Abstract
Assessing pulmonary lesions using computed tomography (CT) images is of great significance to the severity diagnosis and treatment of coronavirus disease 2019 (COVID-19)-infected patients. Such assessment mainly depends on radiologists’ subjective judgment, which is inefficient and presents difficulty for those with low levels of experience, especially in rural areas. This work focuses on developing a radiomics signature to quantitatively analyze whether COVID-19-infected pulmonary lesions are mild (Grade I) or moderate/severe (Grade II). We retrospectively analyzed 1160 COVID-19-infected pulmonary lesions from 16 hospitals. First, texture features were extracted from the pulmonary lesion regions of CT images. Then, feature preselection was performed and a radiomics signature was built using a stepwise logistic regression. The stepwise logistic regression also calculated the correlation between the radiomics signature and the grade of a pulmonary lesion. Finally, a logistic regression model was trained to classify the grades of pulmonary lesions. Given a significance level of α = 0.001, the stepwise logistic regression achieved an R (multiple correlation coefficient) of 0.70, which is much larger than Rα = 0.18 (the critical value of R). In the classification, the logistic regression model achieved an AUC of 0.87 on an independent test set. Overall, the radiomics signature is significantly correlated with the grade of a pulmonary lesion in COVID-19 infection. The classification model is interpretable and can assist radiologists in quickly and efficiently diagnosing pulmonary lesions.
Graphic Abstract

This work aims to develop a CT-based radiomics signature to quantitatively analyze whether COVID-19-infected pulmonary lesions are mild (Grade I) or moderate/severe (Grade II). The logistic regression model established based on this radiomics signature can assist radiologists to quickly and efficiently diagnose the grades of pulmonary lesions. The model calculates a radiomics score for a lesion and is interpretable and appropriate for clinical use
Similar content being viewed by others
1 Introduction
Coronavirus disease 2019 (COVID-19) has spread rapidly in most countries. As of March 29, 2020, there were 722,088 confirmed cases worldwide. Frontier technologies such as IoMT (internet of medical things) and AI (artificial intelligence) are widely used in the diagnosis, treatment, and prevention of COVID-19 [1, 2]. The common diagnosis of COVID-19 is to use RT-PCR (real-time reverse-transcriptase polymerase chain reaction). In addition, CT (computed tomography) also plays an important role in diagnosing COVID-19 cases. To faster examination, techniques for automated diagnoses, such as methods based on AI with deep learning, have been developed [3,4,5,6,7]. Moreover, assessing whether pulmonary lesions are mild or severe using CT images is of great significance to the severity diagnosis and treatment of COVID-19-infected patients. Although radiologists will make a diagnosis report based on the whole CT images of a patient, assessing lesion regions are mild or severe is still an important part of radiologists’ work. Assessing lesion regions that are mild or severe provides radiologists with more detailed diagnostic information. Currently, this assessment mainly relies on the subjective judgment of radiologists, which is inefficient and presents difficultly for radiologists with low levels of experience, especially in rural areas. Few studies have focused on quantitatively analyzing the grades (mild or moderate/severe) of pulmonary lesions in COVID-19 infection. This work labeled mild lesions as Grade I and labeled moderate or severe lesions as Grade II. We used radiomics-based AI technologies to perform binary classification tasks.
In radiomics, texture as a quantitative feature can provide interpretability [8, 9]. CT textures as radiomics features have been widely used to assist physicians in making decisions on lung diseases. In 2015, Coroller et al. [10] extracted CT-based texture features to predict lung adenocarcinoma metastasis. In 2016, Liu et al. [11] extracted CT-based texture features to analyze a mutation status in lung adenocarcinoma. In 2017, Yip et al. [12] investigated associations between semantic and CT-based texture features of nonsmall cell lung adenocarcinomas.
Given that radiomics-based quantitative assessment is objective and has assisted radiologists in the rapid and accurate diagnosis of lung diseases, radiomics-based AI techniques may be also applicable in the assessment of pulmonary lesions in COVID-19-infected patients. Therefore, this work aims to build a radiomics signature (composed of CT-based texture features) and apply this radiomics signature to quantitatively analyze the grade of pulmonary lesions in COVID-19 infection, including (1) assessing the correlation between the radiomics signature and the grade of a pulmonary lesion and (2) classifying the grades of pulmonary lesions.
