Abstract
Purpose
The research endeavors to explore the implications of CD47 in cancer immunotherapy effectiveness. Specifically, there is a gap in comprehending the influence of CD47 on the tumor immune microenvironment, particularly in relation to CD8 + T cells. Our study aims to elucidate the prognostic and immunological relevance of CD47 to enhance insights into its prospective utilities in immunotherapeutic interventions.
Methods
Differential gene expression analysis, prognosis assessment, immunological infiltration evaluation, pathway enrichment analysis, and correlation investigation were performed utilizing a combination of R packages, computational algorithms, diverse datasets, and patient cohorts. Validation of the concept was achieved through the utilization of single-cell sequencing technology.
Results
CD47 demonstrated ubiquitous expression across various cancer types and was notably associated with unfavorable prognostic outcomes in pan-cancer assessments. Immunological investigations unveiled a robust correlation between CD47 expression and T-cell infiltration rather than T-cell exclusion across multiple cancer types. Specifically, the CD47-high group exhibited a poorer prognosis for the cytotoxic CD8 + T cell Top group compared to the CD47-low group, suggesting a potential impairment of CD8 + T cell functionality by CD47. The exploration of mechanism identified enrichment of CD47-associated differentially expressed genes in the CD8 + T cell exhausted pathway in multiple cancer contexts. Further analyses focusing on the CD8 TCR Downstream Pathway and gene correlation patterns underscored the significant involvement of TNFRSF9 in mediating these effects.
Conclusion
A robust association exists between CD47 and the exhaustion of CD8 + T cells, potentially enabling immune evasion by cancer cells and thereby contributing to adverse prognostic outcomes. Consequently, genes such as CD47 and those linked to T-cell exhaustion, notably TNFRSF9, present as promising dual antigenic targets, providing critical insights into the field of immunotherapy.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
The advent of immunotherapies has brought about significant advancements in the survival rates of cancer patients. The upregulation of various immune checkpoints serves as a key mechanism facilitating tumor immune evasion and represents a primary barrier that hampers the effectiveness of immune-based treatments. CD47, a pivotal checkpoint of innate immunity, emerges as a focal point of investigation. Delving into the intricate interplay between CD47 and the tumor immune microenvironment (TIME) to delineate a comprehensive understanding of tumor immune evasion pathways holds promise for the development of tailored therapeutic interventions and the identification of novel immune targets. These endeavors are poised to address challenges associated with immune therapy resistance, ultimately enhancing the survival outcomes of individuals afflicted with malignancies.
CD47 is a transmembrane glycoprotein belonging to the immunoglobulin superfamily, known to interact with signal regulatory protein α(SIRPα). The CD47/SIRPα axis functions to impede myosin accumulation, thereby initiating the "Do not eat me" signal to evade phagocytosis by macrophages [1, 2] and suppress innate immunity [3,4,5]. Overexpression of CD47 in various tumor cell types has been identified as a mechanism for evading innate immunity[4], correlating with diminished survival outcomes and reduced responsiveness to conventional therapies[6,7,8]. Targeting the CD47/SIRPα axis has been a focal point of investigation, with therapeutic strategies including monoclonal antibodies (McAb), bispecific antibodies, fusion proteins, combination chemotherapies, and immunotherapies, such as CD47 McAb and fusion proteins incorporating SIRPα(e.g., TTI-621/2). These approaches have demonstrated notable clinical efficacy and are currently under evaluation in phase II or III clinical trials[9,10,11].
Significantly, the CD47/SIRPα axis exerts influence on adaptive immunity as well. Studies have revealed that inhibition of the CD47/SIRPα axis can directly enhance T-cell responses or act through modulation of myeloid cells [12, 13]. Notably, a subset of CD8 + T cells has been identified to express SIRPα[14], and disrupting the CD47/SIRPα interaction can overcome resistance to CD8-mediated immunotherapy [15, 16]. Additionally, CD47 agonists have demonstrated the ability to promote antigen presentation and facilitate cross-priming of T-lymphocytes [17, 18]. Preclinical mouse models have underscored the role of CD47 monoclonal antibodies in T-cell cross-priming, with a lack of therapeutic response observed in T-cell-deficient mice but rescued in wild-type counterparts [19]. Combining radiotherapy with CD47 blockade has been shown to enhance antitumor immunity by directly impacting CD8 + T cells [20]. The intricate interplay between the tumor microenvironment (TME), tumor immune evasion, cancer prognosis, and therapeutic responses has been elucidated [21, 22], highlighting the significance of targeting the CD47/SIRPα axis to impede phagocytosis and counteract innate immunity checkpoints associated with tumor immune evasion. Nonetheless, a comprehensive understanding of the implications of adaptive immunity in regulating the CD47 checkpoint and its impact on antitumor T-cell immunity remains an imperative area for further exploration.
A promising target of interest is the tumor necrosis factor (TNF) receptor superfamily member TNFRSF9 (CD137, TNFRSF9). TNFRSF9 expression is specifically induced through the interaction between the T cell receptor (TCR) and the major histocompatibility complex (MHC) [23]. Identified as a characteristic marker of tumor-reactive T cell subsets within the tumor microenvironment (TME), TNFRSF9 is notably absent on static T cells present in peripheral blood [24, 25]. DSP107, a fusion protein, exhibits dual immune regulatory capabilities by binding to both targets, thereby stimulating innate and adaptive immune responses and showcasing potent antitumor effects. Through the natural trimerization of DSP107 via the 4-1BBL trimerization domain, binding to CD47 on cancer cells disrupts the CD47-SIRPα interaction. This interaction also facilitates the immobilization of DSP107 on the cancer cell surface, enabling the delivery of the 4-1BBL-4-1BB costimulatory signal to T cells localized within the tumor microenvironment. The dual immunomodulatory mechanism of DSP107 is strategically formulated to activate both innate and adaptive immune responses at the tumor site, ultimately enhancing antitumor immunity [26].
This study conducted a comprehensive analysis of the prognostic and immunological implications of CD47 in pan-cancer, with a specific focus on its association with CD8 + T cell exhaustion. The investigation aimed to elucidate the potential impact of CD47 interaction with TNFRSF9 within the tumor immune microenvironment (TIME) on CD8 + T cell functionality and its consequent effect on the prognosis of patients with malignant tumors. Leveraging a range of bioinformatics tools, the analysis encompassed differential expression assessments, prognosis evaluations, immune cell infiltration patterns, and pathway enrichment analyses across various cancer types. Furthermore, the co-expression relationship between CD47 and the pivotal gene TNFRSF9 was validated utilizing single-cell sequencing data. The research design and technical approach are well-structured and reasonable.
2 Methods
2.1 Analysis tools and data collection
XENA-TCGA GTEx
TCGA (https://portal.gdc.cancer.gov/) and GTEx handling were consolidated by the Toil process in UCSC XENA (https://xenabrowser.net/datapages/). Data (V8.0) conversion: Transcripts per million reads format RNAseq data in TPM format and log2 conversion for analysis and comparison. GTEx, The Genotype-Tissue Expression (https://www.gtexportal.org/home/). After log2 transformation, RNAseq data in TPM (transcripts per million reads) format was examined and contrasted. [27]
UALCAN
A comprehensive OMICS cancer data analysis web portal is located in Ualcan (http://ualcan.path.uab.edu/). The expression level of CD47 was normalized as transcript per million reads. P < 0.05 was considered statistically significant [28].
TIMER2.0
TIMER2.0, Tumor IMmune Estimation Resource (https://cistrome.shinyapps.io/timer/) is a database for comprehensive analysis of tumor-infiltrating immune cells [29]. The TIMER database consists of 10897 samples from 32 TCGA cancer types to evaluate immune infiltrate abundance.
TIDE
TIDE [30] (http://tide.dfci.harvard.edu/) stands for Tumor Immune Dysfunction and Exclusion. It is a computational framework developed to evaluate the potential of tumor immune escape from the gene expression profiles of cancer samples. The TIDE score computed for each tumor sample can be a surrogate biomarker to predict response to immune checkpoint blockade, including anti-PD1 and anti-CTLA4 for melanoma and NSCLC. The highly scored genes in TIDE signatures also present potential regulators of tumor immune escape and resistance to cancer immunotherapies [31].
STRING
STRING database [32]: (string-db.org) is a protein interaction network database based on public database and literature information. It gathers several public databases, including UniProt, KEGG, NCBI, and Gene Ontology, to integrate these data and generate a comprehensive protein interaction network database.
TISCH
TISCH collected data from Gene Expression Omnibus (GEO) [33] and Array Express [34] to formulate its scRNA-seq atlas [35], including 79 databases and 2045746 cells from tumor patients and healthy donors. Data sets were processed uniformly to allow for clarification of the TME components at both the single-cell and annotated cluster levels.
HPA[36] (https://www.proteinatlas.org).
The HPA database (Human Protein Atlas) is based on proteomics, transcriptomics, and systems biology data to map tissues, cells, and organs. It includes not only tumor tissue, but also normal tissue protein expression, and can also check the survival curve of tumor patients.
