Abstract
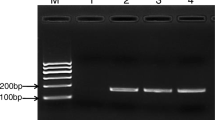
Vibrio parahaemolyticus is part of the natural microflora of estuarine and coastal marine waters and can be also present in seafood, especially shellfish and bivalve molluscs. In this study we compared the reference cultural method ISO 6887-3 with two molecular methods, multiplex PCR and real-time PCR, for the detection of two distinct genetic markers (tlh species-specific gene and tdh virulence gene) of V. parahaemolyticus in bivalve mollusc. The analyses were performed on clams inoculated with V. parahaemolyticus ATCC 43996 at T0 and after a 3 and 6 h of pre-enrichment in alkaline saline peptone water. Counts on agar plates were largely inaccurate, probably due to other Vibrio species grown on the TCBS selective agar. Multiplex PCR assays, performed using primers pairs for tdh and tlh genes, showed a detection limit of 104 CFU/g of shell stock within 6 h of pre-enrichment, respecting however the action level indicated by the National Seafood Sanitation Program guideline. Detection by tdh gene in real-time PCR reached the definitely highest sensitivity in shorter times, 101 CFU/g after 3 h of pre-enrichment, while the sensitivity for the tlh gene was not promising, detecting between 105 and 106 CFU/g after 6 h of pre-enrichment. Our findings provide a rapid routine method of detection of V. parahaemolyticus based on tdh gene by real-time PCR for commercial seafood analysis to identify the risk of gastrointestinal diseases.





Similar content being viewed by others
References
Anonymous (2003) ISO 6887–3: 2003 Microbiology of food and animal feeding stuffs—preparation of test samples, initial suspension and decimal dilutions for microbiological examination—Part 3: Specific rules for the preparation of fish and fishery products. International Organization for Standardization (ISO) 1, Geneva, Switzerland
Baffone W, Tarsi R, Pane L, Campana R, Repetto B, Mariottini GL, Pruzzo C (2006) Detection of free-living and plankton-bound vibrios in coastal waters of the Adriatic Sea (Italy) and study of their pathogenicity-associated properties. Environ Microbiol 8:1299–1305
Barker WHJ (1974) Vibrio parahaemolyticus outbreaks in the United States. Lancet I 303:551–554
Bates TC, Oliver JD (2004) The viable but nonculturable state of Kanagawa positive and negative strains of Vibrio parahaemolyticus. J Microbiol 42:74–79
Bej AK, Patterson DP, Brasher CW, Vickery MC, Jones DD, Kaysner CA (1999) Detection of total and hemolysin-producing Vibrio parahaemolyticus in shellfish using multiplex PCR amplification of tlh, tdh, and trh. J Microbiol Met 36:215–225
Bisha B, Simonson J, Janes M, Bauman K, Goodrige LD (2012) A review of the current status of cultural and rapid detection of Vibrio parahaemolyticus. Int J Food Sci Technol 47:885–899
Campbell MS, Wright AC (2003) Real-time PCR analysis of Vibrio vulnificus from oysters. Appl Environ Microb 69:7137–7144
Cook DW, Bowers JC, DePaola A (2002) Density of total and pathogenic (tdh +) Vibrio parahaemolyticus in Atlhantic and Gulf coast molluscan shellfish at harvest. J Food Protect 65:1873–1880
Croci L, Suffredini E, Cozzi L, Toti L, Ottaviani D, Pruzzo C, Serratore P, Fischetti R, Goffredo E, Loffredo G, Mioni R (2007) Comparison of different biochemical and molecular methods for the identification of Vibrio parahaemolyticus. J Appl Microbiol 102:229–237
DePaola A, Nordstrom J, Bowers C, Wells J, Cook D (2003) Seasonal abundance of total and pathogenic Vibrio parahaemolyticus in Alabama oysters. Appl Environ Microbiol 69:1521–1526
Deter J, Lozach S, Veron A, Chollet J, Derrien A, Hervio-Heath D (2010) Ecology of pathogenic and non-pathogenic Vibrio parahaemolyticus on the French Atlantic coast. Effects of temperature, salinity, turbidity and chlorophylla. Environ Microbiol 12:929–937
Dey MM (2015) World and U.S. demand and supply relationships for seafood: implications for aquaculture producers. Aquac Mag 41(2):44–45
Diana JS (2009) Aquaculture production and biodiversity conservation. J Biosci 59:27–38
