. 115: 247–258.
Altschul SF, Madden TL, Schaffer AA, Zhang J, Zhang Z, Miller W and Lipman DJ (1997) Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res. 25: 3389–3402.
Apweiler R, Attwood TK, Bairoch A, Bateman A, Birney E, Biswas M, Bucher P, Cerutti L, Corpet F, Croning MD, et al. (2001) The InterPro database, an integrated documentation resource for protein families, domains and functional sites. Nucleic Acids Res. 29: 37–40.
Ashburner M and Lewis SE (2002) On ontologies for biologists: the Gene Ontology — uncoupling the web. Novartis Found Symp. 247: 66–80
Bateman A, Birney E, Cerruti L, Durbin R, Etwiller L, Eddy SR, Griffiths-Jones S, Howe KL, Marshall M and Sonnhammer EL (2002) The Pfam protein families database. Nucleic Acids Res. 30: 276–280.
Bellafiore S, Shen Z, Rosso MN, Abad P, Shih P and Briggs SP (2008) Direct identification of the Meloidogyne incognita secretome reveals proteins with host cell reprogramming potential. PLoS. Pathog. 4: e1000192.
Bird DM, Opperman CH, Jones SJ and Baillie DL (1999) The Caenorhabditis elegans genome: A Guide in The Post Genomics Age. Annu. Rev. Phytopathol. 37:247–265
Burke J, Davison D and Hide W (1999) d2_cluster: a validated method for clustering EST and full-length cDNA sequences. Genome Res. 9: 1135–1142.
Dubreuil G, Magliano M, Deleury E, Abad P and Rosso MN (2007) Transcriptome analysis of root knot nematode functions induced in the early stages of parasitism. N Phytol. 176:426–436.
Ewing B and Green P (1998) Base-calling of automated sequencer traces using phred. II. Error probabilities. Genome Res. 8: 186–194.
Fairbairn DJ, Cavallaro AS, Bernard M, Mahalinga-Iyer J, Graham MW and Botella JR (2007) Host-delivered RNAi: an effective strategy to silence genes in plant parasitic nematodes. Planta. 226: 1525–1533.
Falquet L, Pagni M, Bucher P, Hulo N, Sigrist CJ, Hofmann K and Bairoch A (2002) The PROSITE database, its status in 2002. Nucleic Acids Res. 30: 235–238.
Fire A, Xu SQ, Montgomery MK, Kostas SA, Driver SE and Mello CC (1998) Potent and specific genetic interference by double-stranded RNA in Caenorhabditis elegans. Nature 391: 806–811.
Hahn BS and Labouesse M (2001) Tissue integrity: hemidesmosomes and resistance to stress. Curr. Biol. 11: R858–861.
Henikoff S, Henikoff JG and Pietrokovski S (1999) Blocksþ: a non-redundant database of protein alignment blocks derived from multiple compilations. Bioinformatics 15: 471–479.
Hofmann K and Stoffel W (1993) TMbase-A database of membrane spanning proteins segments. Biol. Chem. Hoppe-Seyler 374: 166
Huang X and Madan A (1999) CAP3: A DNA sequence assembly program. Genome Res. 9: 868–877.
Hussey RS and Barker KR (1973) A comparison of methods of collecting inocular of Meloidogyne spp., including a new technique. Plant Dis. Rep. 57: 1025–1028.
Hussey RS (1989) Disease-inducing secretion of plant partasitic nematodes. Annu. Rev. Phytopathol. 27:123–141.
Jones MG and Northcote DH (1972) Nematode-induced syncytium-a multinucleate transfer cell. J. Cell Sci. 10: 789–809.
Kikuchi T, Aikawa T, Kosaka H, Pritchard L, Ogura N and Jones JT (2007) Expressed sequence tag (EST) analysis of the pine wood nematode Bursaphelenchus xylophilus and B mucronatus. Mol. Biochem. Parasitol. 155:9–17.
Kimber MJ, McKinney S, McMaster S, Day TA, Fleming CC and Maule AG (2007) flp gene disruption in a parasitic nematode reveals motor dysfunction and unusual neuronal sensitivity to RNA interference. FASEB J. 21: 1233–1243.