STATISTIC
P value less than 0.05 was statistically significant. Significance markers: NS, P ≥ 0.05; *, p < 0.05; * *, p < 0.01; * * *, p < 0.001.
2.2 Analysis of differential CD47 expression in average, tumor stages, and protein levels
Differential CD47 expression levels between tumors and normal tissues adjacent to TCGA cancer types were analyzed using R (version 3.6.3) and R packages (mainly GGGlot2 [version 3.3.3]) from the XENA-TCGA GTEx resource. Furthermore, Protein levels between tumors and adjacent normal tissues were also investigated using the UALCAN interactive web resource. Survival curves were presented using predictive analysis. SurvMiner [version 0.4.9] and Survival package [version 3.2–10] were used (grouped by p-best). The type of prognosis was OS (Overall Survival), DSS (Disease-Specific Survival), and prognostic data were also obtained from a Cell article [37], and finally verified by immunohistochemistry of HPA database.
2.3 Analysis of tumor immune and immunosuppressive cell infiltration and comparative biomarker analysis
Using the TIMER2 server, we analyzed the correlation between tumor infiltration and CD47 expression, with four immunosuppressive cell types promoting T-cell rejection, MDSCs, CAF, M2-TAM, and Treg across 39 TCGA cancers. The Spearman partial rho value and p < 0.05 were used for correlation analysis. The results of the study used the algorithm with the best positive results. In addition, we used the GSVA R package [version 1.34.0] [38] to explore correlations between the expression of CD47 and the infiltration of 23 types of immune cells [39] in TCGA cancers. Then, the overall predictive power of CD47 was compared with standardized biomarkers of tumor immune response in terms of treatment response outcome and OS, and the correlation analysis of CD47 with other immune checkpoints, MHC class molecules and other immune-related molecules was explored.
2.4 Analysis of CD47 on cytotoxic CD8 + T cell infiltration influenced tumor prognosis
We used the GSVA R package [version 1.34.0] [38, 39] to explore the difference in Cytotoxic T-cell infiltration in the different CD47 expression situations in TCGA pan-cancer. The median method was used to divide patients into CD47- high group and CD47-low group. We also use the TIDE algorithm to assess the effects of CD47 on Cytotoxic CD8 + T cell [31].
2.5 Analysis of pathway
We identify DEG between high and low-expression CD47 clusters using the DESeq2 R package [1.26.0 version] [40]. Division of patients into high and low CD47 groups by median method. These different genes were enriched in the pathway via the R package cluster profile [41, 42] [3.14.3 version] (for SEEA analysis) [42]. [version 3.14.3] (for GSEA analysis) [42]. Reference gene set: H.all.v7.2.symbols.GMT [Hallmarks]. The species is Homo sapiens. Gene set databases came from MSigDB Collections [43]. It included the BIOCARTA subset of CP (browse 292 gene sets); KEGG subset of CP (browse 186 gene sets); PID subset of CP (browse 196 gene sets); REACTOME subset of CP (browse 1615 gene sets); WikiPathways subset of CP (browse 664 gene sets). Significance: It is generally considered that the conditions of False Discovery Rate (FDR) < 0.25 and p.adjust < 0.05. Visualization: GGploT2 [version 3.3.3]. Furthermore, CBNplot [44] was used to investigate molecular regulatory connections. in PID_CD8_TCR_DOWNSTREAM_PATHWAY, exhibiting a Bayesian network inference approach. STRING database was used to detect the PPI (Protein–Protein Interaction Networks), showing the interactions of CD47 protein and core protein of that pathway in biological systems.
2.6 Analysis of co-expression.
We used Software: R (version 3.6.3) to analyze the correlation of CD47 and T-cell exhaustion in TCGA pan-cancer. The data sets were level 3 HTSeq—FPKM RNAseq data format. Visualization of the results is provided by the R package: GGploT2 [version 3.3.3]. Besides, we used TISCH to determine whether CD47 was mainly expressed primarily on CD8 + Tex and whether CD47 has a relationship with the CD8 + Tex gene expressed primarily on CD8 + Tex. The results were presented in the stacked value bar chart (bars of superimposed proportions) to see the expression levels of CD47 and TNFRSF9 in different cohorts through software R (version 3.6.3) and GGPLOT2 [version 3.3.3] (visualization).
3 Results
3.1 Study flowchart
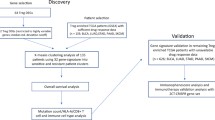
Figure 1 illustrates the schematic diagram of the present study. The study encompasses clinic prognosis and gene data analyses.
The workflow of the study. TCGA, The Cancer Genome Atlas; TME, Tumor microenvironment; TNM, Tumor Node Metastasis; TAMs, tumor-associated macrophages; CAFs, cancer-associated fibroblasts; aDC (activated DC); B cells; CD8 T cells; CTL(Cytotoxic T cells); DC; Eosinophils; iDC (immature DC); Macrophages; Mast cells; Neutrophils; NK CD56bright cells; NK CD56dim cells; NK cells; pDC (Plasmacytoid DC); T cells; T helper cells; Tcm (T central memory); Tem (T effector memory); Tfh (T follicular helper); Tgd (T gamma delta); Th1 cells; Th17 cells; Th2 cells; Treg, regulatory T cells; CAFs, cancer-associated fibroblasts; MDSCs, myeloid-derived suppressor cells; M2-TAMs; M2 subtype of tumor-associated macrophages. MSI, Microsatellite instability; TMB, Tumor mutational burden; CD274, Cluster of differentiation 274; IFNG, interferon-γ; 4-1BB(TNFRSF9), TNF receptor superfamily member 9; CD8 + Tex, exhausted CD8 T lymphocyte cell
3.2 Abnormal expression of CD47 in pan-cancer patients is associated with tumor stages and poor prognosis
We looked into CD47's carcinogenic potential using the XENA-TCGA GTEx pan-cancer database. When comparing nearly all cancer types to normal tissue, we discovered that CD47 gene expression was higher in the former. (ACC, BRCA, BLCA, CHOL, COAD, DLBC, ESCA, GBM, HNSC, KIRC, KIRP, LAML, LGG, LIHC, LUAD, OV, PAAD, PRAD, READ, SARC, SKCM, STAD, THCA, THYM, UCEC, UCS) (Fig. 2a). Additionally, we delved deeper into the CD47 expression of paired samples within the XENA-TCGA database, yielding identical outcomes to the XENA-TCGA GTEx pan-cancer database across various cancer types including BRCA, CHOL, COAD, ESCA, HNSC, KIRC, LIHC, PAAD, STAD, THCA, and UCEC. (Supplementary Fig. 1a).
CD47 is aberrantly overexpressed and is associated with poor cancer prognoses. (A) Boxplots showing differential CD47 expression levels (log2FPKM + 1)/ (log2TPM + 1) between tumors in the XENA-TCGA_GTEx database. Box plots showing differential CD47 expression levels (log2FPKM + 1)/ (log2TPM + 1) between tumor and adjacent normal tissues (Paired Patient) across the TCGA database. CD47 is expressed differently in multiple cancers. B Boxplots illustrating the varying levels of CD47 expression (protein) among tumors in the CPTAC database. C Kaplan–Meier curves of cumulative survival differences between TCGA cancer cohorts with high and those with low expression levels of CD47. The presentation showcases TCGA cancers that exhibit statistically significant variations among the cohorts. UCSC XENA (https://xenabrowser.net/datapages/) by the Toil process unified TCGA RNAseq TPM format data processing. (GTEx)The Genotype-Tissue Expression; Significance representation: ns, p ≥ 0.05; *, p < 0.05; **, p < 0.01; ***, p < 0.001
Significantly, we noted an increase in CD47 protein expression in HNSC, PAAD, UCEC, RCC, and OV compared to the normal levels, as indicated by the UALCAN database (Fig. S2b, Supplementary Fig. 1b). Moreover, in cancer, the expression of CD47 was elevated in advanced tumor stages. For example, patients with M1 stage lung squamous cell carcinoma malignancy expressed more CD47 than patients with M0 stage. The same trend's outcomes were observed in THCA, UCEC, PRAD, KIRC, KIRP, and LIHC. (Supplementary Fig. 1c). Subsequently, we discovered a correlation between excessive CD47 expression and reduced overall survival in ACC, BRCA, LIHC, and KICH; decreased PFI in ACC, LUSC, UVM, and decreased DSS in ACC, LUSC, LGG, and KICH. All of these findings point to CD47 could be an early biomarker for cancer detection, staging, and monitoring. (Fig. 2c, Supplementary Fig. 1d). Lastly, the Human Protein Atlas (HPA) database was used to detect the expression of CD47 in human normal tissues. Representative IHC images of CD47 expression in BRCA, HNSC, LUSC, OV, SKCM. (Supplementary Fig. 1e). From the protein expression level, it was again proved that CD47 was highly expressed in tumor tissues, especially on the cell membranes of tumor cells.
