Abstract
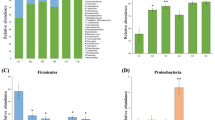
The intestinal microbiota has been identified as a crucial regulator of the overall health, with studies describing its influence in a variety of disorders and developmental processes throughout the body. A widely accepted approach of influencing the microbiota and regulating its functionality in health or disease is the consumption of probiotics. In this study, we aimed to identify the impact of probiotic Lacticaseibacillus casei ATCC393 on the intestinal microbiota of mice and circulating soluble products of microbial origin or the immune system. Investigation of the gut microflora using next-generation sequencing analysis revealed alterations in the microbial populations following consumption of the probiotic. Abundance of taxa classified as Muribaculaceae was increased in lactobacilli-fed animals, while abundance of taxa classified as Lachnospiraceae and Oscillospiraceae was decreased. In addition, the composition of the intestinal microbiota was modified by the administration of L. casei, as evident by the clustering of test subjects when inspecting beta diversity, without however any significant effect on the alpha diversity of the animals. Finally, production of IgA in the intestinal lumen of mice that had received the microorganism was significantly increased, as was the concentration of lactic acid, while levels of acetic acid were noticeably lower in the L. casei group. The findings suggest that L. casei can be considered a potential candidate strain for the modulation of intestinal homeostasis and a component of dietary interventions aiming to improve overall health.






Similar content being viewed by others
Data Availability
All data generated or analysed during this study are available from the corresponding author on request.
References
Rodríguez JM, Murphy K, Stanton C, Ross RP, Kober OI, Juge N, Avershina E, Rudi K et al (2015) The composition of the gut microbiota throughout life, with an emphasis on early life. Microb Ecol Health Dis 26:26050. https://doi.org/10.3402/mehd.v26.26050
Yadav M, Shukla P (2019) Recent systems biology approaches for probiotics use in health aspects: a review. 3 Biotech 9(12):1–10. https://doi.org/10.1007/s13205-019-1980-5
Terpou A, Papadaki A, Lappa IK, Kachrimanidou V, Bosnea LA, Kopsahelis N (2019) Probiotics in food systems: significance and emerging strategies towards improved viability and delivery of enhanced beneficial value. Nutrients 11(7):1591. https://doi.org/10.3390/nu11071591
Erdman SE, Poutahidis T (2017) Gut microbiota modulate host immune cells in cancer development and growth. Free Radic Biol Med 105:28–34. https://doi.org/10.1016/j.freeradbiomed.2016.11.013
Sheflin AM, Borresen EC, Wdowik MJ, Rao S, Brown RJ, Heuberger AL, Broeckling CD, Weir TL et al (2015) Pilot dietary intervention with heat-stabilized rice bran modulates stool microbiota and metabolites in healthy adults. Nutrients 7(2):1282–1300. https://doi.org/10.3390/nu7021282
Raymond F, Ouameur AA, Déraspe M, Iqbal N, Gingras H, Dridi B, Leprohon P, Plante P-L et al (2016) The initial state of the human gut microbiome determines its reshaping by antibiotics. ISME J 10(3):707–720. https://doi.org/10.1038/ismej.2015.148
Bokulich NA, Chung J, Battaglia T, Henderson N, Jay M, Li H, D Lieber A, Wu F et al (2016) Antibiotics, birth mode, and diet shape microbiome maturation during early life. Sci Transl Med 8(343):343ra82–343ra82. https://doi.org/10.1126/scitranslmed.aad7121
Ramos GP, Papadakis KA (2019) Mechanisms of disease: inflammatory bowel diseases. Mayo Clin Proc 94(1):155–165. https://doi.org/10.1016/j.mayocp.2018.09.013
Myles IA (2014) Fast food fever: reviewing the impacts of the Western diet on immunity. Nutr J 13:61. https://doi.org/10.1186/1475-2891-13-61