2 Materials and Methods
This work is a retrospective study based on the CT images of COVID-19-infected patients and was approved by the Ethics Committee of West China Hospital of Sichuan University (number 2020190). Figure 1 shows the framework of this work.
2.1 Patients and Acquisition of ROIs
Chest CT images of eighty-four COVID-19-infected patients were collected. In total, 1160 pulmonary lesions were retrospectively analyzed. The patients were selected from 16 hospitals in Sichuan Province, China, from January 1, 2020 to February 29, 2020, including 49 males and 35 females. In the female patients, the minimum age, the maximum age, and the median age were 28, 74, and 45.2 years old, respectively. In the male patients, the minimum age, the maximum age and the median age were 20, 76, and 42.4 years old, respectively. All patients were confirmed by RT-PCR examinations and received nonenhanced CT scans. Figure 2 illustrates the inclusion and exclusion of patients and the acquisition of ROIs (regions of interest).
Nine senior radiologists with more than 6 years of experience in chest CT diagnosis at West China Hospital filtered the CT images and delineated the bounding boxes. Given the complexity of the prevalent grading system, two of the radiologists assessed the bounding boxes independently. Discrepancies were solved by discussion or consulting a third radiologist. Briefly, a bounding box with scattered GGOs (ground-glass nodules) was regarded as a mild bounding box (Grade I), and a high-density bounding box with continuous GGOs or even large areas of GGOs was regarded as a moderate or severe bounding box (Grade II) [13, 14]. Figure 3 illustrates delineating bounding boxes from DICOM images. Multiple bounding boxes with overlapping areas were defined as a lesion region. For a lesion region, the bounding box with the largest area was selected and regarded as the ROI. The grade of the selected bounding box was the grade of this ROI. An ROI represented a pulmonary lesion (lesion region). In total, 1160 ROIs were acquired, of which 910 ROIs were Grade I and 250 ROIs were Grade II.
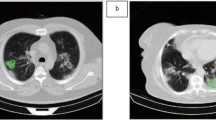
Examples of delineating bounding boxes. a An example with a mild bounding box (Grade I); the patient’s age was 43 years, female. b An example with a moderate bounding box (Grade II); the patient’s age was 36 years, male. c An example with a severe bounding box (Grade II); the patient’s age was 57 years, male
2.2 Texture Feature Extraction
The unit of a pixel value in CT images is HU (Hounsfield unit). This paper omits the unit to express the pixel values concisely. We scaled the pixel values of ROIs to [1 128] based on [− 1000 200], as shown in Eq. (1).
Here, c is a pixel value, s is the scaled value of c, [l h] is [− 1000 200], and [1 n] is [1 128]. In this work, 936 texture features were extracted from each ROI. These 936 features were used as candidate features, including coefficient statistics features, histogram features, gray-level co-occurrence matrix (GLCM) features, gray-level run-length matrix (GLRLM) features, Laplacian of Gaussian (LoG) features, wavelet features, contourlet features, angle cooccurrence matrix (ACM) features, absolute gradient features, autoregression features, and gray-level differential matrix (GLDM) features [15,16,17,18,19,20,21,22,23,24,25]. The value of a feature extracted from an ROI was normalized based on the number of pixels in this ROI. Section A of the supplemental material describes these texture analysis methods in detail.
2.3 Feature Preselection
We performed feature preselection using the least absolute shrinkage and selection operator (LASSO) algorithm on the candidate features [26,27,28]. This work randomly partitioned the dataset into a training set and a test set at a ratio of 7:3. The classification model was built based on the training set and tested using the independent test set. In the training set, 637 ROIs were Grade I, and 175 ROIs were Grade II. In the independent test set, 273 ROIs were Grade I, and 75 ROIs were Grade II. Feature preselection and the subsequent building of radiomics signature were both calculated on the training set. In the implementation of the LASSO algorithm, tenfold cross-validation was used, and those features corresponding to the smallest MSE (mean squared error) value are selected as the pre-selected features.
2.4 Building a Radiomics Signature
Next, we performed stepwise logistic regression on the preselected features to build a radiomics signature. Logistic regression and stepwise logistic regression are both generalized linear regression approaches, and the regression results can be statistically tested and have significant interpretability. Stepwise logistic regression, as the name suggests, will produce a series of models step by step. We used the features in the final model to constitute the radiomics signature.