3.3 CD47 is related to tumor immune evasion through infiltration by T lymphocyte cells
Due to its association with tumor immunity evasion, we assessed the associations between CD47 expression levels and the infiltration of MDSCs, CAFs, M2-TAMs, and Treg cells through six algorithms (QUANTISEQ, XCELL, CIBERSORT, CIBERSORT-ABS, TIDE, MCPCOUNTER). These types of immune cells could promote T-cell exclusion. Treg and CAF in BRCA-LumA, Treg and CAF in LICH, Treg, MDSC, and CAF in PRAD, and Treg and CAF in THYM were found to positively correlate with CD47 expression. (r > 0.2, p < 0.05, every cell type ≥ 2 algorithms positive, and at least two types of these cells are positive) (Fig. 3a).
The differential expression of CD47 in tumor microenvironment. A, B The heatmap chart showed correlations of CD47 expression with infiltration by different immune cell types and different immunosuppressive cell types in various TCGA cancer types. C The differential expression of CD47 in Pan-cancer was predominantly associated with CD8 + T cells, CD4 + T cells, DC cells, and macrophages, as demonstrated by the lollipop. Correlation is depicted with a purity-corrected partial (D) Bar plot showing the biomarker relevance of CD47 compared to standardized cancer immune evasion biomarkers in immune checkpoint blockade (ICB) sub-cohorts. The AUC was utilized to assess the predictive efficacy of the test biomarkers in determining the ICB response status. Spearman’s rho values and statistical significance were used. (A. B TIMER database) (C. R3.6.3 ssGSEA). Significance representation: ns, p ≥ 0.05; *, p < 0.05; **, p < 0.01; ***, p < 0.001
Subsequently, we employ the identical cognitive approach to identify the association with T cell infiltration and expression of CD47 through seven different algorithms (QUANTISEQ, XCELL, CIBERSORT, CIBERSORT-ABS, EPIC, MCPCOUNTER, TIMER). The results showed that CD47 expression was positively correlated with infiltration of CD8 + T cell in almost all cancer types. (BLCA, BRCA, BRCA-Basal, COAD, DLBC, ESCA, KIRC, KIRP, LIHC, LUSC, PAAD, PRAD, READ, SKCM, SKCM-Metastasis, STAD, TGCT, THCA, UVM) (r > 0.2, p < 0.05, at least two types of these cells are positive, and at least two types of calculation methods). Interestingly, the infiltration of CD8 + T cell effector memory is comparatively lower in the majority of cancer species compared to other types such as CD8 + T cell central memory and CD8 + T cell naive. As for CD4 + T cell, there were still a lot of CD4 + Tcells closely related to CD47, but it depends on the types of CD4 + T cell. The presence of CD47 in the majority of cancer types was observed to have a positive correlation with the infiltration of CD4 + T cell memory resting and CD4 + T cell Th2, in contrast to CD4 + T cell (non-regulatory) and CD4 + T cell Th1(Fig. 3b).
We further explored which types of T-cell infiltration in tumors were most associated with CD47 using another way. It is worth mentioning that the correlation between CD8 + T cells infiltration and CD47 is the highest in these cancer types (COAD, DLBC, ESCA, HNSC-HPV-, LIHC, LUAD, LUSC, PAAD, STAD) (Fig. 3b). CD47 expression had strong positive correlations with T cell infiltration in various cancer types including BLCA, CHOL, COADREAD, DLBC, GBM, HNSC, KIRC, KIRP, LAML, LUSC, PAAD, PRAD, SKCM, TGCT, READ, STAD, UCS, and UVM. The T cell subtypes that displayed significant associations included T helper cells, CD8 + T cells, CD4 T cells, Cytotoxic cells, Th1, Th2, Th17, as well as Tgd, Tcm, Tem, and TFH. (Fig. 3c, Supplementary Fig. 2a). Furthermore, it suggested a strong association between the infiltration of CD8 + T cells and CD47 in numerous cancer types (DLBC, ESCA, LUSC, OV, SKCM, STAD, TGCT, THCA, UCS, UVM). (r > 0.2, p < 0.05).
Then, we assessed CD47 biomarker relevance by comparing CD47 with standardized biomarkers based on its response outcomes to ICB sub-cohorts and OS predictive ability. Interestingly, we found that in 16 of the 25 ICB sub-cohorts, CD47 alone had an area greater than 0.5% under the AUC. CD47 was predicted to be more valuable than TMB, T. Clonality, B. Clonality, and MSI. Seven, nine, seven, and 13 ICB subgroups had more significant AUC values than 0.5. However, CD47 is lower than CD274, TIDE, IFNG, CD8, and Merck18. Based on these results, it is strongly indicated that CD47 plays a pivotal role in the immune microenvironment of tumors and exhibits a strong association with T-cell infiltration(Fig. 3d). Lastly, We analysed the correlation of CD47 in pan-cancer with other immune checkpoints including immunoinhibitor molecule, MHCmolecule and immunostimula molecule. We found CD47 had strong correlation with CD274, LAG3, IDO1 and TNF receptor superfamily in pan-cancer(r > 0.2, p < 0.05)( Supplementary Fig. 2b).
3.4 CD47 on CD8 + T cells infiltration had an impact on tumor prognosis.
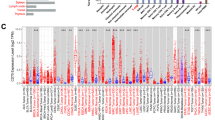
Drawing from our prior findings, it can be inferred that the presence of CD8 + T cells exhibited a strong correlation with CD47. Interestingly, we found that in the CD47 High group, the level of Cytotoxic CD8 + T cell was more frequently observed in BLCA, BRCA, CESC, COAD, COADREAD, ESAD, ESCA, GBM, HNSC, KIRC, LUSC, OV, SKCM, STAD, TGCT, THCA, UCS types compared to the CD47 low group based on TCGA database. (Fig. 4a, Supplemental Fig. 3a) Then we further identified the same conclusion in Multiple immunotherapy cohorts (Nathanson2017_CTLA4-OS, Gide2019_PD1-OS, Gide2019_PD1 + CTLA4-OS, Miao2018_ICB-OS, Mariathasan2018_PDL1-OS, Riaz2017_PD1-OS, Liu2019_PD1-OS Zhao2019_PD1-OS, VanAllen2015_CTLA4-OS) (Supplemental Fig. 3b).
CD47 on CD8 + T cells Infiltration Influenced Tumor Prognosis. A The violin diagram illustrates the disparity in CD47 expression within the infiltration of Cytotoxic CD8 + T-cell. Kaplan–Meier curves (the first picture on the left) of survival ratios as a measure of the BTCGA cohorts C immunotherapeutic response (immune checkpoint blockade) between cancer cohorts D GEO and other cohorts with high and those with low expression levels of CD47. The remainder (the second and third picture on the left) of the graph illustrates the prognosis for the two different sets of CTL expressions with CD47-high group and CD47-low group in above cohorts. Only cancers with statistically significant differences between the cohorts are presented. Significance representation: ns, p ≥ 0.05; *, p < 0.05; **, p < 0.01; ***, p < 0.001
Furthermore, it was observed that the Cytotoxic CD8 + T cell Top group had a poorer prognosis in CD47-high group than the CD47-low group (Fig. 4b). Especially in these cohorts, high Cytotoxic CD8 + T cell infiltration did not suggest a better prognosis. In the same, this phenomenon was also shown in GSE13507@PRECOG Bladder, GSE10886@PRECOG Breast, Roepman Lung Cancer @PRECOG cohorts, GSE17536 Colorectal OS, E-MTAB-3267-Kidney, GSE31684-Bladder, OV GSE31245@PRECOG, GSE49997 OV, METABRIC BreastLumA, Prostate GSE16560@PRECOG, Gide2019-PD1 + CTLA4 Melanomas (Fig. 4c, Supplemental Fig. 3c). The same results happened in KIRC, SARC, LIHC in TCGA database (Fig. 4d). It is well known that T cell dysfunction can negatively impact the prognosis even in the presence of cytotoxic CD8 + T cell, while T cell rejection might negatively impact the prognosis because of the absence of cytotoxic CD8 + T cell infiltration. Consequently, we strongly suggest that CD47 might impair CD8 + T cell function and so negatively impact tumor patients' prognosis.
3.5 CD47 differential genes enrichment in CD8 + Tex pathway.
We categorized the TCGA cohort data into high-expression group and low-expression group, and performed pathway enrichment analysis for both groups of differently expressed genes (p < 0.05). We found that some of the same pathways are present in cancers(Fig. 5a), which includes PID CD8 TCR Downstream Pathway in BLCA, BRCA, ESAD, ESCA, GBM, KIRP, LIHC, LUSC, OV, PRAD, SKCM, STAD, UCS, UVM; PID CD8 TCR Pathway in BLCA, BRCA, ESCA, GBM, LIHC, LUSC, OV, PRAD, SKCM, STAD, TGCT, UCS, UVM; WP T cell Antigen Receptor TCR Signal Pathway in BLCA, BRCA, ESCA, GBM, KIRP, LIHC, LUSC, OV, PRAD, SKCM, STAD, TGCT, UCS, UVM; BIOCARTA CTLA4 Pathway and WP Cancer Immunotherapy By PD1 Blockade in BLCA, BRCA, ESCA, LIHC, LUSC, OV, PRAD, SKCM, STAD, TGCT, UCS, UVM. Furthermore, CD8 + Tcell exhaustion correlated pathways comprise IL2, IL10, IL12, IL17, INF-gamma, T cell, TCR, and JAK-STAT.