Minihane AM, Vinoy S, Russell WR, Baka A, Roche HM, Tuohy KM, Teeling JL, Blaak EE et al (2015) Low-grade inflammation, diet composition and health: current research evidence and its translation. Br J Nutr 114(7):999–1012. https://doi.org/10.1017/S0007114515002093
Hill C, Guarner F, Reid G, Gibson GR, Merenstein DJ, Pot B, Morelli L, Canani RB et al (2014) Expert consensus document: the international scientific association for probiotics and prebiotics consensus statement on the scope and appropriate use of the term probiotic. Nat Rev Gastroenterol Hepatol 11(8):506–514. https://doi.org/10.1038/nrgastro.2014.66
Kota RK, Ambati RR, Y V V AK, Srirama K, Reddy PN (2018) Recent advances in probiotics as live biotherapeutics against gastrointestinal diseases. Curr Pharm Des 24(27):3162–3171. https://doi.org/10.2174/1381612824666180717105128
Khangwal I, Shukla P (2019) Prospecting prebiotics, innovative evaluation methods, and their health applications: a review. 3 Biotech 9(5):187. https://doi.org/10.1007/s13205-019-1716-6
Cunningham M, Azcarate-Peril MA, Barnard A, Benoit V, Grimaldi R, Guyonnet D, Holscher HD, Hunter K et al (2021) Shaping the future of probiotics and prebiotics. Trends Microbiol 29(8):1–19. https://doi.org/10.1016/j.tim.2021.01.003
Plaza-Diaz J, Ruiz-Ojeda FJ, Gil-Campos M, Gil A (2019) Mechanisms of action of probiotics. Adv Nutr 10:S49–S66. https://doi.org/10.1093/advances/nmy063
Lebeer S, Bron PA, Marco ML, Van Pijkeren J-P, O’Connell Motherway M, Hill C, Pot B, Roos S et al (2018) Identification of probiotic effector molecules: present state and future perspectives. Curr Opin Biotechnol 49:217–223. https://doi.org/10.1016/j.copbio.2017.10.007
Li S, Xiong Q, Lai X, Li X, Wan M, Zhang J, Yan Y, Cao M et al (2016) Molecular modification of polysaccharides and resulting bioactivities. Compr Rev Food Sci Food Saf 15(2):237–250. https://doi.org/10.1111/1541-4337.12161
Song B, Zheng C, Zha C, Hu S, Yang X, Wang L, Xiao H (2020) Dietary leucine supplementation improves intestinal health of mice through intestinal SIgA secretion. J Appl Microbiol 128(2):574–583. https://doi.org/10.1111/jam.14464
Wu M, Xiao H, Liu G, Chen S, Tan B, Ren W, Bazer FW, Wu G et al (2016) Glutamine promotes intestinal SIgA secretion through intestinal microbiota and IL-13. Mol Nutr Food Res 60(7):1637–1648. https://doi.org/10.1002/mnfr.201600026
Wells JM, Brummer RJ, Derrien M, MacDonald TT, Troost F, Cani PD, Theodorou V, Dekker J et al (2017) Homeostasis of the gut barrier and potential biomarkers. Am J Physiol Gastrointest Liver Physiol 312(3):G171–G193. https://doi.org/10.1152/ajpgi.00048.2015
Planer JD, Peng Y, Kau AL, Blanton LV, Ndao IM, Tarr PI, Warner BB, Gordon JI (2016) Development of the gut microbiota and mucosal IgA responses in twins and gnotobiotic mice. Nature 534(7606):263–266. https://doi.org/10.1038/nature17940
Masahata K, Umemoto E, Kayama H, Kotani M, Nakamura S, Kurakawa T, Kikuta J, Gotoh K et al (2014) Generation of colonic IgA-secreting cells in the caecal patch. Nat Commun 5:3704. https://doi.org/10.1038/ncomms4704
Pabst O, Slack E (2020) IgA and the intestinal microbiota: the importance of being specific. Mucosal Immunol 13(1):12–21. https://doi.org/10.1038/s41385-019-0227-4
Luck H, Khan S, Kim JH, Copeland JK, Revelo XS, Tsai S, Chakraborty M, Cheng K et al (2019) Gut-associated IgA(+) immune cells regulate obesity-related insulin resistance. Nat Commun 10(1):3650. https://doi.org/10.1038/s41467-019-11370-y
Sun Y, Liu Y, Ai C, Song S, Chen X (2019) Caulerpa lentillifera polysaccharides enhance the immunostimulatory activity in immunosuppressed mice in correlation with modulating gut microbiota. Food Funct 10(7):4315–4329. https://doi.org/10.1039/c9fo00713j