Eddabra R, Piemont Y, Scheftel JM (2011) Evaluation of a new chromogenic medium, chromID Vibrio, for the isolation and presumptive identification of Vibrio cholerae and Vibrio parahaemolyticus from human clinical specimens. Eur J Clin Microbiol 30:733–737
FAO (2015) Fishery statistical collections: global capture production. Food and Agriculture Organization of the United Nations, Rome
Garrido A, Chapela MJ, Ferreira M, Atanassova M, Fajardo P, Lago J, Vieites JM, Lago J, Vieites JM, Cabado AG (2012) Development of a multiplex real-time PCR method for pathogenic Vibrio parahaemolyticus detection (tdh + and trh +). Food Control 24:128–135
Garrido-Maestu A, Chapela MJ, Peñaranda E, Vieites JM, Cabado AG (2014) In-house validation of novel multiplex real-time PCR gene combination for the simultaneous detection of the main human pathogenic vibrios (Vibrio cholerae, Vibrio parahaemolyticus, and Vibrio vulnificus). Food Control 37:371–379
Hayat-Mahmud Z, Kassu A, Mohammad A, Yamato M, Bhuiyan NA, BalakrishNair G, Ota F (2006) Isolation and molecular characterization of toxigenic Vibrio parahaemolyticus from the Kii Channel, Japan. Microbiol Res 161:25–37
He P, Chen Z, Luo J, Wang H, Yan Y, Chen L, Gao W (2014) Multiplex real-time PCR assay for detection of pathogenic Vibrio parahaemolyticus strains. Mol Cell Probe 28:246–250
Honda T, Iida T (1993) The pathogenicity of Vibrio parahaemolyticus and the role of thermostable direct haemolysin and related haemolysins. Rev Med Microbiol 4:106–113
Honda S, Goto I, Minematsu I, Ikeda I, Asano N, Ishibashi M, Kinoshita Y, Nishibuchi M, Honda T, Miwatani T (1987a) Vibrio parahaemolyticus infectious disease caused by Kanagawa phenomenon-negative O3:K6 originated from Maldives. Jpn J Infect Dis 61:1070–1078
Honda S, Goto I, Minematsu I, Ikeda I, Asano N, Ishibashi M, Kinoshita Y, Nishibuchi M, Honda T, Miwatani T (1987b) Gastroenteritis due to Kanagawa negative Vibrio parahaemolyticus. Lancet 29:331–332
Jones JL, Ludeke CM, Bowers JC, Garrett N, Fischer M, Parsons MB, Bopp CA, DePaola A (2012) Biochemical, serological, and virulence characterization of clinical and oyster Vibrio parahaemolyticus isolates. J Clin Microbiol 51:2343–2352
Kim JY, Lee JL (2014) Multipurpose assessment for the quantification of Vibrio spp. and total bacteria in fish and seawater using multiplex real-time polymerase chain reaction. J Sci Food Agric 94:2807–2817
Kim YB, Okuda J, Matsumoto C, Takahashi N, Hashimoto S, Nishibuchi M (1999) Identification of Vibrio parahaemolyticus strains at the species level by PCR targeted to the toxR gene. J Clin Microbiol 37:1173–1177
Koh EG, Huyn JH, LaRock PA (1994) Pertinence of indicator organisms and sampling variables to Vibrio concentrations. Appl Environ Microbiol 60:3897–3900
Lee SKY, Wang HZ, Law SHW, Wu RSS, Kong RYC (2002) Analysis of the 16S–23S rDNA intergenic spacers (IGSs) of marine vibrios for species-specific signature DNA sequences. Mar Pollut Bull 44:412–420
Leiva GE, Castilla JC (2002) A review of the world marine gastropod fishery: evolution of catches, management and the Chilean experience. Rev Fish Biol Fish 11:283–300
Lin ZK, Kumagai K, Baba J, Mekalanos J, Nishibuchi M (1993) Identification of Vibrio parahaemolyticus strains at the species level by PCR targeted to the toxR gene. J Clin Microbiol 37:1173–1177
Lin YT, Labbe RG, Shetty K (2005) Inhibition of Vibrio parahaemolyticus in seafood systems using oregano and cranberry phytochemical synergies and lactic acid. Innov Food Sci Emerg 6:453–458
Liu B, He X, Chen W, Yu S, Shi C, Zhou X, Chen J, Wang D, Shi X (2012) Development of a real time PCR assay for rapid detection of Vibrio parahaemolyticus from seafood. Protein Cell 3:204–212. https://doi.org/10.1007/s13238-012-2017-6
Messelhäusser U, Colditz J, Thärigen D, Kleih W, Höller C, Busch U (2010) Detection and differentiation of Vibrio spp. in seafood and fish samples with cultural and molecular methods. Int J Food Microbiol 142:360–364
Naylor RL, Goldburg RJ, Primavera JH, Kautsky N, Beveridge MCM, Clay J et al (2000) Effect of aquaculture on world fish supplies. Nature 405:1017–1024
