Abstract
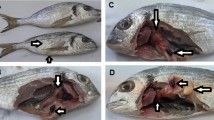
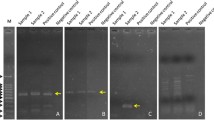
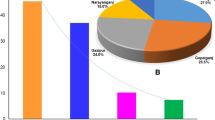
Intensive fish farming systems have led to increase in disease incidence, due to higher stocking density, high organic matter levels, and poor quality of the aquatic environment. Diseased fish samples showing hemorrhages and reddish lesions were collected from different freshwater fish farms located at three different districts of West Bengal, India (Burdwan, North 24 Parganas, and Nadia). The present study was conducted to evaluate the genetic diversity of ten different Klebsiella pneumoniae strains isolated from different infected freshwater fish samples based on 16S rRNA gene sequence analysis. Primarily, Klebsiella-specific media was used for the isolation and characterization of Klebsiella pneumoniae. Further, through a biochemical test, all the strains were confirmed as K. pneumoniae. PCR analysis of 16S–23S internal transcribed spacer (PCR ribotyping) was carried out to study the species variation within different Klebsiella pneumoniae isolates. For all the isolates, a conserved PCR ribotype pattern was observed while differing from other bacterial species. Phylogenetic study showed the high degree of homology with diverse source of other strains. The multiple antibiotic resistance (MAR) values of the present study for the isolates were found to be 0.468. MAR value above 0.2 indicates that the source of isolation was highly contaminated with antibiotics. Based on the 16S rRNA gene sequence analysis, the present study revealed the genetic diversity of Klebsiella pneumoniae isolated from the different diseased fish farms of West Bengal. All the strains were found to be hypermucoviscous and multidrug-resistant, thus making it pathogenic towards the host organisms. Further, the study revealed a high prevalence of K. pneumoniae in aquaculture farms, representing a risk towards successful aquaculture.






Similar content being viewed by others
References
Abbott JG, Campbell LM, Hay CJ, Næsje TF, Purvis J (2007) Market-resource links and fish vendor livelihoods in the upper Zambezi river floodplains. Hum Ecol 35:559–574
Abdul-Razzaq MS, Al-Khafaji JK, Al-Maamory EH (2014) Molecular characterization of capsular polysaccharide genes of Klebsiella pneumoniae in Iraq. Int J Curr Microbiol App Sci 3:224–234
Aher T, Roy A, Kumar P (2012) Molecular detection of virulence genes associated with pathogenicity of Klebsiella spp. isolated from the respiratory tract of apparently healthy as well as sick goats. Isr J Vet Med 67:249–252
Akond MA, Alam S, Hassan SM, Shirin M (2009) Antibiotic resistance of Escherichia coli isolated from poultry and poultry environment of Bangladesh. Int J food safety 11:19–23
Al-Imarah EA (2008) Distribution of some aerobic bacteria in an infected Cyprinus carpio L. fish farm in Basrah and its resistance to antibiotics. JKU 6:209–215
Apun K, Yusof AM, Jugang K (1999) Distribution of bacteria in tropical freshwater fish and ponds. Int J Environ Health Res 9:285–292
Bagley ST, Seidler RJ (1978) Primary Klebsiella identification with MacConkey-inositol-carbenicillin agar. Appl Environ Microbiol 36:536–538
Bauer AW, Kirby WM, Sherris JC, Turck M (1966) Antibiotic susceptibility testing by a standardized single disk method. Am J Clin Pathol 45:493–496
Behera BK, Paria P, Das A, Bhowmick S, Sahoo AK, Das BK (2017) Molecular characterization and pathogenicity of a virulent Acinetobacter baumannii associated with mortality of farmed Indian major carp Labeo rohita (Hamilton 1822). Aquaculture 471:157–162
Brisse S, Verhoef J (2001) Phylogenetic diversity of Klebsiella pneumoniae and Klebsiella oxytoca clinical isolates revealed by randomly amplified polymorphic DNA, gyrA and parC genes sequencing and automated ribotyping. Int J Syst Evol Microbiol 51:915–924
Das A, Acharya S, Behera BK, Paria P, Bhowmick S, Parida PK, Das BK (2018) Isolation, identification and characterization of Klebsiella pneumoniae from infected farmed Indian major carp Labeo rohita (Hamilton 1822) in West Bengal, India. Aquaculture 482:111–116
de Pádua SB, Marques DP, Sebastião FA, Pilarski F, Martins ML, Ishikawa MM (2014) Isolation, characterization and pathology of Citrobacter freundii infection in native Brazilian catfish Pseudo platystoma. BJVP 7:151–157
Deshpande RG, Khan MB (1999) Purification and characterization of hemolysin from Porphyromonasgingivalis A7436. FEMS Microbiol Lett 176:387–394
Diana T, Manjulatha C (2012) Incidence and identification of Klebsiella pneumoniae in mucosal buccal polyp of Nemipterus japonicus of Visakhapatnam Coast. India J Fish Aquat Sci 7:454–460
Dorsch MR (2007) Rapid detection of bacterial antibiotic resistance: preliminary evaluation of PCR assays targeting tetracycline resistance genes (no. DSTO-TR-2059)
