Abstract
Various Genome-wide association studies (GWAS) have reported the association of variant rs2494938 with lung cancer. However, genetic association of LRFN2 genetic variation with non-small cell lung cancer (NSCLC) in North Indian population remained unexplored. We conducted a case–control association study using TaqMan-based chemistry in which a total of 619 individuals, 189 NSCLC cases and 430 controls, were genotyped to explore the association of rs2494938 genetic variant of the LRFN2 gene with NSCLC patients from North India. The allele ‘G’ (risk allele) of the genetic variant rs2494938 was significantly associated with the NSCLC [OR = 1.51 (1.18–1.93 at 95% CI); p value = 0.0009]. Genetic association was also explored by applying different genetic models (Dominant, Additive). These results suggest that rs2494938 polymorphism of the LRFN2 gene is a risk factor in the North Indian populations to develop NSCLC. The LD (Linkage Disequilibrium) plot demonstrates the variant and its LD SNPs (r2 > 0.8) and the variant has direct regulatory effect, which could affect the overall physiology of the gene. These findings could be used as diagnostic and prognostic markers in clinical studies of lung cancer patients in North Indian population groups. The present study also provides an important evidence on the genetic etiology of NSCLC in North Indian populations and further expounds GWAS findings on the role of LRFN2 in lung cancer risk. This study provides the holistic view about the non-small cell lung cancer in Jammu and Kashmir, North Indian population and it can be a hallmark of cancer if verified on a very large sample size (cohort).

Similar content being viewed by others
References
Arnold M, Raffler J, Pfeufer A, Suhre K, Kastenmüller G (2014) SNiPA: an interactive, genetic variant-centered annotation browser. Bioinformatics 31:1334–1336
Castellanos A, Lang G, Frampton J, Weston K (2007) Regulation of erythropoiesis by the neuronal transmembrane protein Lrfn2. Exp Hematol 35:724–734
Dhar GM, Shah GN, Naheed B, Hafiza A (1993) Epidemiological trend in the distribution of cancer in Kashmir Valley. J Epidemiol Community Health 47:290–292
Fernandes HB, Baimbridge KG, Church J, Hayden MR, Raymond LA (2007) Mitochondrial sensitivity and altered calcium handling underlie enhanced NMDA-induced apoptosis in YAC128 model of Huntington's disease. J Neurosci 27:13614–13623
Ganesh B, Sushama S, Monika S, Suvarna P (2011) A case-control study of risk factors for lung cancer in Mumbai. India Asian Pac J Cancer Prev 12:357–362
Jemal A, Bray F, Center MM, Ferlay J, Ward E, Forman D (2011) Global cancer statistics. CA Cancer J Clin 61:69–90
Jin G et al (2012) Genetic variants at 6p21. 1 and 7p15. 3 are associated with risk of multiple cancers in Han Chinese. Am J Hum Genet 91:928–934
Kim MS et al (2006) N-methyl-D-aspartate receptor type 2B is epigenetically inactivated and exhibits tumor-suppressive activity in human esophageal cancer. Cancer Res 66:3409–3418
Ko J et al (2006) SALM synaptic cell adhesion-like molecules regulate the differentiation of excitatory synapses. Neuron 50:233–245
Malik PS, Raina V (2015) Lung cancer: Prevalent trends and emerging concepts. Indian J Med Res 141:5
Miko E et al (2011) miR-126 inhibits proliferation of small cell lung cancer cells by targeting SLC7A5. FEBS Lett 585:1191–1196
Nam J, Mah W, Kim E (2011) The SALM/Lrfn family of leucine-rich repeat-containing cell adhesion molecules. Seminars in cell and developmental biology. Elsevier, Amsterdam, pp 492–498
Noronha V, Pinninti R, Patil VM, Joshi A, Prabhash K (2016) Lung cancer in the Indian subcontinent. South Asian J Cancer 5:95
Pandith AA, Siddiqi MA (2012) Burden of cancers in the valley of Kashmir: 5 year epidemiological study reveals a different scenario. Tumor Biol 33:1629–1637
Skeberdis VA et al (2006) Protein kinase A regulates calcium permeability of NMDA receptors. Nat Neurosci 9:501
Stelzer G et al (2016) The GeneCards suite: from gene data mining to disease genome sequence analyses. Curr Protoc Bioinform 54:31–31
Szklarczyk D et al (2015) STRING v10: protein-protein interaction networks, integrated over the tree of life. Nucleic Acids Res 43:D447–D452. https://doi.org/10.1093/nar/gku1003
Tamura H, Suzuki M, Moriya Y, Hoshino H, Okamoto T, Yoshida S, Yoshino I (2011) Aberrant methylation of N-methyl-D-aspartate receptor type 2B (NMDAR2B) in non-small cell carcinoma. BMC Cancer 11:220
The Human Gene Database (1996) The Human Gene Database: GeneCards®: 1996–2018 Weizmann Institute of Science,v4.7.1 Build 10. https://www.genecards.org/cgi-bin/carddisp.pl?gene=LRFN2
Torre LA, Bray F, Siegel RL, Ferlay J, Lortet-Tieulent J, Jemal A (2015) Global cancer statistics, 2012. Cancer J Clin 65:87–108
Verma S et al (2020) Genetic variants of DNAH11 and LRFN2 genes and their association with ovarian and breast cancer. Int J Gynecol Obstet 148:118–122. https://doi.org/10.1002/ijgo.12997
Wang C-Y, Chang K, Petralia RS, Wang Y-X, Seabold GK, Wenthold RJ (2006) A novel family of adhesion-like molecules that interacts with the NMDA receptor. J Neurosci 26:2174–2183
Wu Y-H, Graff RE, Passarelli MN, Hoffman JD, Ziv E, Hoffmann TJ, Witte JS (2018) Identification of pleiotropic cancer susceptibility variants from genome-wide association studies reveals functional characteristics Cancer Epidemiology and Prevention. Biomarkers 27:75–85
Acknowledgements
RK and GRB acknowledge Department of science & technology, Government of India project (DST/SSTP/J&K/459) for financial assistance.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Electronic supplementary material
Below is the link to the electronic supplementary material.
13205_2020_2403_MOESM1_ESM.tif
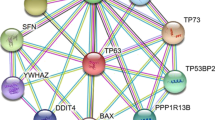
Supplementary file1 Supplementary Fig1: Showing the interaction of LRFN2 with other important genes using String Software (https://string-db.org/) v.11.0. The interaction among the genes depends upon the thickness of the nodes. Greater the thickness of nodes, greater is the relationship among the genes. LRFN2 strongly interacts with DLG3 and DLG4. (TIF 235 kb)
13205_2020_2403_MOESM2_ESM.tif
Supplementary file2 Supplementary figure 2: showing the LD of the genetic variant rs2494938 of LRFN2 using genetic variant centered annotation browser SNIPA 3.0. Linkage disequilibrium plot shows the amount of correlation between a sentinel variant (blue coloured) and its surrounding variants (red coloured). The plot demonstrates the variant and its LD SNP’s (r2 > 0.8) and the variant has direct regulatory effect, which could affect the overall physiology of the gene. (TIF 73 kb)
Rights and permissions
About this article
Cite this article
Bhat, G.R., Verma, S., Bhat, A. et al. Genetic variant rs2494938 of LRFN2 gene is associated with non-small cell lung cancer risk in North-Indian population. 3 Biotech 10, 410 (2020). https://doi.org/10.1007/s13205-020-02403-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s13205-020-02403-1