CD47 Differential Genes Enriched in CD8 + Tex Pathway. (A) GSEA enrichment analysis results based on CD47 differentially expressed genes in Pan-cancer. (B) The mountain map showcased the path enrichment outcomes, encompassing PID CD8 TCR Downstream Pathway, PID CD8 TCR Pathway, WP Tcell Receptor Signal Pathway, BIOCARTA CTLA4 Pathway, and WP Cancer Immunotherapy by PD1 Blockade, along with the gene regulatory network for the PID CD8 TCR Downstream Pathway. (D) The PPI network demonstrated the correlation between the CD47 protein and the central protein of PID CD8 TCR Downstream Pathway. The representation of significance is as follows: ns, p ≥ 0.05; *, p < 0.05; **, p < 0.01; ***, p < 0.001
Furthermore, mountain maps presented a visualization of the above first 5 pathways enrichment results (Fig. 5b), in BRCA, ESCA, LUSC, OV, and SKCM to further demonstrate the distribution of corresponding numbers of differential genes enriched. The figure illustrates that NES (normalized enrichment score) exhibited positivity, with the majority of the differential genes exhibiting enrichment in the high-expression group. We explore further the gene regulatory network of PID CD8 TCR Downstream Pathway (Fig. 5c, Supplemental Fig. 4a), we discovered that TNFRSF9 and CD8A were expressed at a high level in above all cancers. Taking into account the regulatory networks deduced from the enriched outcomes, we can direct our attention towards a regulatory pathway extending from TNFRSF9 to IL2RA/B/G, via CD8A. Subsequently, we employ PPI(Fig. 5d) to determine the association between the CD47 protein and the proteins linked to the core-enriched molecules. The findings indicated a direct interaction between CD47 protein and CD8A, TNFRSF9, IFNG, B2M, and GZMB. In light of our observations, we deduced that the activation of the CD47 within the signaling pathway could potentially exert a crucial influence on the regulation of CD8 + T cell functionality—a phenomenon that may potentially collaborate with TNFRSF9.
3.6 CD47 expression is related to the CD8 + Tex in pan-cancer.
We delved deeper into the connection between CD47 and the exhaustion of T-cells. The co-expression heat map showed a relationship between CD47 expression in pan-cancer and T-cell exhausted genes. [45] CD274, IDO1, CTLA4, ICOS, TIGIT, IL10, TNFRSF9, HAVCR2 exhibited significant co -expression with CD47 in BLCA, BRCA, CESC, COAD, READ, CRC, ESCA, ESCC, GBM, HNSC, LUSC, OSCC, OV, SKCM, STAD, TGCT, THCA, UCS; NFKB1, GRB2, NFATC3, YY1, NFATC2IP, PRDM1, and FOXO1 in other cancers. Additionally, it was demonstrated that TNFRSF9 ranked among the top three genes in the tumors listed below: BRCA, COAD, ESCA, ESAD, ESCC, GBM, LUSC, OV, and DLBC(r > 0.5 but not the top); whereas CD274 held the top three positions in BLCA, BRCA, CESC, COAD, READ, CRC, LAML, LGG, LUAD, OSCC, PCPG, PRAD, READ, SARC, SKCM, STAD, THCA, UCS (Supplemental Table 1). (Fig. 6a, b, Supplemental 4c).
CD47 expression is related to the CD8 + Tex in many cancer types. A, B Co-expression heat map shows the relationship between the expression of CD47 in pan-cancer and the exhausted genes of T cells, especially TNFRSF9, CD274, IDO1, and ICOS. C The bar chart displayed a wide range of CD47 expression in CD8 + Tex and CD8 + T cell from a single-cell database in a pan-cancerous environment. (D) The histogram illustrates the manifestation of CD47 and TNFRSF on CD8 + Tex.Significance representation: ns, p ≥ 0.05; *, p < 0.05; **, p < 0.01; ***, p < 0.001
After that, we used the single-cell analysis to evaluate the expression of CD47 in CD8 + Tex and CD8 + T cells from the TISCH database and used the embedded bar chart to show the Data distribution. We found CD47 mainly expressed on CD8 + Tex(> 50%) in AML, BRCA, CHOL, CLL, CRC, ESCA, Glioma, HNSC, KICH, LIHC, MCC, MM, NHL, NSCLC, OS, OV, PRAD, SCC, SCLC, SKCM, THCA, UVM (Fig. 6c). And we further detect the expression of TNFRSF9 and CD47. It was observed that both of them exhibited expression on CD8 + Tex in BRCA, BCC, CHOL, CLL, CRC, Glioma, KIRC, LIHC, MCC, NHL, NSCLC, PAAD, SKCM, UCEC, and UVM (Fig. 6d). The findings indicate that the association of CD47 and TNFRSF9 with CD8 + Tex in pan-cancer suggests their involvement in the dysregulation of CD8 + Tex in this type of cancer.
4 Discussion
These findings highlight a strong association between CD47 and the initiation and progression of multiple cancer types. Previous studies have also documented functional associations between CD47 and tumorigenesis, including its role in maintaining immune system homeostasis [2, 46]. Furthermore, our investigation focuses on examining the correlation between CD47 expression levels and various parameters such as prognosis [48, 49], the tumor microenvironment (TME) [50], immune escape mechanisms [51], adaptive immunity [47], and notably, T-cell exhaustion [52], within the pan-cancer landscape of TCGA. The substantial heterogeneity and distinctive clinical characteristics observed across different cancer types and subtypes carry significant implications for further research and clinical applications.
In this study, we investigated the oncogenic role of CD47 within TCGA dataset. Our results reveal that CD47 mRNA or protein levels exhibit notable overexpression across a broad spectrum of TCGA cancers, prominently including BRCA, CHOL, COAD, ESCA, HNSC, KIRC, LIHC, PAAD, STAD, THCA, and UCEC. Furthermore, high levels of CD47 were associated with advanced tumor staging or poorer prognosis in ACC, HNSC, KICH, KIRC, LGG, LICH, LUSC, OV, PAAD, RCC, THCA, UCEC, and UVM. These findings corroborate existing clinical and preclinical evidence highlighting elevated CD47 expression in cancer and its correlation with high-risk tumor features [45, 53,54,55]. Our observations suggest CD47 as a potential biomarker for cancer diagnosis, staging, and post-treatment monitoring.
Subsequently, we evaluated the relationship between the expression levels of CD47 and the infiltration of four immunosuppressive cell populations. Specifically, these cell types, including CAFs, Tregs, M2-TAMs, and MDSCs, have been recognized as biomarkers associated with T-cell exclusion within the tumor microenvironment (TME) [56,57,58,59].The interaction between CAFs and CD47 involves the receptor THBS2/THBS3 on CAFs interacting with CD47 expressed on cancer cells, promoting further cancer progression. Another receptor on the cell membrane of CAFs, MDK, interacts with NCL/SDC2/SDC on cancer cells, potentially serving as therapeutic targets [60]. Regarding the current understanding of the relationship between regulatory T cells (Tregs) and CD47, research has indicated that the ligand of CD47, SIRPγ, is expressed on T cells and varies with differentiation. Some studies have shown that although SIRPγ is expressed on Tregs, it does not participate in their suppressive function [61]. Regarding M2-TAMs, reprogramming TAMs into pro-inflammatory M1 macrophages or inhibiting the M2 polarization of macrophages can disrupt the interaction between tumor cells and macrophages, thereby influencing immune function. Furthermore, it has been reported that the CD47/SIRPα axis plays a crucial role in mediating the interaction involving MDSCs in this context [62]. Inhibition of CXCR2 in G-MDSCs augments the efficacy of CD47 blockade in promoting melanoma tumor cell clearance [63]. Interestingly, we observed a robust correlation between the expression levels of immunosuppressive cells and CD47 in BRCA-LumA, LICH, LUAD, PRAD, and THYM. (≥ 2 immunosuppressive cell types, every cell type ≥ 2 calculated method positive, r > 0.2 and p < 0.05). Therefore, we hypothesize that one of the primary mechanisms through which CD47 regulates tumor immune evasion, tumor progression, and metastasis is through T-cell rejection.
