Abstract
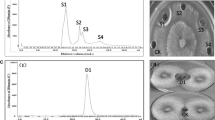
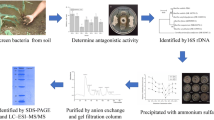
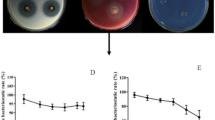
A newly discovered alkaline antifungal protease named P6 from Bacillus subtilis N7 was purified and partially characterized. B. subtilis N7 culture filtrates were purified by 30–60% (NH4)2SO4 precipitation, anion-exchange chromatography and gel filtration chromatography. Sodium dodecyl sulfate-polyacrylamide gel electrophoresis (SDS-PAGE) revealed a single band of 41.38 kDa. Peptide sequence of protease P6 was determined using a 4800 Plus MALDI TOF/TOF™ Analyzer System. Self-Formed Adaptor PCR (SEFA-PCR) was used to amplify the 1,149 bp open read frame of P6. Dimensional structure prediction using Automatic Modeling Mode software showed that the protease P6 consisted of two β-barrel domains. Purified P6 strongly inhibited spore and mycelium growth of Fusarium oxysporum f. sp. cucumerium (FOC) by causing hypha lysis when the concentration was 25 μg/ml. Characterization of the purified protease indicated that it had substrate specificity for gelatin and was highly active at pH 8.0–10.6 and 70°C. The P6 protease was inhibited by EDTA (2 mmol/L), phenyl methyl sulfonyl fluoride (PMSF, 1 mmol/L), Na+, Fe3+, Cu2+, Mg2+ (5 mmol/L each) and H2O2 (2%, v/v). However, protease activity was activated by Ca2+, K+, Mn2+ (5 mmol/L each), mercaptoethanol (2%, v/v) and Tween 80 (1%, v/v). In additon, activity was also affected by organic solvents such as acetone, normal butanol and ethanol, but not hexane (25%, v/v each).
Similar content being viewed by others
References
Adinarayana, K., Ellaiah, P., and Prasad, D.S. 2003. Purification and partial characterization of thermostable serine alkaline protease from a newly isolated Bacillus subtilis pe-11. AAPS PharmSciTech. 4, E56.
Becher, D., Buttner, K., Moche, M., Hessling, B., and Hecker, M. 2011. From the genome sequence to the protein inventory of Bacillus subtilis. Proteomics 11, 2971–2980.
Bhaskar, N., Sudeepa, E., Rashmi, H., and Tamil Selvi, A. 2007. Partial purification and characterization of protease of Bacillus proteolyticus cfr3001 isolated from fish processing waste and its antibacterial activities. Bioresour. Technol. 98, 2758–2764.
Borisova, S.A., Circello, B.T., Zhang, J.K., van der Donk, W.A., and Metcalf, W.W. 2010. Biosynthesis of rhizocticins, antifungal phosphonate oligopeptides produced by Bacillus subtilis ATCC 6633. Chem. Biol. 17, 28–37.
Bradford, M.M. 1976. A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal. Biochem. 72, 248–254.
Broekaert, W.F., Terras, F.R.G., Cammue, B., and Vanderleyden, J. 1990. An automated quantitative assay for fungal growth inhibition. FEMS Microbiol. Lett. 69, 55–59.
Chen, X., Li, J., Sun, Q., Tong, Y., and Xu, J. 2010. Isolation, purification and characterization of antifungal protein from rice endophytic bacterim Bacillus subtilis G87. Acta Microbiol. Sinica 50, 1353.
Cho, K.M., Math, R.K., Hong, S.Y., Asraful Islam, S.M., Mandanna, D.K., Cho, J.J., Yun, M.G., Kim, J.M., and Yun, H.D. 2009. Iturin produced by Bacillus pumilus Hy1 from Korean soybean sauce (kanjang) inhibits growth of aflatoxin producing fungi. Food Control. 20, 402–406.
Chung, S., Kong, H., Buyer, J.S., Lakshman, D.K., Lydon, J., Kim, S.D., and Roberts, D.P. 2008. Isolation and partial characterization of Bacillus subtilis ME488 for suppression of soilborne pathogens of cucumber and pepper. Appl. Microbiol. Biotechnol. 80, 115–123.
Das, S., Mishra, B., Gill, K., Ashraf, M.S., Singh, A.K., Sinha, M., Sharma, S., Xess, I., Dalal, K., Singh, T.P., and et al. 2011. Isolation and characterization of novel protein with anti-fungal and anti-inflammatory properties from aloe vera leaf gel. Int. J. Biol. Macromol. 48, 38–43.
Earl, A.M., Losick, R., and Kolter, R. 2008. Ecology and genomics of Bacillus subtilis. Trends Microbiol. 16, 269–275.
Guangrong, H., Tiejing, Y., Po, H., and Jiaxing, J. 2010. Purification and characterization of a protease from thermophilic Bacillus strain HS08. Afr. J. Biotechnol. 5, 2433–2438.
Gupta, R., Beg, Q., and Lorenz, P. 2002. Bacterial alkaline proteases: Molecular approaches and industrial applications. Appl. Microbiol. Biotechnol. 59, 15–32.
Harwood, C.R. 1992. Bacillus subtilis and its relatives: Molecular biological and industrial workhorses. Trends Biotechnol. 10, 247–256.