Equation (2) demonstrates a regression equation, where y = 1 represents that its corresponding ROI is Grade II and y = 0 represents that its corresponding ROI is Grade I. The radiomics signature can be expressed by (x1, x2, …, xm).
To test the linear correlation between y and (x1, x2, …, xm), the following should be tested:
where H0 is the null hypothesis. To test H0, F test can be used. Equation (4) shows the definition of statistic F.
Here, U denotes the regression sum of squares, Q denotes the residual sum of squares, n denotes the number of samples, and R denotes the multiple correlation coefficient used to assess the regression effect. The regression effect represents the quantitative correlation between (x1, x2, …, xm) and y. The closer the value of R is to 1, the stronger is the correlation. Equation (5) defines R.
We can find Fα (the critical value of F, α denotes a significance level) from the F distribution table and calculate Rα (the critical value of R) using Eq. (6). If R > Rα, the regression effect (the correlation) is statistically significant at significance level α.
2.5 Classification
Section 2.3 describes how the dataset was partitioned. We used this partitioning to train a logistic regression model on the training set and classify the lesions in the independent test set. We also conducted a preliminary classification experiment and trained other machine learning models. We calculated the average AUC of the tenfold cross-validation for each model. Considering the interpretability of the models and their AUCs, we chose the logistic regression model to perform a further classification task and constructed a nomogram. Section C of the supplementary material describes the preliminary experiment.
3 Results
The LASSO algorithm preselected 40 features from 936 candidate features. The stepwise logistic regression ultimately selected nine features from these 40 features to constitute the radiomics signature. Table 1 shows the features of the radiomics signature. For more details of the texture analysis methods, please refer to section A of the supplemental material.
Using the nine features, the stepwise logistic regression achieved an R of 0.6996. Given α = 0.001, Fα and Rα were calculated based on Eqs. (4) and (6), respectively.
Using the regression equation shown in Eq. (2), the estimated coefficients (b, β1, β2, …, βm) obtained through the stepwise logistic regression are shown in Table 2. The t test was performed on these estimated regression coefficients. According to Eq. (2) and Table 2, the grade of a pulmonary lesion (i.e., the radiomics score for a lesion) can be expressed by
where y is the grade and (x1, x2, …, x9) is the radiomics signature. We trained a logistic regression model based on the radiomics signature to classify the lesions in the independent test set. Figure 4 depicts the ROC (receiver operating characteristic) curves and AUCs (areas under the ROC curves) of the classification results.
To simultaneously obtain relatively high values of sensitivity and relatively high values of specificity, we adjusted the threshold of the classification output from 0.1 to 0.9. The results are illustrated in Fig. 5.
Figure 5 shows that high sensitivity values and high specificity values appear simultaneously when the threshold is varied from 0.7 to 0.8. Table 3 lists some thresholds and their sensitivity and specificity values.
To assist radiologists in quickly diagnosing patients infected with COVID-19, object detection is also very important. Object detection finds the locations of objects in images and classifies the objects. A YOLO model can perform object detection and classify the detected objects [29]. If a YOLO model can achieve a promised classification performance, it is very appropriate to use the objects detected by the YOLO model as ROIs (lesion regions) and to use the classification results of this YOLO model to classify the grades of ROIs. For comparison with the logistic regression model, we trained a YOLO v3 model to perform one-class object detection (only detecting lesion regions and not classifying the grades of these lesion regions) and the classification of grades (detecting lesion regions and classifying the grades of these lesion regions). The mAP (mean average precision) indicator is frequently used to assess an object detection model in training.
In the experiment involving one-class object detection (only detecting lesion regions and not classifying the grades of these lesion regions), a mAP of 0.81 was achieved in the training, and an accuracy of 0.948 was achieved in the testing. In the experiment involving the classification of grades (detecting lesion regions and classifying the grades of these lesion regions), the training achieved a mAP of 0.050 for Grade I and a mAP of 0.59 for Grade II, and the testing achieved an accuracy of 0.264 for Grade I and an accuracy of 0.836 for Grade II. Table 4 shows the testing results as the threshold varies.