Consequently, our investigation into the association between CD47 and T-cell infiltration utilized seven distinct algorithms (QUANTISEQ, XCELL, CIBERSORT, CIBERSORT-ABS, EPIC, MCPCOUNTER, TIMER, including ≥ 2 immune types, every cell type ≥ 2 calculated method positive, r > 0.2, p < 0.05). These findings demonstrated significant positive correlations in BLCA, BRCA, BRCA-Basal, COAD, DLBC, ESCA, KIRC, KIRP, LIHC, LUSC, PAAD, PRAD, READ, SKCM, SKCM-Metastasis, STAD, TGCT, THCA, UVM. CD47 blockade has been implemented in various models and clinical trials to enhance phagocytosis, augment T cell infiltration into tumors, and reduce tumor burden both in vitro and in vivo [64, 65]. For example, the co-delivery nanocarrier aCD47-DMSN was designed by encapsulating DOX within the mesoporous cavity of MSN and adsorbing aCD47 onto the MSN surface. Following intravenous administration, aCD47-DMSN exhibited robust antitumor efficacy by enhancing the infiltration of CD8 + T cells into the tumor. These results offer another hypothesis that CD47 plays a role in activating dysfunctional T-cell phenotypes to regulate immune escape mechanisms and patient outcomes [66,67,68].
While the prognostic implications of CD47-mediated CD8 + T cell activity are yet to be fully elucidated, our focus was on examining the relationship between CD47 expression and levels of infiltrating cytotoxic CD8 + T cells. Analysis of the TCGA database revealed a notable increase in cytotoxic CD8 + T cell infiltration in BLCA, BRCA, CESC, COAD, COADREAD, ESAD, ESCA, GBM, HNSC, KIRC, LUSC, OV, SKCM, STAD, TGCT, THCA, and UCS when comparing the CD47 high and low expression groups. Subsequently, we investigated the potential impact of these findings on treatment response and patient outcomes. Interestingly, no significant differences in prognosis were observed based on CD47 expression status alone. Thus, we further assessed the prognostic implications of cytotoxic CD8 + T cell infiltration in both high and low CD47 expression groups. Notably, in cases where CD47 expression was high, we also explored the prognostic significance of cytotoxic CD8 + T cell infiltration in both the CD47-high and CD47-low subgroups across BLCA, BRCA, COAD, KIRC, LIHC, LUNG CANCER (Adeno, Large, Squamous), OV, PAAD, PRAD, ARC, SKCM, TCGA, PRECOG, and the METABIC cohort. It is widely recognized that the efficacy of immunotherapy is closely associated with the infiltration of cytotoxic CD8 + T cells [66,67,68] (Supplemental Fig. 3a), and good Cytotoxic CD8 + T cell infiltration usually represents a better prognosis [69]. CD47 has been shown to impede the recruitment and activation of effector T cells, leading to intratumoral immunosuppressive effects. By blocking CD47, the suppression of cytotoxic CD8 + T cell function is alleviated, thereby sustaining the anti-tumor response. This blockade operates indirectly by thwarting immunosuppressive signals expressed in antigen-presenting cells or by shielding tumor-infiltrating cytotoxic CD8 + T cells from the local tumor microenvironment's irradiation [20]. Hence, the expression level of CD47 was associated with dysfunctional T-cells in those cohorts.
Then, utilizing TCGA data, we stratified these tumors into high and low-CD47 expression groups and identified differentially expressed genes between the two cohorts based on median CD47 expression levels. These genes were subjected to pathway enrichment analysis to uncover the correlation between high CD47 expression and immunomodulatory effects. High CD47 expression in BLCA, BRCA, ESAD, ESCA, GBM, KIRP, LIHC, LUSC, OV, PRAD, SKCM, STAD, UCS, and UVM was associated with pathways involving interactions among CD8 + T cells, cytokines, and key immunomodulators such as PD-1, CTLA4, IL-10, INF gamma, and cytotoxic CD8 + T cells. These findings suggest that CD47 may play a crucial role in modulating adaptive immunity, particularly in the context of CD8 + T cell exhaustion. Notably, PD-1, a well-established checkpoint pathway present in both immune and tumor cells, was among the pathways implicated in this immunomodulatory network [70]. The blockade of PD-1 signaling influences the TME [71] by initiating an immune response within tumor cells. Pathways induced by PD-1 blockade in cancer immunotherapy have been closely linked to the upregulation of CD47 expression, indicating that elevated CD47 levels post PD-1 inhibitor treatment lead to a reduction in CD8 + T cells, ultimately contributing to drug resistance. Targeting these factors represents a promising adjunctive strategy for patients undergoing anti-programmed death-1 receptor (PD-1)/anti-programmed death-ligand 1 (PD-L1) therapy. Additionally, IL-10 has been demonstrated to contribute to T cell dysfunction. CD8 + T cells play a pivotal role in immune response, metastasis, and tumorigenesis [72]. A notable association was observed between IL-2 levels and the prognosis of patients diagnosed with pan-cancer [73, 74]. However, our analysis did not reveal a significant correlation with T-dysfunction-related pathways in other cancer types. This discrepancy could potentially be attributed to limited data availability or other factors, such as the absence of a key regulatory gene like LAYN that modulates T cell function, or variations in the TME [75].
To enhance comprehension of the regulatory network, we focused on the enriched pathway-PID CD8 TCR DOWNSTREAM PATHWAY. Utilizing Bayesian network inference through CBNplot, we identified heightened expression of TNFRSF9 and established a regulatory pathway connecting CD8A, IL2RA/B/G to TNFRSF9 in BRCA, ESCA, LUSC, OV, and SKCM. This led to the inference that CD47 might modulate the CD8 TCR DOWNSTREAM pathway, inducing CD8 + T cell exhaustion, potentially in conjunction with TNFRSF9. Subsequently, employing protein–protein interaction (PPI) analysis (Fig. 5d), we elucidated a direct interaction between the CD47 protein and the proteins associated with the core-enriched molecules. Furthermore, investigation into the CD47 ligand SIRPA and the PPI network of the pathway core genes revealed its association with IL2, IL2RA, CD8A, B2M, IFNG, and GZMB, thereby substantiating our deduction. (Supplemental Fig. 4b) (all combined score > 0.3).
Additionally, we conducted an analysis of the expression profile and co-expression patterns of the T-cell depletion gene set [43] CD47 across pan-cancer cohorts. Notably, CD274, IDO1, CTLA4, ICOS, TIGIT, IL10, TNFRSF9, and HAVCR2 demonstrated significant co-expression with CD47 in Group 1 (BLCA, BRCA, CESC, COAD, READ, CRC, ESCA, ESCC, GBM, HNSC, LUSC, OSCC, OV, SKCM, STAD, TGCT, THCA, UCS), while NFKB1, GRB2, NFATC3, YY1, NFATC2IP, PRDM1, and FOXO1 exhibited co-expression in other cancer types designated as Group 2. Noteworthy findings included TNFRSF9 ranking among the top three co-expressed genes in BRCA, COAD, ESCA, ESAD, ESCC, GBM, LUSC, OV, and DLBC (with correlation coefficients exceeding 0.5 but not ranking first), and CD274 ranking among the top three in BLCA, BRCA, CESC, COAD, READ, CRC, LAML, LGG, LUAD, OSCC, PCPG, PRAD, READ, SARC, SKCM, STAD, and THCA. These observations suggest potential functional partnerships between these genes and CD47 across various cancer types, a phenomenon also corroborated by previous studies in select cancer types [24, 76, 77].
The TME is widely known for its substantial heterogeneity [78]. Utilizing the TISCH single-cell database, we explored the impact of CD47 on the TME. Notable variations in immune cell profiles were observed across primary and metastatic tumor sites. Specifically, our analysis revealed that CD47 was prominently expressed on CD8 + exhausted T cells (CD8 + Tex), exhibiting significantly higher expression levels compared to other CD8 + T cell subsets in AML, BRCA, CHOL, CLL, CRC, ESCA, Glioma, HNSC, KICH, LIHC, MCC, MM, NHL, NSCLC, OS, OV, PRAD, SCC, SCLC, SKCM, THCA, and UVM (Fig. 6c). Furthermore, in BRCA, CHOL, CLL, CRC (COADREAD), GBM, HNSC, KIRC, LIHC, MCC, NHL, NSCLC, PAAD, SCC, SKCM, UCEC, and UVM, both CD47 and TNFRSF9 were co-expressed on CD8 + exhausted T cells. These findings suggest a potential functional partnership between CD47 and TNFRSF9 in regulating CD8 + exhausted T cells, particularly in BRCA, ESCA, LUSC, OV, and SKCM (Supplemental Table 2).
Currently, extensive research efforts are focused on the development of dual-targeting strategies involving CD47, with notable progress in the development of CD47 and TNFRSF9 targeting fusion protein (DSP107) [24, 79], PD1 and CD47 bispecific fusion molecules, bispecific antibody CD47xPD-L1, CD47 x HER2, CD47xICAM-1, and other approaches [80,81,82,83,84]. These novel strategies not only aim to mitigate the hematologic toxicity and adverse effects associated with CD47 monoclonal antibodies but also seek to enhance the anti-tumor efficacy of CD47 blockade. By elucidating the cell-intrinsic mechanisms regulated by CD47, we gain insights into its role in promoting tumor biology. Collectively, these advancements position CD47 as a promising therapeutic target in the treatment of both solid and non-solid tumors.