Kusumo PD, Bela B, Wibowo H, Munasir Z, Surono IS (2019) Lactobacillus plantarum IS-10506 supplementation increases faecal sIgA and immune response in children younger than two years. Benef Microbes 10(3):245–252. https://doi.org/10.3920/BM2017.0178
Fontana L, Bermudez-Brito M, Plaza-Diaz J, Muñoz-Quezada S, Gil A (2013) Sources, isolation, characterisation and evaluation of probiotics. Br J Nutr 109(SUPPL. 2) https://doi.org/10.1017/S0007114512004011
Kumar M, Nagpal R, Verma V, Kumar A, Kaur N, Hemalatha R, Gautam SK, Singh B (2013) Probiotic metabolites as epigenetic targets in the prevention of colon cancer. Nutr Rev 71(1):23–34. https://doi.org/10.1111/j.1753-4887.2012.00542.x
Ahmadi S, Ghollasi M, Hosseini HM (2017) The apoptotic impact of nisin as a potent bacteriocin on the colon cancer cells. Microb Pathog 111:193–197. https://doi.org/10.1016/j.micpath.2017.08.037
Faghfoori Z, Pourghassem Gargari B, Saber A, Seyyedi M, Fazelian S, Khosroushahi AY (2020) Prophylactic effects of secretion metabolites of dairy lactobacilli through downregulation of ErbB-2 and ErbB-3 genes on colon cancer cells. Eur J cancer Prev Off J Eur Cancer Prev Organ 29(3):201–209. https://doi.org/10.1097/CEJ.0000000000000393
Jacouton E, Chain F, Sokol H, Langella P, Bermúdez-Humarán LG (2017) Probiotic strain Lactobacillus casei BL23 prevents colitis-associated colorectal cancer. Front Immunol 8:1–10. https://doi.org/10.3389/fimmu.2017.01553
Escamilla J, Lane MA, Maitin V (2012) Cell-free supernatants from probiotic Lactobacillus casei and Lactobacillus rhamnosus GG decrease colon cancer cell invasion in vitro. Nutr Cancer 64(6):871–878. https://doi.org/10.1080/01635581.2012.700758
Aindelis G, Chlichlia K (2020) Modulation of anti-tumour immune responses by probiotic bacteria. Vaccines 8(2):329. https://doi.org/10.3390/vaccines8020329
Dimitrellou D, Kandylis P, Kourkoutas Y, Kanellaki M (2017) Novel probiotic whey cheese with immobilized lactobacilli on casein. LWT - Food Sci Technol 86:627–634. https://doi.org/10.1016/j.lwt.2017.08.028
Sidira M, Kandylis P, Kanellaki M, Kourkoutas Y (2016) Effect of curing salts and probiotic cultures on the evolution of flavor compounds in dry-fermented sausages during ripening. Food Chem 201:334–338. https://doi.org/10.1016/j.foodchem.2016.01.084
Abdel-Hamid M, Romeih E, Gamba RR, Nagai E, Suzuki T, Koyanagi T, Enomoto T (2019) The biological activity of fermented milk produced by Lactobacillus casei ATCC 393 during cold storage. Int Dairy J 91:1–8. https://doi.org/10.1016/j.idairyj.2018.12.007
Casas-Solís J, del Huizar-López M, R, Irecta-Nájera CA, Pita-López ML, Santerre A, (2020) Immunomodulatory effect of Lactobacillus casei in a murine model of colon carcinogenesis. Probiotics Antimicrob Proteins 12(3):1012–1024. https://doi.org/10.1007/s12602-019-09611-z
Irecta-Nájera CA, del Rosario H-López M, Casas-Solís J, Castro-Félix P, Santerre A (2017) Protective effect of Lactobacillus casei on DMH-induced colon carcinogenesis in mice. Probiotics Antimicrob Proteins 9(2):163–171. https://doi.org/10.1007/s12602-017-9253-2
Tiptiri-Kourpeti A, Spyridopoulou K, Santarmaki V, Aindelis G, Tompoulidou E, Lamprianidou EE, Saxami G, Ypsilantis P et al (2016) Lactobacillus casei exerts anti-proliferative effects accompanied by apoptotic cell death and up-regulation of TRAIL in colon carcinoma cells. PLoS One 11(2):1–20. https://doi.org/10.1371/journal.pone.0147960
Aindelis G, Tiptiri-Kourpeti A, Lampri E, Spyridopoulou K, Lamprianidou E, Kotsianidis I, Ypsilantis P, Pappa A et al (2020) Immune responses raised in an experimental colon carcinoma model following oral administration of Lactobacillus casei. Cancers (Basel) 12(2):368. https://doi.org/10.3390/cancers12020368
Martin M (2013) Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet. Journal 17(1):10–12. https://doi.org/10.14806/ej.17.1.200