4 Discussion
Assessing whether pulmonary lesions are mild or severe using CT images is of great significance to the severity diagnosis and treatment of COVID-19-infected patients. Currently, this assessment is subjective, which is inefficient and presents difficultly for radiologists with low levels of experience, especially in rural areas. Relatively, AI models can quantitatively and objectively analyze images. Recently, some radiomics-based AI models have been developed for aided diagnosis, efficacy evaluation, or prognosis analysis of COVID-19. Wu et al. [30] developed a CT-based signature to perform prognostic analysis in patients with COVID-19. Fang et al. [31] developed a radiomics model to predict COVID-19 pneumonia. Fu et al. [32] used a machine learning-based tool to develop radiomics signatures and perform prognosis analysis of COVID-19 patients. Ozturk et al. [33] developed a COVID-19 detection model based on X-ray images to diagnosis COVID-19. However, few studies have focused on quantitatively analyzing the grades (mild or moderate/severe) of pulmonary lesions in COVID-19 infection.
Applications of radiomics-based AI models can greatly save time for radiologists in producing image reports and can reduce the workload of radiologists. Radiologists combined with AI models for diagnosis can reduce the possibility of misdiagnosis and missed diagnosis. AI applications can extend the knowledge and experience of senior experts to medical institutions in less developed regions. Unfortunately, many machine learning-based models, including deep learning models, mainly focus on the accuracy, AUC, etc., and rarely pay attention to interpretability. There are two main schemes for radiomics including deep learning and feature engineering combining classic machine learning methods [8, 9]. Deep learning has achieved good results in some image recognition problems and image segmentation problems [34,35,36,37]. But deep learning has substantial difficulties and challenges in AI applications involving small sample sizes, small regions, or expecting interpretability [34, 35]. A deep learning model generally has an N-layer structure. It is difficult to determine which layer is more appropriate for extracting features, and the extracted features are abstract. Although deep learning features are highly versatile, their ability to solve specific problems is relatively weak [38]. In contrast, building an interpretable AI model based on feature engineering is relatively easy. The output of the model is expected to be understood by physicians in clinical applications [39]. We built a nomogram based the logistic regression and used the nomogram to classify ROIs. The nomogram is illustrated Fig. 6.
Nomogram of classifying ROIs and its calibration curve. a Nomogram: for an unknown lesion, a vertical line of xi upward to axis “Points” to assign the score indicating the probability of Grade II. The process is repeated for each variable (from x1 to x9), and the assigned scores are summed. The sum is located on axis “Total Points”, and a vertical line downward to axis “Risk” to find the lesion’s probability of Grade II. b Calibration curve of a: the x-axis represents the nomogram-estimated probabilities and the y-axis represents the observed probabilities. A perfect estimation of an ideal model is represented by the diagonal dotted line. In the diagonal dotted line, the estimated outcome perfectly corresponds to the actual outcome. The performance of a is represented by the solid line. In the solid line, a closer to the diagonal dotted line indicates a better estimation
Figure 6a shows an interpretable classification process. Figure 6b is the calibration curve of Fig. 6a. It visually evaluates the classification performance of Fig. 6a. We also calculated the index of concordance (C-index) to evaluate the classification performance of the nomogram (Fig. 6a). The nomogram achieved a C-index of 0.875. Both the calibration curve and the C-index were calculated based on the independent test set. Figure 6 indicates that the logistic regression model can achieve a promised classification result and its classification process is interpretable. It interprets what drives the identification, and the identification can be quantitatively assessed by a multiple correlation coefficient. As can be seen from Eq. (2), a logistic regression model can clearly express a value called the radiomics score. In future studies, we can use the radiomics score as a factor and combine it with other clinic factors or demographic factors to perform aided diagnoses.
In this work, senior experts in the radiology department of West China Hospital labeled the grades of pulmonary lesions in COVID-19 infected patients. The diagnostic ability of the radiology department of West China Hospital ranks among the best in China’s medical institutions. We quantitatively analyzed the grades of pulmonary lesions in COVID-19 infection using a CT-based radiomics signature. The regression analysis showed that the radiomics signature significantly correlated with the grade of a pulmonary lesion (the value of R was much larger than Rα, α = 0.001), and the trained model achieved the promised AUC for the independent test set. This indicates that the model can be used to assist radiologists, especially those with low levels of experience, in diagnosing the grades of pulmonary lesions in COVID-19 infection.