5 Limitation
In our study, we encountered negative and opposite findings in certain types of cancers. The majority of the studies utilized public databases available on the internet, which may have contained slightly outdated data. Future research should focus on utilizing real-world cohorts and conducting multicenter data analysis to enhance the robustness and relevance of the findings. Additionally, the small sample size and dearth of reliable data contributed to limitations in the generalizability of the results. Moreover, methodological deficiencies, such as constraints in the technology and instruments used for data collection, further weakened the study's overall findings. The high expression levels of CD47 and TNFRSF9 in cancer cells and their correlation can be demonstrated through multiple immunofluorescence staining. Additionally, utilizing WB-PCR, it can be shown that CD47 wild-type cancer cells express a higher amount of T cell depletion factors compared to CD47 knockout cancer cells. Through T cell co-culture experiments, it can be established that CD47 knockout cancer cells exhibit reduced T cell depletion compared to wild-type cancer cells. The analysis revealed a poor prognosis in the low CD47 expression group in SKCM. Moreover, the expression of CD47 in adjacent tissues was observed to be higher than in cancer tissues, as evidenced in KICH and LUSC. These findings suggest potential factors contributing to these observations, including intrinsic tumor heterogeneity, the tumor immune microenvironment, specifically telomere Tex cell richness [85, 86], or data insufficiency, warranting further experimental validation and investigation.
6 Conclusions
CD47 plays a pivotal role in the TME, prognosis, and immunotherapy not only through its interactions with macrophages but also with tumor-infiltrating T-lymphocyte cells, particularly CD8 + T cells across various cancer types. The interaction between CD47 and TNFRSF9 triggers exhaustion in CD8 + T cells, leading to an unfavorable prognosis. A therapeutic approach targeting both CD47 and TNFRSF9 has the potential to activate both innate and adaptive immune responses, presenting a significant advancement in treatment modalities, especially for patients with BRCA, ESCA, LUSC, OV, and SKCM.
Data availability
The datasets provided in this study can be found in online repositories.
Abbreviations
- 4-1BB:
-
TNFRSF9 (tumor necrosis factor receptor superfamily member 9), CD137, ILA
- ACC:
-
Adrenocortical carcinoma
- ALL:
-
Acute lymphoblastic leukemia
- AUC:
-
Area under curve
- BCC:
-
Basal cell carcinoma
- BIOCARTA:
-
A subset of cancer pathway( icluding 292 gene sets)https://maayanlab.cloud/Harmonizome/dataset/Biocarta
- BLCA:
-
Bladder urothelial carcinoma
- BRCA:
-
Breast invasive carcinoma
- BRCA-Lum A:
-
LumA positive pathological type of breast invasive carcinoma
- BRCA_SRP114962:
-
SRP114962 cohort of non-small cell lung cancer
- CAF:
-
Cancer associated fibroblasts
- CBNplot:
-
Ayesian network plots for enrichment analysis
- CD274:
-
PD-L1 (Programmed death-ligand 1)
- CESC:
-
Cervical squamous cell carcinoma and endocervical adenocarcinoma
- CHOL:
-
Cholangiocarcinoma
- CLL:
-
Chronic lymphocytic leukemia
- COAD:
-
Colon adenocarcinoma
- CP:
-
Cancer pathway
- CRC:
-
Colerectal cancer
- CTL:
-
Cytotoxic T lymphocytes
- CTLA4:
-
Cytotoxic T-lymphocyte-associated protein 4
- DEG:
-
Differential Expression Analysis
- DLBC:
-
Lymphoid neoplasm diffuse large B-cell lymphoma
- DSP107:
-
A Novel Bi-Functional Fusion Protein That Combines Inhibition of CD47 with Targeted Activation of 4-1BB
- DSS:
-
Disease-Specific Survival
- ESAD:
-
Esophageal adenocarcinoma
- ESCA:
-
Esophageal carcinoma
- ESCC:
-
Esophageal cell squamous carcinoma
- FCGR3A:
-
Fc gamma receptor IIIa
- FDR:
-
False discovery rate
- FPKM:
-
Fragments Per Kilobase of exon model per Million mapped fragments
- GBM:
-
Glioblastoma multiforme
- GEO:
-
Gene Expression Omnibus (https://www.ncbi.nlm.nih.gov/geo/)
- GRB2:
-
Growth factor receptor bound protein 2
- GSE:
-
GEO Series
- GTEx:
-
Genotype-Tissue Expression Project (https://commonfund.nih.gov/GTEx)
- HAVCR2:
-
Hepatitis A virus cellular receptor 2
- HER2:
-
Human epidermal growth factor receptor 2
- HTSeq:
-
High-throughput sequence analysis (https://pypi.org/project/HTSeq/)
- ICAM-1:
-
Intercellular Cell Adhesion Molecule-1
- ICB:
-
Immune checkpoint blockade
- ICOS:
-
CD278 (Inducible Co-Stimulator)
- IDO1:
-
Indoleamine 2,3-dioxygenase 1
- IFNG:
-
Interferon γ
- IL:
-
Interleukin
- JAK:
-
Janus kinase
- KEGG:
-
Kyoto Encyclopedia of Genes and Genomes (https://www.genome.jp/kegg/)
- KICH:
-
Kidney chromophobe
- KIRC:
-
Kidney renal clear cell carcinoma
- KIRP:
-
Kidney renal papillary cell carcinoma
- LAML:
-
Acute myeloid leukemia
- LAYN:
-
Layilin
- LGG:
-
Brain lower grade glioma
- LIHC:
-
Liver hepatocellular carcinoma
- LUAD:
-
Lung adenocarcinoma
- LUSC:
-
Lung squamous cell carcinoma
- M2-TAM:
-
M2-tumor-associated macrophages
- McAb:
-
Monoclonal antibody
- MCC:
-
Merkel cell carcinoma
- MDSC:
-
Myeloid-derived suppressor cells
- MESO:
-
Mesothelioma
- METABRIC:
-
Molecular Taxonomy of Breast Cancer International Consortium (https://www.mercuriolab.umassmed.edu/metabric)
- MHC:
-
Main histocompatibility complex
- MSI:
-
Microsatellite instability
- MSigDB:
-
Molecular Signatures Database (https://www.gsea-msigdb.org/gsea/msigdb/)
- MYC:
-
Gene
- NFATC2IP:
-
Nuclear factor of activated T cells 2 interacting protein
- NFATC3:
-
Nuclear factor of activated T cells 3
- NF-κB:
-
Nuclear factor κB
- NHL:
-
Non-Hodgkin lymphoma
- NS:
-
No significance
- NSCLC:
-
Non-small cell lung cancer
- NSCLC_EMTAB6149:
-
EMTAB6149 cohort of non-small cell lung cancer
- OS:
-
Overall survival
- OSCC:
-
Oral squamous cell carcinoma
- OV:
-
Ovarian serous cystadenocarcinoma
- PAAD:
-
Pancreatic adenocarcinoma
- PCPG:
-
Pheochromocytoma and paraganglioma
- PD1:
-
Programmed cell death protein 1
- PFI:
-
Progress Free Interval
- PID:
-
A subset of cancer pathway( icluding 196 gene sets)
- PPI:
-
Protein–Protein Interaction Networks
- PRAD:
-
Prostate adenocarcinoma
- PRDM1:
-
PR domain zinc finger protein 1
- PRECOG:
-
PREsentation and Characterization Of Growth-data (https://precog.stanford.edu/)
- PTEN:
-
Phosphatase and tensin homolog
- REACTOME:
-
A subset of cancer pathway (browse 1615 gene sets) https://reactome.org/
- READ:
-
Pectum adenocarcinoma
- SARC:
-
Sarcoma
- SCC:
-
Squamous cell carcinoma
- SEEA:
-
System of Environmental Economic Accounting
- SIRPα:
-
Signal regulatory protein α
- SKCM:
-
Skin cutaneous melanoma
- STAD:
-
Stomach adenocarcinoma
- STAT:
-
Signal transducer and activator of transcription
- STRING:
-
A database of functional protein association networks
- TCGA:
-
The Cancer Genome Atlas
- Tcm:
-
Central memory T cell
- TCR:
-
T cell receptor
- Tem:
-
Effector memory T cell
- Tex:
-
Exhausted T cell
- Tfh:
-
Follicular helper T cell
- TGCT:
-
Testicular germ cell tumors
- Tgd:
-
Gamma delta T cell
- Th:
-
T helper cells
- THCA:
-
Thyroid carcinoma
- THYM:
-
Thymoma
- TIDE:
-
Tumor immune dysfunction and exclusion
- TIM:
-
Translocation induced circling mutation
- TIME:
-
Tumor immune microenvironment
- TIMER:
-
Tumor immune estimation resource
- TISCH:
-
Tumor Immune Single-cell Hub
- TLR:
-
Toll-like receptors
- TMB:
-
Tumor mutational burden
- TME:
-
Tumor microenvironment
- TNF:
-
Tumor necrosis factor
- TPM:
-
Transcripts per million reads
- Treg:
-
Regulatory T cell
- TTI-621:
-
SIRPα Fc
- UCEC:
-
Uterine corpus endometrial carcinoma
- UCS:
-
Uterine carcinosarcoma
- UVM:
-
Uveal melanoma
- WT:
-
Wild type
- WP:
-
A subset of cancer pathway (icluding 664 gene sets)
- XENA-TCGA GTEx:
-
Xena.ucsc.edu
- YY1:
-
YIN-YANG-1
References
Tsai RK, Discher DE. Inhibition of “self” engulfment through deactivation of myosin-II at the phagocytic synapse between human cells. J Cell Biol. 2008;180:989–1003.