R Core Team. R: a language and environment for statistical computing. Published online 2020. https://www.r-project.org
Callahan BJ, McMurdie PJ, Rosen MJ, Han AW, Johnson AJA, Holmes SP (2016) DADA2: high-resolution sample inference from Illumina amplicon data. Nat Methods 13(7):581–583. https://doi.org/10.1038/nmeth.3869
Quast C, Pruesse E, Yilmaz P, Gerken J, Schweer T, Yarza P, Peplies J, Glöckner FO (2013) The SILVA ribosomal RNA gene database project: improved data processing and web-based tools. Nucleic Acids Res 41(D1):590–596. https://doi.org/10.1093/nar/gks1219
Wright ES (2016) Using DECIPHER v2.0 to analyze big biological sequence data in R. R J 8(1):352–359. https://doi.org/10.32614/rj-2016-025
Schliep KP (2011) Phangorn: phylogenetic analysis in R. Bioinformatics 27(4):592–593. https://doi.org/10.1093/bioinformatics/btq706
McMurdie PJ, Holmes S (2013) Phyloseq: an R package for reproducible interactive analysis and graphics of microbiome census data. PLoS One 8(4):e61217. https://doi.org/10.1371/journal.pone.0061217
Love MI, Huber W, Anders S (2014) Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol 15(12):1–21. https://doi.org/10.1186/s13059-014-0550-8
Benjamini Y, Hochberg Y (1995) Controlling the false discovery rate: a practical and powerful approach to multiple testing. J R Stat Soc Ser B 57(1):289–300. https://doi.org/10.1111/j.2517-6161.1995.tb02031.x
Jalili V, Afgan E, Gu Q, Clements D, Blankenberg D, Goecks J, Taylor J, Nekrutenko A (2020) Corrigendum: the galaxy platform for accessible, reproducible and collaborative biomedical analyses: 2020 update. Nucleic Acids Res 48(14):8205–8207. https://doi.org/10.1093/nar/gkaa554
Wickham H (2016) Ggplot2: elegant graphics for data analysis. Springer-Verlag New York https://ggplot2.tidyverse.org
Rinninella E, Raoul P, Cintoni M, Franceschi F, Miggiano GAD, Gasbarrini A, Mele MC (2019) What is the healthy gut microbiota composition? A changing ecosystem across age, environment, diet, and diseases. Microorganisms 7(1):14. https://doi.org/10.3390/microorganisms7010014
Zitvogel L, Ma Y, Raoult D, Kroemer G, Gajewski TF (2018) The microbiome in cancer immunotherapy: diagnostic tools and therapeutic strategies. Science 359(6382):1366–1370. https://doi.org/10.1126/science.aar6918
Shi CW, Cheng MY, Yang X, Lu YY, Yin HD, Zeng Y, Wang RY, Jiang YL et al (2020) Probiotic Lactobacillus rhamnosus GG promotes mouse gut microbiota diversity and T cell differentiation. Front Microbiol 11:1–16. https://doi.org/10.3389/fmicb.2020.607735
Wang G, Yu Y, Garcia-gutierrez E, Jin X, He Y, Wang L, Tian P, Liu Z et al (2020) Lactobacillus acidophilus JCM 1132 strain and its mutant with different bacteriocin-producing behaviour have various in situ effects on the gut microbiota of healthy mice. Microorganisms 8(1):49. https://doi.org/10.3390/microorganisms8010049
Aktas B, De Wolfe TJ, Tandee K, Safdar N, Darien BJ, Steele JL (2015) The effect of Lactobacillus casei 32G on the mouse cecum microbiota and innate immune response is dose and time dependent. PLoS One 10(12):e0145784–e0145784. https://doi.org/10.1371/journal.pone.0145784
Warda AK, Rea K, Fitzgerald P, Hueston C, Gonzalez-Tortuero E, Dinan TG, Hill C (2019) Heat-killed lactobacilli alter both microbiota composition and behaviour. Behav Brain Res 362:213–223. https://doi.org/10.1016/j.bbr.2018.12.047
Sibai M, Altuntaş E, Yıldırım B, Öztürk G, Yıldırım S, Demircan T (2020) Microbiome and longevity: high abundance of longevity-linked Muribaculaceae in the gut of the long-living rodent Spalax leucodon. OMICS 24(10):592–601. https://doi.org/10.1089/omi.2020.0116
Smith BJ, Miller RA, Ericsson AC, Harrison DC, Strong R, Schmidt TM (2019) Changes in the gut microbiome and fermentation products concurrent with enhanced longevity in acarbose-treated mice. BMC Microbiol 19(1):130. https://doi.org/10.1186/s12866-019-1494-7