The grades of pulmonary lesions are considered critical indicators for assessing a patient’s condition in COVID-19 infection or progression, as well as determinants for subsequent treatment strategies. Empirically, patients with more mild pulmonary lesions (Grade I) merely need supportive treatment with close surveillance, while for patients with more moderate or severe pulmonary lesions (Grade II), symptomatic treatment or even ventilator treatment is usually required. Nevertheless, the accurate assessment of the grades of lesions highly relies on the profound knowledge of a radiologist. Although radiologists can also visually confirm ROIs one by one as Grade I or Grade II, it is subjective and is difficult for hospitals in rural areas or radiologists with high workloads. Obviously, our work can greatly improve the diagnosis efficiency, and the calculated aided-diagnosis information is objective.
Texture-based radiomics signatures can deeply mine the heterogeneous data contained in CT images and other medical images at the tissue level and even molecular level [40]. We also performed one-class object detection and the classification of grades using a YOLO v3 deep learning model. However, regarding the classification of grades, the YOLO model yields poor classification of the detected objects, and its testing results are better than the training results. This is because the trained YOLO v3 deep learning model lacks interpretability, so the model’s generalization ability may be poor. Section B of the supplemental material describes more details. However, the experiment involving one-class object detection achieved an accuracy of 0.948. Although the YOLO model can detect the ROIs accurately, it yielded a poor classification of the detected objects (ROIs): It failed to classify the ROIs into Grade I and Grade II. By contrast, the developed logistical regression model yielded a promised result for classifying the ROIs into Grade I and Grade II. This work aimed to develop a radiomics signature and explore an interpretable model, and then use this model to calculate the diagnosis information of lesion grades. However, it can be inferred that combining the one-class object detection of the YOLO v3 model and the logistic regression model developed in this paper may greatly assist radiologists in quickly and efficiently diagnosing COVID-19 pulmonary infections.
This work is a retrospective study of multicenter institutions. There are also some limitations: (1) more samples need to be collected; (2) we use a bounding box to mark a lesion region such that the ROI includes non-lesion regions, which may affect the results of quantitative analysis; (3) this work aimed to give the diagnosis information of lesion grades, we will collect more information to conduct some patient-level classifications for more comprehensive diagnosing in future studies; and (4) although the trained YOLO model yields poor classification on the detected objects, it achieves a high accuracy in object detection; thus, we may combine the object detection of the YOLO model with the logistic model developed in this paper in further works.
5 Conclusion
This work built a CT-based radiomics signature to quantitatively analyze the grades of pulmonary lesions in COVID-19 infection. The experimental results indicated that the developed radiomics signature is significantly correlated with the grade of a pulmonary lesion. The logistic regression model established based on this radiomics signature achieved a promised classification performance for Grade I and Grade II. This result indicated that this model can assist radiologists in quickly and efficiently diagnosing the grades of pulmonary lesions in COVID-19 infection. Furthermore, the nomogram based on the logistic regression model showed an interpretable classification process, which is rewarding for clinical use.
Availability of Data and Material
The datasets during and/or analyzed during the current study are available from the corresponding author on reasonable request with the approval of the institution and trial/study investigators who contributed to the dataset.