Li D, et al. SLAMF3 and SLAMF4 are immune checkpoints that constrain macrophage phagocytosis of hematopoietic tumors. Sci Immunol. 2022;7:eabj5501.
Okazawa H, et al. Negative regulation of phagocytosis in macrophages by the CD47-SHPS-1 system. J Immunol. 2005;174:2004–11.
Yang H, et al. Engineering macrophages to phagocytose cancer cells by blocking the CD47/SIRPɑ axis. Cancer Med. 2019;8:4245–53.
Andrechak JC, Dooling LJ, Discher DE. The macrophage checkpoint CD47: SIRPα for recognition of “self” cells: from clinical trials of blocking antibodies to mechanobiological fundamentals. Philos Trans R Soc Lond B Biol Sci. 2019;374:20180217.
Bouwstra R, et al. CD47 expression defines efficacy of rituximab with CHOP in non-germinal center B-cell (Non-GCB) diffuse large b-cell lymphoma patients (DLBCL), but not in GCB DLBCL. Cancer Immunol Res. 2019;7:1663–71.
Uger R, Johnson L. Blockade of the CD47-SIRPα axis: a promising approach for cancer immunotherapy. Expert Opin Biol Ther. 2020;20:5–8.
Li Y, et al. Overexpression of CD47 predicts poor prognosis and promotes cancer cell invasion in high-grade serous ovarian carcinoma. Am J Transl Res. 2017;9:2901–10.
Feng R, Zhao H, Xu J, Shen C. CD47: the next checkpoint target for cancer immunotherapy. Crit Rev Oncol Hematol. 2020;152: 103014.
Petrova PS, et al. TTI-621 (SIRPα Fc): a CD47-blocking innate immune checkpoint inhibitor with broad antitumor activity and minimal erythrocyte binding. Clin Cancer Res. 2017;23:1068–79.
Puro RJ, et al. Development of AO-176, a next-generation humanized anti-CD47 antibody with novel anticancer properties and negligible red blood cell binding. Mol Cancer Ther. 2020;19:835–46.
van Duijn A, Van der Burg SH, Scheeren FA. CD47/SIRPα axis: bridging innate and adaptive immunity. J Immunother Cancer. 2022;10:e004589.
Cham LB, et al. Immunotherapeutic blockade of CD47 inhibitory signaling enhances innate and adaptive immune responses to viral infection. Cell Rep. 2020;31: 107494.
Strizova Z, et al. Tumoral and peritumoral NK cells and CD8(+) T cells of esophageal carcinoma patients express high levels of CD47. Sci Rep. 2020;10:13936.
Barkal AA, et al. Engagement of MHC class I by the inhibitory receptor LILRB1 suppresses macrophages and is a target of cancer immunotherapy. Nat Immunol. 2018;19:76–84.
van der Burg SH, Arens R, Ossendorp F, van Hall T, Melief CJ. Vaccines for established cancer: overcoming the challenges posed by immune evasion. Nat Rev Cancer. 2016;16:219–33.
Dheilly E, et al. Tumor-directed blockade of CD47 with bispecific antibodies induces adaptive antitumor immunity. Antibodies. 2018;7:3.
Tseng D, et al. Anti-CD47 antibody-mediated phagocytosis of cancer by macrophages primes an effective antitumor T-cell response. Proc Natl Acad Sci USA. 2013;110:11103–8.
Liu X, et al. CD47 blockade triggers T cell-mediated destruction of immunogenic tumors. Nat Med. 2015;21:1209–15.
Soto-Pantoja DR, et al. CD47 in the tumor microenvironment limits cooperation between antitumor T-cell immunity and radiotherapy. Can Res. 2014;74:6771–83.
Hong W, Xue M, Jiang J, Zhang Y, Gao X. Circular RNA circ-CPA4/ let-7 miRNA/PD-L1 axis regulates cell growth, stemness, drug resistance and immune evasion in non-small cell lung cancer (NSCLC). J Exp Clin Cancer Res. 2020;39:149.
Anand P, et al. Single-cell RNA-seq reveals developmental plasticity with coexisting oncogenic states and immune evasion programs in ETP-ALL. Blood. 2021;137:2463–80.
Chester C, Sanmamed MF, Wang J, Melero I. Immunotherapy targeting 4–1BB: mechanistic rationale, clinical results, and future strategies. Blood. 2018;131:49–57.
Bagheri S, Safaie Qamsari E, Yousefi M, Riazi-Rad F, Sharifzadeh Z. Targeting the 4–1BB costimulatory molecule through single chain antibodies promotes the human T-cell response. Cell Mol Biol Lett. 2020;25:28.
Chu DT, et al. An update on anti-CD137 antibodies in immunotherapies for cancer. Int J Mol Sci. 2019;20:1822.
Cendrowicz E, et al. DSP107 combines inhibition of CD47/SIRPα axis with activation of 4–1BB to trigger anti-cancer immunity. J Exp Clin Cancer Res CR. 2022;41:97.
Vivian J, et al. Toil enables reproducible, open source, extensive biomedical data analyses. Nat Biotechnol. 2017;35:314–6.
Chandrashekar DS, et al. UALCAN: A portal for facilitating tumor subgroup gene expression and survival analyses. Neoplasia. 2017;19:649–58.
Li T et al. TIMER2.0 for analysis of tumor-infiltrating immune cells. Nucleic Acids Res 48, W509-w514 (2020).
Fu J, et al. Large-scale public data reuse to model immunotherapy response and resistance. Genome Med. 2020;12:21.
Jiang P, et al. Signatures of T cell dysfunction and exclusion predict cancer immunotherapy response. Nat Med. 2018;24:1550–8.
Szklarczyk et al. Nucleic acids research 47.D1 (2018): D607-D613.2
Sun D, et al. TISCH: a comprehensive web resource enabling interactive single-cell transcriptome visualization of tumor microenvironment. Nucleic Acids Res. 2021;49:D1420-d1430.
Athar A, et al. ArrayExpress update—from bulk to single-cell expression data. Nucleic Acids Res. 2019;47:D711-d715.
Barrett T, et al. NCBI GEO: archive for functional genomics data sets–update. Nucleic Acids Res. 2013;41:D991-995.
Liu J, et al. An integrated TCGA pan-cancer clinical data resource to drive high-quality survival outcome analytics. Cell. 2018;173:400-416.e411.
Hanzelmann S, Castelo R, Guinney J. GSVA: gene set variation analysis for microarray and RNA-seq data. BMC Bioinf. 2013;14:7.
Bindea G, et al. Spatiotemporal dynamics of intratumoral immune cells reveal the immune landscape in human cancer. Immunity. 2013;39:782–95.
Love MI, Huber W, Anders S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 2014;15:550.
Yu G, Wang LG, Han Y, He QY. clusterProfiler: an R package for comparing biological themes among gene clusters. OMICS. 2012;16:284–7.
Subramanian A, et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc Natl Acad Sci USA. 2005;102:15545–50.
Guo NL, Wan YW. Network-based identification of biomarkers co-expressed with multiple pathways. Cancer Inform. 2014;13:37–47.
Sato N, Tamada Y, Yu G, Okuno Y. CBNplot : Bayesian network plots for enrichment analysis. Bioinformatics. 202 https://doi.org/10.1093/bioinformatics/btac175.
Yu L, et al. Significance of CD47 and its association with tumor immune microenvironment heterogeneity in ovarian cancer. Front Immunol. 2021;12: 768115.
Hayat SMG, et al. CD47: role in the immune system and application to cancer therapy. Cell Oncol. 2020;43:19–30.
Chen SH, et al. Dual checkpoint blockade of CD47 and PD-L1 using an affinity-tuned bispecific antibody maximizes antitumor immunity. J Immunother Cancer. 2021;9:e003464.
Zhang W, et al. Advances in anti-tumor treatments targeting the CD47/SIRPα axis. Front Immunol. 2020;11:18.
Jiang Y, et al. Noninvasive imaging evaluation of tumor immune microenvironment to predict outcomes in gastric cancer. Ann Oncol. 2020;31:760–8.
Pan Y, et al. Single-cell RNA sequencing reveals compartmental remodeling of tumor-infiltrating immune cells induced by anti-CD47 targeting in pancreatic cancer. J Hematol Oncol. 2019;12:124.