Chung YW, Gwak HJ, Moon S, Rho M, Ryu JH (2020) Functional dynamics of bacterial species in the mouse gut microbiome revealed by metagenomic and metatranscriptomic analyses. PLoS One 15(1):1–19. https://doi.org/10.1371/journal.pone.0227886
Qiu L, Tao X, Xiong H, Yu J, Wei H (2018) Lactobacillus plantarum ZDY04 exhibits a strain-specific property of lowering TMAO via the modulation of gut microbiota in mice. Food Funct 9(8):4299–4309. https://doi.org/10.1039/c8fo00349a
Yin X, Yan Y, Kim EB, Lee B, Marco ML (2014) Short communication: effect of milk and milk containing Lactobacillus casei on the intestinal microbiota of mice. J Dairy Sci 97(4):2049–2055. https://doi.org/10.3168/jds.2013-7477
Li S, Qi C, Zhu H, Yu R, Xie C, Peng Y, Yin SW, Fan J et al (2019) Lactobacillus reuteri improves gut barrier function and affects diurnal variation of the gut microbiota in mice fed a high-fat diet. Food Funct 10(8):4705–4715. https://doi.org/10.1039/c9fo00417c
Sun J, Qi C, Zhu H, Zhou Q, Xiao H, Le G, Chen D, Yu R (2019) IgA-targeted Lactobacillus jensenii modulated gut barrier and microbiota in high-fat diet-fed mice. Front Microbiol 10:1–13. https://doi.org/10.3389/fmicb.2019.01179
Yue Y, Xu X, Yang B, Lu J, Zhang S, Liu L, Nassar K, Zhang C et al (2020) Stable colonization of orally administered Lactobacillus casei SY13 alters the gut microbiota. Biomed Res Int 2020:5281639. https://doi.org/10.1155/2020/5281639
Cani PD (2019) Targeting gut microbiota with a complex mix of dietary fibers improves metabolic diseases. Kidney Int 95(1):14–16. https://doi.org/10.1016/j.kint.2018.11.012
Cani PD, Jordan BF (2018) Gut microbiota-mediated inflammation in obesity: a link with gastrointestinal cancer. Nat Rev Gastroenterol Hepatol 15(11):671–682. https://doi.org/10.1038/s41575-018-0025-6
Certo M, Tsai CH, Pucino V, Ho PC, Mauro C (2020) Lactate modulation of immune responses in inflammatory versus tumour microenvironments. Nat Rev Immunol 21(3):151–161. https://doi.org/10.1038/s41577-020-0406-2
Sun S, Li H, Chen J, Qian Q (2017) Lactic acid: no longer an inert and end-product of glycolysis. Physiology 32(6):453–463. https://doi.org/10.1152/physiol.00016.2017
Ratter JM, Rooijackers HMM, Hooiveld GJ, Hijmans AGM, de Galan BE, Tack CJ, Stienstra R (2018) In vitro and in vivo effects of lactate on metabolism and cytokine production of human primary PBMCs and monocytes. Front Immunol 9:2564. https://doi.org/10.3389/fimmu.2018.02564
Ehrlich AM, Pacheco AR, Henrick BM, Taft D, Xu G, Huda MN, Mishchuk D, Goodson ML et al (2020) Indole-3-lactic acid associated with Bifidobacterium-dominated microbiota significantly decreases inflammation in intestinal epithelial cells. BMC Microbiol 20(1):1–13. https://doi.org/10.1186/s12866-020-02023-y
Iraporda C, Romanin DE, Bengoa AA, Errea AJ, Cayet D, Foligné B, Sirard JC, Garrote GL et al (2016) Local treatment with lactate prevents intestinal inflammation in the TNBS-induced colitis model. Front Immunol 7:1–9. https://doi.org/10.3389/fimmu.2016.00651
Lee YS, Kim TY, Kim Y, Lee SH, Kim S, Kang SW, Yang JY, Baek IJ et al (2018) Microbiota-derived lactate accelerates intestinal stem-cell-mediated epithelial development. Cell Host Microbe 24(6):833-846.e6. https://doi.org/10.1016/j.chom.2018.11.002
Schofield WB, Palm NW (2018) Gut microbiota: IgA protects the pioneers. Curr Biol 28(18):R1117–R1119. https://doi.org/10.1016/j.cub.2018.08.019
Cheng Y, Tang S, Huang Y, Liang F, Fang Y, Pan S, Wu T, Xu X (2020) Lactobacillus casei-fermented blueberry pomace augments sIgA production in high-fat diet mice by improving intestinal microbiota. Food Funct 11(7):6552–6564. https://doi.org/10.1039/d0fo01119c