References
Singh RP, Javaid M, Haleem A, Vaishya R, Ali S (2020) Internet of Medical Things (IoMT) for orthopaedic in COVID-19 pandemic: roles, challenges, and applications. J Clin Orthop Trauma 11(4):713–717. https://doi.org/10.1016/j.jcot.2020.05.011
Vaishya R, Javaid M, Khan IH, Haleem A (2020) Artificial Intelligence (AI) applications for COVID-19 pandemic. Diabetes Metab Syndrome Clin Res Rev 14(4):337–339. https://doi.org/10.1016/j.dsx.2020.04.012
Wang S, Kang B, Ma J, Zeng X, Xiao M, Guo J, Cai M, Yang J, Li Y, Meng X, Xu B (2020) A deep learning algorithm using CT images to screen for Corona Virus Disease (COVID-19. MedRxiv. https://doi.org/10.1101/2020.02.14.20023028
Mei X, Lee HC, Diao KY, Huang M, Lin B, Liu C, Xie Z, Ma Y, Robson PM, Chung M, Bernheim A, Mani V, Calcagno C, Li K, Li S, Shan H, Lv J, Zhao T, Xia J, Long Q, Steinberger S, Jacobi A, Deyer T, Luksza M, Liu F, Little BP, Fayad ZA, Yang Y (2020) Artificial intelligence–enabled rapid diagnosis of patients with COVID-19. Nat Med 26:1224–1228. https://doi.org/10.1038/s41591-020-0931-3
Chowdhury MEH, Rahman T, Khandakar A, Mazhar R, Kadir MA, Mahbub ZB, Islam KR, Khan MS, Lqbal A, AI-Emadi N, Reaz MBI (2020) Can AI help in screening viral and COVID-19 pneumonia. IEEE Access 8:132665–132676. https://doi.org/10.1109/ACCESS.2020.3010287
Gozes O, Frid-Adar M, Greenspan H, Browning PD, Zhang H, Ji W, Bernheim A, Siegel E (2020) Rapid ai development cycle for the coronavirus (covid-19) pandemic: initial results for automated detection & patient monitoring using deep learning ct image analysis. arXiv preprint arXiv:2003.05037
Butt C, Gill J, Chun D, Babi BA (2020) Deep learning system to screen coronavirus disease 2019 pneumonia. Appl Intell 22:1–7. https://doi.org/10.1007/s10489-020-01714-3
Bi WL, Hosny A, Schabath MB, Giger ML, Birkbak NJ, Mehrtash A, Allison T, Arnaout O, Abbosh C, Dunn LF, Schwartz LH, Gillies RJ, Huang RY, Aerts HJWL (2019) Artificial intelligence in cancer imaging: clinical challenges and applications. CA 69(2):127–157. https://doi.org/10.3322/caac.21552
Gillies RJ, Kinahan PE, Hricak H (2016) Radiomics: images are more than pictures, they are data. Radiology 278(2):563–577. https://doi.org/10.1148/radiol.2015151169
Coroller TP, Grossmann P, Hou Y, Velazquez ER, Leijenaar RTH, Hermann G, Lambin P, Haibe-Kains B, Mak RH, Aerts HJWL (2015) CT-based radiomic signature predicts distant metastasis in lung adenocarcinoma. Radiother Oncol 114(3):345–350. https://doi.org/10.1016/j.radonc.2015.02.015
Liu Y, Kim J, Balagurunathan Y, Li Q, Garcia AL, Stringfield O, Ye Z, Gillies RJ (2016) Radiomic features are associated with EGFR mutation status in lung adenocarcinomas. Clin Lung Cancer 17(5):441–448. e6. https://doi.org/10.1016/j.cllc.2016.02.001
Yip SSF, Liu Y, Parmar C, Li Q, Liu S, Qu F, Ye Z, Gillies RJ, Aerts HJWL (2017) Associations between radiologist-defined semantic and automatically computed radiomic features in non-small cell lung cancer. Sci Rep 7(1):1–11. https://doi.org/10.1038/s41598-017-02425-5
Kazerooni EA, Gross BH (2004) Cardiopulmonary imaging. Lippincott Williams & Wilkins, Philadelphia, p 379
Wong HYF, Lam HYS, Fong AHT, Leung ST, Chin TWY, Lo CSY, Lui MMS, Lee JCY, Chiu KWH, Chung T, Lee EYP, Wan EYF, Huang FNI, Lam TPW, Ng KMY (2020) Frequency and distribution of chest radiographic findings in COVID-19 positive patients. Radiology 2020:201160. https://doi.org/10.1148/radiol.2020201160
Gonzalez RC, Woods RE, Eddins SL (2009) Digital image processing using Matlab, 2nd edn. Gatesmark Publishing, Knoxville, pp 644–656
Haralick RM, Shanmugam K, Dinstein IH (1973) Textural features for image classification. IEEE T Syst Man CY B 6:610–621. https://doi.org/10.1109/TSMC.1973.4309314
Haralick RM (1979) Statistical and structural approaches to texture. Proc IEEE 67(5):786–804. https://doi.org/10.1109/PROC.1979.11328
Tang X (1998) Texture information in run-length matrices. IEEE Trans Image Process 7(11):1602–1609. https://doi.org/10.1109/83.725367