Klink VP, Martins VE, Alkharouf NW, Overall CC, MacDonald MH and Matthews BF (2007). A decline in transcript abundance for Heterodera glycines homologs of Caenorhabditis elegans uncoordinated genes accompanies its sedentary parasitic phase. BMC Dev. Biol. 7:35.
Maeda I, Kohara Y, Yamamoto M and Sugimoto A (2001) Large-scale analysis of gene function in Caenorhabditis elegans by high-throughput RNAi. Curr. Biol. 11: 171–176.
Martin DM, Berriman M and Barton GJ (2004) GOtcha: a new method for prediction of protein function assessed by the annotation of seven genomes. BMC Bioinformatics 5: 178.
Maruyama K and Sugano S (1994) Oligo-capping: a simple method to replace the cap structure of eukaryotic mRNAs with oligoribonucleotides. Gene 138: 171–174.
McCarter JP, Mitreva MD, Martin J, Dante M, Wylie T, Rao U, Pape D, Bowers Y, Theising B, Murphy CV, Kloek AP, Chiapelli BJ, Clifton SW, Bird DM and Waterston RH (2003) Analysis and functional classification of transcripts from the nematode Meloidogyne incognita. Genome Biol. 4: 26.
Nagaraj SH, Gasser RB and Ranganathan S (2008) Needles in the EST Haystack: Large-scale identification and analysis of excretory-secretory (ES) proteins in parasitic nematodes using expressed sequence tags (ESTs). PLoS. Negl. Trop. Dis. 2: e301.
Oh JH, Kim YS and Kim NS (2003) An improved method for constructing a full-length enriched cDNA library using small amounts of total RNA as a starting material. Exp. Mol. Med. 35: 586–590.
Opperman CH, Bird DM, Williamson VM, Rokhsar DS, Burke M, Cohn J, Cromer J, Diener S, Gajan J, Graham S, Houfek TD, Liu Q, Mitros T, Schaff J, Schaffer R, Scholl E, Sosinski BR, Thomas VP and Windham E (2008) Sequence and genetic map of Meloidogyne hapla: A compact nematode genome for plant parasitism. Proc. Natl. Acad. Sci. U.S.A. 105: 14802–14807.
Roze E, Hanse B, Mitreva M, Vanholme B, Bakker J and Smant G (2008) Mining the secretome of the root-knot nematode Meloidogyne chitwoodi for candidate parasitism genes. Mol. Plant Pathol. 9:1–10.
Sasser JN (1980) Root-knot nematodes: A global menace to crop production. Plant Dis. 64: 36–41.
Sindhu AS, Maier TR, Mitchum MG, Hussey RS, Davis EL and Baum TJ (2009) Effective and specific in planta RNAi in cyst nematodes: expression interference of four parasitism genes reduces parasitic success. J. Exp. Bot. 60: 315–324.
Soares MB, Bonaldo MF, Jelene P, Su L, Lawton L and Efstratiadis A (1994) Construction and characterization of a normalized cDNA library. Proc. Natl. Acad. Sci. U.S.A. 91: 9228–9232.
Timmons L and Fire A (1998) Specific interference by ingested dsRNA. Nature 395: 854.
Vanholme B, Mitreva M, Van Criekinge W, Logghe M, Bird D, McCarter JP and Gheysen G (2006) Detection of putative secreted proteins in the plant-parasitic nematode Heterodera schachtii. Parasitol. Res. 98: 414–424.
Vieira J and Messing J (1987) Production of single-stranded plasmid DNA. Methods. Enzymol. 153: 3–11.
Wiggers RJ, Starr JL and Price HJ (1990) DNA content and variation in chromosome number in plant cells affected by Meloidogyne incognita and M. arenaria. Phytopath.ology 80: 1391–1395.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Kang, MJ., Kim, YH. & Hahn, BS. Expressed sequence tag analysis generated from a normalized full-length cDNA library of the root-knot nematode (Meloidogyne incognita). Genes Genom 32, 553–562 (2010). https://doi.org/10.1007/s13258-010-0065-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13258-010-0065-y