Hedstrom, L. 2002. Serine protease mechanism and specificity. Chem. Rev. 102, 4501–4524.
Kim, P. and Chung, K.C. 2004. Production of an antifungal protein for control of colletotrichum lagenarium by Bacillus amyloliquefaciens MET0908. FEMS Microbiol. Lett. 234, 177–183.
Kugler, M., Loeffler, W., Rapp, C., Kern, A., and Jung, G. 1990. Rhizocticin a, an antifungal phosphono-oligopeptide of Bacillus subtilis ATCC 6633: Biological properties. Arch. Microbiol. 153, 276–281.
Laemmli, U.K. 1970. Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature 227, 680–685.
Li, J., Yang, Q., Zhao, L.H., Zhang, S.M., Wang, Y.X., and Zhao, X.Y. 2009. Purification and characterization of a novel antifungal protein from Bacillus subtilis strain B29. J. Zhejiang Univ. Sci. B. 10, 264–272.
Liu, Y., Chen, Z., Ng, T.B., Zhang, J., Zhou, M., Song, F., and Lu, F. 2007. Bacisubin, an antifungal protein with ribonuclease and hemagglutinating activities from Bacillus subtilis strain B-916. Peptides 28, 553–559.
Liu, B., Huang, L., Buchenauer, H., and Kang, Z. 2010. Isolation and partial characterization of an antifungal protein from the endophytic Bacillus subtilis strain EDR4. Pestic. Biochem. Physiol. 98, 305–311.
Manjula, K., Kishore, G.K., and Podile, A. 2004. Whole cells of Bacillus subtilis AF 1 proved more effective than cell-free and chitinase-based formulations in biological control of citrus fruit rot and groundnut rust. Can. J. Microbiol. 50, 737–744.
Mizumoto, S. and Shoda, M. 2007. Medium optimization of antifungal lipopeptide, iturin a, production by Bacillus subtilis in solid-state fermentation by response surface methodology. Appl. Microbiol. Biotechnol. 76, 101–108.
Nilegaonkar, S., Zambare, V., Kanekar, P., Dhakephalkar, P., and Sarnaik, S. 2007. Production and partial characterization of dehairing protease from Bacillus cereus MCM B-326. Bioresour. Technol. 98, 1238–1245.
Oberoi, R., Beg, Q.K., Puri, S., Saxena, R., and Gupta, R. 2001. Characterization and wash performance analysis of an SDS-stable alkaline protease from a Bacillus sp. World J. Microbiol. Biotechnol. 17, 493–497.
Oman, T.J., Boettcher, J.M., Wang, H., Okalibe, X.N., and van der Donk, W.A. 2011. Sublancin is not a lantibiotic but an S-linked glycopeptide. Nature Chem. Biol. 7, 78–80.
Özcengiz, G. and Alaeddinoglu, N. 1991. Bacilysin production by Bacillus subtilis: effects of bacilysin, pH and temperature. Folia Microbiol. (Praha) 36, 522–526.
Perona, J.J. and Craik, C.S. 2008. Structural basis of substrate specificity in the serine proteases. Protein Sci. 4, 337–360.
Pillay, V., Polya, G.M., and Spangenberg, G.C. 2011. Optimisation of an in vitro antifungal protein assay for the screening of potential antifungal proteins against Leptosphaeria maculans. J. Microbiol. Methods 84, 121–127.
Rapp, C., Jung, G., Kugler, M., and Loeffler, W. 1988. Rhizocticins-new phosphono-oligopeptides with antifungal activity. Liebigs Annalen der Chemie 1988, 655–661.
Reddy, L.V.A., Wee, Y.-J., and Ryu, H.-W. 2008. Purification and characterization of an organic solvent and detergent-tolerant novel protease produced by Bacillus sp. RKY3. J. Chem. Technol. Biotechnol. 83, 1526–1533.
Riffel, A., Ortolan, S., and Brandelli, A. 2003. De-hairing activity of extracellular proteases produced by keratinolytic bacteria. J. Chem. Technol. Biotechnol. 855-859.
Stein, T. 2005. Bacillus subtilis antibiotics: Structures, syntheses and specific functions. Mol. Microbiol. 56, 845–857.
Toure, Y., Ongena, M., Jacques, P., Guiro, A., and Thonart, P. 2004. Role of lipopeptides produced by Bacillus subtilis GA1 in the reduction of grey mould disease caused by Botrytis cinerea on apple. J. Appl. Microbiol. 96, 1151–1160.
Wang, S., He, J., Cui, Z., and Li, S. 2007. Self-formed adaptor PCR: a simple and efficient method for chromosome walking. Appl. Environ. Microbiol. 73, 5048–5051.
Wang, H. and Ng, T.B. 2005. An antifungal protein from ginger rhizomes. Biochem. Biophys. Res. Commun. 336, 100–104.
Xie, D., Peng, J., Wang, J., Hu, J., and Wang, Y. 1998. Purification and properties of antifungal protein x98iii from Bacillus subtilis. Acta Microbiol. Sinica 38, 13.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Luo, Y., Sun, L., Zhu, Z. et al. Identification and characterization of an anti-fungi Fusarium oxysporum f. sp. cucumerium protease from the Bacillus subtilis strain N7. J Microbiol. 51, 359–366 (2013). https://doi.org/10.1007/s12275-013-2627-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12275-013-2627-6