Mallat SG (1989) A theory for multiresolution signal decomposition: the wavelet representation. IEEE Trans Pattern Anal Mach Intell 11(7):674–693. https://doi.org/10.1109/34.192463
Do MN, Vetterli M (2005) The contourlet transform: an efficient directional multiresolution image representation. IEEE Trans Image Process 14(12):2091–2106. https://doi.org/10.1109/TIP.2005.859376
Qiu JJ, Wu Y, Hui B, Huang ZX, Chen J (2018) Texture analysis of computed tomography images in the classification of pancreatic cancer and normal pancreas: a feasibility study. J Med Imag Health Inform 8(8):1539–1545. https://doi.org/10.1166/jmihi.2018.2507
Qiu JJ, Wu Y, Hui B, Chen J, Ji L, Wang M (2018) A novel texture analysis method based on reverse biorthogonal wavelet and co-occurrence matrix applied for classification of hepatocellular carcinoma and hepatic hemangioma. J Med Imag Health Inform 8(9):1835–1843. https://doi.org/10.1166/jmihi.2018.2516
Qiu JJ, Wu Y, Chen J, Hui B, Huang ZX, Ji L (2018) A texture analysis method based on statistical contourlet coefficient applied to the classification of pancreatic cancer and normal pancreas. Int Symp Big Dat Artif Intell. https://doi.org/10.1145/3305275.3305278
Chakraborty J, Rangayyan RM, Banik S, Mukhopadhyay S, Desautels JEL (2012) Statistical measures of orientation of texture for the detection of architectural distortion in prior mammograms of interval-cancer. J Electron Imaging 21(3):033010. https://doi.org/10.1117/1.JEI.21.3.033010
Szczypinski P, Strzelecki M, Materka A, Klepaczko A (2009) MaZda-A software package for image texture analysis. Comput Methods Programs Biomed 94(1):66–76. https://doi.org/10.1016/j.cmpb.2008.08.005
Breiman L (1995) Better subset regression using the nonnegative garrote. Technometrics 37(4):373–384. https://doi.org/10.1080/00401706.1995.10484371
Tibshirani R (1996) Regression shrinkage and selection via the lasso. J R Stat Soc Ser B (Methodological) 58(1):267–288. https://doi.org/10.1111/j.2517-6161.1996.tb02080.x
Sauerbrei W, Royston P, Binder H (2007) Selection of important variables and determination of functional form for continuous predictors in multivariable model building. Stat Med 26(30):5512–5528. https://doi.org/10.1002/sim.3148
Redmon J, Farhadi A (2018) Yolov3: an incremental improvement. arXiv preprint arXiv:1804.02767v
Wu Q, Wang S, Li L, Wu Q, Qian W, Hu Y, Li L, Zhou X, Ma H, Li H, Wang M, Qiu X, Zha Y, Tian J (2020) Radiomics Analysis of Computed Tomography helps predict poor prognostic outcome in COVID-19. Theranostics 10(16):7231. https://doi.org/10.7150/thno.46428
Fang X, Li X, Bian Y, Ji X, Lu J (2020) Radiomics nomogram for the prediction of 2019 novel coronavirus pneumonia caused by SARS-CoV-2. Eur Radiol. https://doi.org/10.1007/s00330-020-07032-z
Fu L, Li Y, Cheng A, Pang P, Shu Z (2020) A novel machine learning-derived radiomic signature of the whole lung differentiates stable from progressive COVID-19 infection: a retrospective cohort study. J Thorac Imaging. https://doi.org/10.1097/RTI.0000000000000544
Ozturk T, Talo M, Yildirim EA, Baloglu UB, Yildirim O, Acharya UR (2020) Automated detection of COVID-19 cases using deep neural networks with X-ray images. Comput Biol Med 121:103792. https://doi.org/10.1016/j.compbiomed.2020.103792
Litjens G, Kooi T, Bejnordi BE, Setio AAA, Ciompi F, Ghafoorian M, Laak JVD, Ginneken BV, Sanchez CI (2017) A survey on deep learning in medical image analysis. Med Image Anal 42:60–88. https://doi.org/10.1016/j.media.2017.07.005
Zhao ZQ, Zheng P, Xu S, Wu X (2019) Object detection with deep learning: a review. IEEE Trans Neural Netw Learn Syst 30(11):3212–3232. https://doi.org/10.1109/TNNLS.2018.2876865
Hesamian MH, Jia W, He X, Kennedy P (2019) Deep learning techniques for medical image segmentation: achievements and challenges. J Dig Imaging 32(4):582–596. https://doi.org/10.1007/s10278-019-00227-x