Raya Tonetti F, Arce L, Salva S, Alvarez S, Takahashi H, Kitazawa H, Vizoso-Pinto MG, Villena J (2020) Immunomodulatory properties of bacterium-like particles obtained from immunobiotic lactobacilli: prospects for their use as mucosal adjuvants. Front Immunol 11:1–13. https://doi.org/10.3389/fimmu.2020.00015
Dogi C, García G, Moreno De, de LeBlanc A, Greco C, Cavaglieri L (2016) Lactobacillus rhamnosus RC007 intended for feed additive: immune-stimulatory properties and ameliorating effects on TNBS-induced colitis. Benef Microbes 7(4):539–547. https://doi.org/10.3920/BM2015.0147
Xiong E, Li Y, Min Q, Cui C, Liu J, Hong R, Lai N, Wang Y et al (2019) MZB1 promotes the secretion of J-chain–containing dimeric IgA and is critical for the suppression of gut inflammation. Proc Natl Acad Sci U S A 116(27):13480–13489. https://doi.org/10.1073/pnas.1904204116
Wang P, Li Y, Xiao H, Shi Y, Le G-W, Sun J (2016) Isolation of Lactobacillus reuteri from Peyer’s patches and their effects on sIgA production and gut microbiota diversity. Mol Nutr Food Res 60(9):2020–2030. https://doi.org/10.1002/mnfr.201501065
Harbige LS, Pinto E, Allgrove J, Thomas LV (2016) Immune response of healthy adults to the ingested probiotic Lactobacillus casei Shirota. Scand J Immunol 84(6):353–364. https://doi.org/10.1111/sji.12495
Plaza-Diaz J, Gomez-Llorente C, Campaña-Martin L, Matencio E, Ortuño I, Martínez-Silla R, Gomez-Gallego C, Periago MJ et al (2013) Safety and immunomodulatory effects of three probiotic strains isolated from the feces of breast-fed infants in healthy adults: SETOPROB study. PLoS One 8(10):e78111. https://doi.org/10.1371/journal.pone.0078111
Acknowledgements
The authors would like to thank Associate Professor Nicholas M Glykos for his help during the Bioinformatics analysis and Dr. Anastasios Nikolaou for the analysis of samples in the HPLC studies.
Funding
This research is co-financed by Greece and the European Union (European Social Fund-ESF) through the Operational Programme «Human Resources Development, Education and Lifelong Learning» in the context of the project “Strengthening Human Resources Research Potential via Doctorate Research” (MIS-5000432), implemented by the State Scholarships Foundation (ΙΚΥ).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Ethics Approval
Animal experiments were approved by the Animal Care and Use Committee of the Veterinary Department of Evros Prefecture (license number 4766/28–3-2013) since it complied with the requirements set by Directive 86/609/EEC and PD 160/91.
Conflict of Interest
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
12602_2021_9864_MOESM1_ESM.png
Supplementary file1 (PNG 233 KB): SI Fig. 1 Rarefaction curves of all samples. The sequencing depth of all samples was sufficient to capture most of the diversity
12602_2021_9864_MOESM2_ESM.png
Supplementary file2 (PNG 237 KB): SI Fig. 2 Bar plots representing class and order relative abundance in Lactobacillus or PBS administered mice. Each bar corresponds to the relative abundance of taxa from individual animals. No statistically significant (fdr<0.05) differences were observed
12602_2021_9864_MOESM3_ESM.png
Supplementary file3 (PNG 156 KB): SI Fig. 3 Beta diversity principal coordinates analysis (PCoA) using unweighted UniFrac distance. Differences between the groups were assessed on the inferred ASVs with PERMANOVA (9999 permutations) and were found to not be statistically significant (p>0.05). Indicators represent microbial compositions of individual mice
Rights and permissions
About this article
Cite this article
Aindelis, G., Ypsilantis, P. & Chlichlia, K. Alterations in Faecal Microbiota and Elevated Levels of Intestinal IgA Following Oral Administration of Lacticaseibacillus casei in mice. Probiotics & Antimicro. Prot. 15, 524–534 (2023). https://doi.org/10.1007/s12602-021-09864-7
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12602-021-09864-7