Shimizu A, et al. Exosomal CD47 plays an essential role in immune evasion in ovarian cancer. Mol Cancer Res. 2021;19:1583–95.
Jiang TT, et al. Clinical response to anti-CD47 immunotherapy is associated with rapid reduction of exhausted bystander CD4(+) BTLA (+) T cells in tumor microenvironment of mycosis fungoides. Cancers. 2021;13:5982.
Dagogo-Jack I, Shaw AT. Tumour heterogeneity and resistance to cancer therapies. Nat Rev Clin Oncol. 2018;15:81–94.
Yang Y, Yang Z, Yang Y. Potential Role of CD47-Directed Bispecific Antibodies in Cancer Immunotherapy. Front Immunol. 2021;12: 686031.
Yu WB, Ye ZH, Chen X, Shi JJ, Lu JJ. The development of small-molecule inhibitors targeting CD47. Drug Discov Today. 2021;26:561–8.
Hanley CJ, Thomas GJ. T-cell tumor exclusion and immunotherapy resistance: a role for CAF targeting. Br J Cancer. 2020;123:1353–5.
Verzella D, et al. GADD45β loss ablates innate immunosuppression in cancer. Can Res. 2018;78:1275–92.
Vonderheide RH, Bear AS. Tumor-derived myeloid cell chemoattractants and T cell exclusion in pancreatic cancer. Front Immunol. 2020;11: 605619.
Togashi Y, Shitara K, Nishikawa H. Regulatory T cells in cancer immunosuppression - implications for anti-cancer therapy. Nat Rev Clin Oncol. 2019;16:356–71.
Logtenberg MEW, et al. Glutaminyl cyclase is an enzymatic modifier of the CD47- SIRPα axis and a target for cancer immunotherapy. Nat Med. 2019;25:612–9.
Carvalho RF, do Canto LM, Abildgaard C, et al. Single-cell and bulk RNA sequencing reveal ligands and receptors associated with worse overall survival in serous ovarian cancer. Cell Commun Signal. 2022;20(1):176. https://doi.org/10.1186/s12964-022-00991-4.
Dehmani S, Nerrière-Daguin V, Néel M, et al. SIRPγ-CD47 interaction positively regulates the activation of human T cells in situation of chronic stimulation. Front Immunol. 2021. https://doi.org/10.3389/fimmu.2021.732530.
Zhou Y, Qian M, Li J, et al. The role of tumor-associated macrophages in lung cancer: from mechanism to small molecule therapy. Biomed Pharmacother. 2024;170: 116014. https://doi.org/10.1016/j.biopha.2023.116014.
Banuelos A, Zhang A, Berouti H, et al. CXCR2 inhibition in G-MDSCs enhances CD47 blockade for melanoma tumor cell clearance. Proc Natl Acad Sci USA. 2024;121(5): e2318534121. https://doi.org/10.1073/pnas.2318534121.
Chao MP, Alizadeh AA, Tang C, Myklebust JH, Varghese B, Gill S, et al. Anti-CD47 antibody synergizes with rituximab to promote phagocytosis and eradicate non-hodgkin lymphoma. Cell. 2010;142:699–713. https://doi.org/10.1016/j.cell.2010.07.044.
Liu X, Pu Y, Cron K, Deng L, Kline J, Frazier WA, et al. CD47 blockade triggers T cell-mediated destruction of immunogenic tumors. Nat Med. 2015;21:1209–15. https://doi.org/10.1038/nm.3931.
Papalampros A, et al. Unique spatial immune profiling in pancreatic ductal adenocarcinoma with enrichment of exhausted and senescent t cells and diffused CD47-SIRPα expression. Cancers. 2020;12:1825.
Borst J, Ahrends T, Bąbała N, Melief CJM, Kastenmüller W. CD4(+) T cell help in cancer immunology and immunotherapy. Nat Rev Immunol. 2018;18:635–47.
Farhood B, Najafi M, Mortezaee K. CD8(+) cytotoxic T lymphocytes in cancer immunotherapy: a review. J Cell Physiol. 2019;234:8509–21.
Yu Y, et al. Association of long noncoding RNA biomarkers with clinical immune subtype and prediction of immunotherapy response in patients with cancer. JAMA Netw Open. 2020;3: e202149.
Xie F, Xu M, Lu J, Mao L, Wang S. The role of exosomal PD-L1 in tumor progression and immunotherapy. Mol Cancer. 2019;18:146.
Luchini C, et al. ESMO recommendations on microsatellite instability testing for immunotherapy in cancer, and its relationship with PD-1/PD-L1 expression and tumour mutational burden: a systematic review-based approach. Ann Oncol. 2019;30:1232–43.
Yu AI, et al. Gut microbiota modulate CD8 T Cell responses to influence colitis-associated tumorigenesis. Cell Rep. 2020;31: 107471.
Propper DJ, Balkwill FR. Harnessing cytokines and chemokines for cancer therapy. Nat Rev Clin Oncol. 2022;19:237–53.
Lan T, Chen L, Wei X. Inflammatory cytokines in cancer: comprehensive understanding and clinical progress in gene therapy. Cells. 2021;10:100.
Pan JH, et al. LAYN Is a prognostic biomarker and correlated with immune infiltrates in gastric and colon cancers. Front Immunol. 2019;10:6.
Hsieh RC, et al. ATR-mediated CD47 and PD-L1 up-regulation restricts radiotherapy-induced immune priming and abscopal responses in colorectal cancer. Science Immunol. 2022;7:eabl9330.
Lian S, et al. Simultaneous blocking of CD47 and PD-L1 increases innate and adaptive cancer immune responses and cytokine release. EBio Med. 2019;42:281–95.
Biffi G, Tuveson DA. Diversity, and biology of cancer-associated fibroblasts. Physiol Rev. 2021;101:147–76.
A. Saeed et al. Phase 1 dose escalation study of DSP107, a first - in - class CD47 and 4–1BB targeting fusion protein, in combination with atezolizumab in patients with advanced solid tumors. The 2023 ASCO (American Society of Clinical Oncology) Annual Meeting, Chicago, American, June 2–6, 2023; 2632.
H.P. Rui et al. D3L-001, a novel bispecific antibody targeting HER2 and CD47, demonstrates potent preclinical efficacy in solid tumors. AACR (American Association for cancer research) Annual Meeting 2023, Orlando, Florida, April 14–19, 2023; 1873.
X. Chauchet et al. NI-2901, an affinity-optimized CD47xPD-L1 bispecific antibody for dual immune checkpoint blockade. AACR (American Association for cancer research) Annual Meeting 2023, Orlando, Florida, April 14–19, 2023; 2951.
S.M. Liu et al. A novel pegylated bispecific antibody-drug conjugate (P-BsADCpbadc) targeting cancers co-expressing PD-L1 and CD47. AACR (American Association for cancer research) Annual Meeting 2023, Orlando, Florida, April 14–19, 2023; 6307.
M. Ma et al. BSI-508, a novel bispecific fusion molecule targeting PD1 and CD47 for cancer immunotherapy. AACR (American Association for cancer research) Annual Meeting 2023, Orlando, Florida, April 14–19, 2023; 2958.
O.K. Wang et al. VBI-002, a CD47xICAM-1 bispecific antibody for the treatment of hepatocellular carcinoma, melanoma and non-small cell lung cancers. AACR (American Association for cancer research) Annual Meeting 2023, Orlando, Florida, April 14–19, 2023; 6334.
Zheng L, et al. Pan-cancer single-cell landscape of tumor-infiltrating T cells. Science. 2021;374:abe6474.
Zhang Z, et al. Pan-cancer landscape of T-cell exhaustion heterogeneity within the tumor microenvironment revealed a progressive roadmap of hierarchical dysfunction associated with prognosis and therapeutic efficacy. EBio Med. 2022;83: 104207.
Acknowledgements
We extend our gratitude to the public databases utilized in our investigation, whose platform facilitated and assisted in the uploading of their valuable datasets.
Funding
This work was supported by [Science and Technology Program of Guangzhou, China] (Grant numbers [202201011052]).
Author information
Authors and Affiliations
Contributions
HX Liang, Yong Zheng, and ZK Huang were major contributors who conceived the main idea. JC Dai, LT Yao and DP Xie helped design the study; Duo Chen and Hao Li helped with data collection. HL Wang and HR Qiu helped adjust the image format; JH Leng and ZM Tang helped write the manuscript; All authors approved the submitted version.
Corresponding authors
Ethics declarations
Ethics approval and consent to participate
Approval was granted by the Ethics Committee at Guangdong Provincial People’s Hospital. (No KY2023-137-01) The research conducted here adhered to the principles outlined in the Declaration of Helsinki.
Competing interests
All of the authors state that there are no conflicts of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Liang, H., Zheng, Y., Huang, Z. et al. Pan-cancer analysis for the prognostic and immunological role of CD47: interact with TNFRSF9 inducing CD8 + T cell exhaustion. Discov Onc 15, 149 (2024). https://doi.org/10.1007/s12672-024-00951-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s12672-024-00951-z