Yang J, Dong X, Hu Y, Peng Q, Tao G, Ou Y, Cai H, Yang X (2020) Fully automatic arteriovenous segmentation in retinal images via topology-aware generative adversarial networks. Interdiscip Sci Comput Life Sci 12(3):323–334. https://doi.org/10.1007/s12539-020-00385-5
He K, Zhang X, Ren S, Sum J (2016) Deep residual learning for image recognition. Proc IEEE Conf Comput Vis Pattern Recogn. https://doi.org/10.1109/CVPR.2016.90
Zhang Z, Xie Y, Xing F, McGough M, Yang L (2017) Mdnet: a semantically and visually interpretable medical image diagnosis network. IEEE Conf Comput Vis Pattern Recogn. https://doi.org/10.1109/CVPR.2017.378
Lambin P, Leijenaar RTH, Deist TM, Peerlings J, de Jong EEC, Timmeren JV, Sanduleanu S, Larue RTHM, Even AJG, Jochems A, Wijk YV, Woodruff H, Soest JV, Lustberg T, Roelofs E, Elmpt WV, Dekker A, Mottaghy FM, Wildberger JE, Walsh S (2017) Radiomics: the bridge between medical imaging and personalized medicine. Nat Rev Clin Oncol 14(12):749–762. https://doi.org/10.1038/nrclinonc.2017.141
Acknowledgements
This work was supported by the Department of Science and Technology of Sichuan Province (no. 2020YFS0556, no. 2020YFS0007). The data collection for this work was supported by Chengdu Public Health Clinical Medical Center, Luzhou People’s Hospital, Ya’an People’s Hospital, Wusheng County People’s Hospital, Dazhou People’s Hospital, Yuechi County People’s Hospital, Zigong First People’s Hospital, Sichuan Mianyang 404 Hospital, Qionglai Medical Center Hospital Chengdu First People’s Hospital, Guangyuan Third People Hospital, Guang’an People’s Hospital, Neijiang Second People’s Hospital, Panzhihua Fourth People’s Hospital, and Chengdu Pidu District People’s Hospital.
Funding
This work was supported by the Department of Science and Technology of Sichuan Province (No. 2020YFS0556, No. 2020YFS0007).
Author information
Authors and Affiliations
Contributions
(1) Substantial contributions to conception and design, acquisition of data, or analysis and interpretation of data: (a) substantial contributions to conception and design: JQ, SP, HS, and WZ; (b) acquisition of data: ZL, JY, JW, and JJ; (c) analysis and interpretation of data: JQ. (2) Drafting the article of revising it critically for important intellectual content: JQ and HS. (3) Final approval of the version to be published: all. (4) Agreement to be accountable for all aspects of the work: JQ, HS.
Corresponding author
Ethics declarations
Conflict of interest
We declare that we have no financial and personal relationships with other people or organizations that can inappropriately influence our work, and there is no professional or other personal interest of any nature or kind in any product, service and/or company that could be construed as influencing the position presented in or the review of the manuscript entitled “A radiomics signature to quantitatively analyze COVID-19-infected pulmonary lesions”.
Ethics approval
This work was approved by the Ethics Committee of West China Hospital of Sichuan University (number 2020190).
Consent to participate
This work is a retrospective study approved by the Ethics Committee of West China Hospital of Sichuan University (number 2020190), and the need to obtain informed consent was waived.
Consent for publication
This work is a retrospective study approved by the Ethics Committee of West China Hospital of Sichuan University (number 2020190), and the need to obtain informed consent was waived.
Code availability
The codes during and/or analyzed during the current study are available from the corresponding author on reasonable request.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Qiu, J., Peng, S., Yin, J. et al. A Radiomics Signature to Quantitatively Analyze COVID-19-Infected Pulmonary Lesions. Interdiscip Sci Comput Life Sci 13, 61–72 (2021). https://doi.org/10.1007/s12539-020-00410-7
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12539-020-00410-7