Nishibuchi M, Kaper JB (1995) Thermostable direct haemolysin gene of Vibrio parahaemolyticus: a virulence gene acquired by a marine bacterium. Infect Immun 63:2093–2099
Nordstrom JL, Vickery MCL, Blackstone GM, Murray SL, DePaola A (2007) Development of a multiplex real-time PCR assay with an internal amplification control for the detection of total and pathogenic Vibrio parahaemolyticus bacteria in oysters. Appl Environ Microbiol 73:5840–5847
NSSP: National Shellfish Sanitation Program (1997) Sanitation of the harvesting, processing and distribution of shellfish. In: Manual of Operations Part II, U.S. Department of Health and Human Services, Food and Drug Administration, Washington, DC
O’Hara CM, Sowers EG, Bopp CA, Duda SB, Strockbine NA (2003) Accuracy of six commercially available systems for identification of members of the family Vibrionaceae. J Clin Microbiol 41:5654–5659
Oliver JD, Nilsson L, Kjelleberg S (1991) Formation of nonculturable vulnificus cells and its relationship to the starvation state. Appl Environ Microbiol 57:2640–2644
Park JY, Jeon S, Kim JY, Park M, Kim S (2013) Multiplex real-time polymerase chain reaction assays for simultaneous detection of Vibrio cholerae, Vibrio parahaemolyticus, and Vibrio vulnificus. Osong Public Health Res Perspect 4:133–139
Raghunath P, Acharya S, Bhanumathi A, Karunasagar I (2008) Detection and molecular characterization of Vibrio parahaemolyticus isolated from seafood harvested along the southwest coast of India. Food Microbiol 25:824–830
Rizvi AV, Bej AK (2010) Multiplexed real-time PCR amplification of tlh, tdh and trh genes in Vibrio parahaemolyticus and its rapid detection in shellfish and Gulf of Mexico water. Antonie Van Leeuwenhoek 98:279–290. https://doi.org/10.1007/s10482
Robert-Pillot A, Guenole A, Lesne J, Delesmont R, Fournier JM, Quilici ML (2004) Occurrence of the tdh and trh genes in Vibrio parahaemolyticus isolates from waters and raw shellfish collected in two French coastal areas and from seafood imported into France. Int J Food Microbiol 91:319–325
Rosec JP, Causse V, Cruz B, Rauzier J, Carnat L (2012) The international standard ISO/TS 21872-1 to study the occurence of total and pathogenic Vibrio parahaemolyticus and Vibrio cholerae in seafood: its improvement by use of a chromogenic medium and PCR. Int J Food Microbiol 157:189–194
Shen XS, Cai YQ, Liu CC, Liu WW, Hui YH, Su YC (2009) Effect of temperature on uptake and survival of Vibrio parahaemolyticus in oysters (Crassostrea plicatula). Int J Food Microbiol 136:129–132
Suffredini E, Cozzi L, Ciccaglioni G, Croci L (2014) Development of a colony hybridization method for the enumeration of total and potentially enteropathogenic Vibrio parahaemolyticus in shellfish. Int J Food Microbiol 186:22–31
Tall A, Teillon A, Boisset C, Delesmont R, Touron-Bodilis A, Hervio-Heath D (2012) Real-time PCR optimization to identify environmental Vibrio spp. strains. J Appl Microbiol 113:361–372
Venkateswaran K, Dohmoto N, Harayama S (1998) Cloning and nucleotide sequence of the gyrB gene of Vibrio parahaemolyticus and its application in detection of this pathogen in shrimp. Appl Environ Microbiol 64:681–687
Wong HC, Wang P, Chen SY, Chiu SW (2004) Resuscitation of viable but non-culturable Vibrio parahaemolyticus in a minimum salt medium. FEMS Microbiol Lett 233:269–275
Zhang XH, Austin B (2005) Haemolysins in Vibrio species. J Appl Microbiol 98:1011–1019
Zhu RG, Li TP, Jia YF, Song LF (2012) Quantitative study of viable Vibrio parahaemolyticus cells in raw seafood using propidium monoazide in combination with quantitative PCR. J Microbiol Meth 90:262–266
Acknowledgements
The Authors would like to thank MARE. A s.r.l. (Cattolica, FC, Italy) to have provided the clam samples.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Federici, S., Serrazanetti, D.I., Guerzoni, M.E. et al. Development of a rapid PCR protocol to detect Vibrio parahaemolyticus in clams. J Food Sci Technol 55, 749–759 (2018). https://doi.org/10.1007/s13197-017-2986-9
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13197-017-2986-9