Fang CT, Chuang YP, Shun CT, Chang SC, Wang JT (2004) A novel virulence gene in Klebsiella pneumoniae strains causing primary liver abscess and septic metastatic complications. J Exp Med 199:697–705
Garcı́a-Martı́nez J, Acinas SG, Anton AI, Rodrı́guez-Valera F (1999) Use of the 16S–23S ribosomal genes spacer region in studies of prokaryotic diversity. J Microbiol Methods 36:55–64
Gee JE, Sacchi CT, Glass MB, De BK, Weyant RS, Levett PN, Whitney AM, Hoffmaster AR, Popovic T (2003) Use of 16S rRNA gene sequencing for rapid identification and differentiation of Burkholderia pseudomallei and B. mallei. J Clin Microbio l4:4647–4654
Gharrah MM, Mostafa El-Mahdy A, Barwa RF (2017) Association between virulence factors and extended spectrum beta-lactamase producing Klebsiella pneumoniae compared to nonproducing isolates. Interdiscip Perspect Infect Dis 2017:1–14
Gopi M, Kumar TT, Prakash S (2016) Opportunistic pathogen Klebsiella pneumoniae isolated from Maldive’s clown fish Amphiprion nigripes with hemorrhages at Agatti Island, Lakshadweep archipelago. Int J Fisheries Aquatic Studies 4:464–467
Gupta A, Ampofo K, Rubenstein D, Saiman L (2003) Extended spectrum β lactamase-producing Klebsiella pneumoniae infections: a review of the literature. J Perinatol 23:439–443
Gürtler V, Stanisich VA (1996) New approaches to typing and identification of bacteria using the 16S-23S rDNA spacer region. Microbiol 142:3–16
Hansen DS, Aucken HM, Abiola T, Podschun R (2004) Recommended test panel for differentiation of Klebsiella species on the basis of a trilateral interlaboratory evaluation of 18 biochemical tests. J Clin Microbiol 42:3665–3669
Izquierdo L, Merino S, Regué M, Rodriguez F, Tomás JM (2003) Synthesis of a Klebsiella pneumoniae O-antigen heteropolysaccharide (O12) requires an ABC 2 transporter. J Bacteriol 185:1634–1641
Jang S, Wheeler L, Carey RB, Jensen B, Crandall CM, Schrader KN, Jessup D, Colegrove K, Gulland FM (2010) Pleuritis and suppurative pneumonia associated with a hypermucoviscosity phenotype of Klebsiella pneumoniae in California sea lions (Zalophus californianus). Vet Microbiol 141:174–177
Kang CI, Kim SH, Bang JW, Kim HB, Kim NJ, Kim EC, Oh MD, Choe KW (2006) Community-acquired versus nosocomial Klebsiella pneumoniae bacteremia: clinical features, treatment outcomes, and clinical implication of antimicrobial resistance. J Korean Med Sci 21:816–822
Karunasagar I, Karunasagar I, Pai R (1992) Systemic Citrobacter freundii infection in common carp, Cyprinus carpio L., fingerlings. J Fish Dis 15:95–98
Kathleen MM, Samuel L, Felecia C, Reagan EL, Kasing A, Lesley M, Toh SC (2016) Antibiotic resistance of diverse bacteria from aquaculture in Borneo. Int J Microbiol 2016:1–9
Kawai T (2006) Hypermucoviscosity: an extremely sticky phenotype of Klebsiella pneumoniae associated with emerging destructive tissue abscess syndrome. Clin Infect Dis 42:1359–1361
Kelly RF, Severn WB, Richards JC, Perry MB, MacLean LL, Tomás JM, Merino S, Whitfield C (1993) Structural variation in the O-specific polysaccharides of Klebsiella pneumoniae serotype O1 and O8 lipopolysaccharide: evidence for clonal diversity in rfb genes. Mol Microbiol 10:615–625
Klappenbach JA, Saxman PR, Cole JR, Schmidt TM (2001) rrndb: the ribosomal RNA operon copy number database. Nucleic Acids Res 29:181–184
Knothe H, Shah P, Krcmery V, Antal M, Mitsuhashi S (1983) Transferable resistance to cefotaxime, cefoxitin, cefamandole and cefuroxime in clinical isolates of Klebsiella pneumoniae and Serratia marcescens. Infection 11:315–317
Kostman JR, Edlind TD, LiPuma JJ, Stull TL (1992) Molecular epidemiology of Pseudomonas cepacia determined by polymerase chain reaction ribotyping. JCM 30:2084–2087
Laith AR, Najiah M (2013) Aeromonas hydrophila: antimicrobial susceptibility and histopathology of isolates from diseased catfish, Clarias gariepinus (Burchell). J Aquac Res Development 5:215. https://doi.org/10.4172/2155-9546.1000215
Lee IR, Molton JS, Wyres KL, Gorrie C, Wong J, Hoh CH, Teo J, Kalimuddin S, Lye DC, Archuleta S, Holt KE (2016) Differential host susceptibility and bacterial virulence factors driving Klebsiella liver abscess in an ethnically diverse population. Sci Rep 13:29316
Lopes AC, Rodrigues JF, Clementino M, Miranda CA, Nascimento AP, MoraisJúnior MA (2007) Application of PCR ribotyping and tDNA-PCR for Klebsiella pneumoniae identification. Mem Inst Oswaldo Cruz 102:827–832
Maroncle N, Rich C, Forestier C (2006) The role of Klebsiella pneumoniae urease in intestinal colonization and resistance to gastrointestinal stress. Res Microbiol 157:184–193
Mobley HL, Warren JW (1996) Urinary tract infections: molecular pathogenesis and clinical management. ASM press, Washington
Nadasy KA, Domiati-Saad R, Tribble MA (2007) Invasive Klebsiella pneumoniae syndrome in North America. Clin Infect Dis 45:25–28
Oliveira RV, Peixoto PG, Ribeiro DD, Araujo MC, do Santos CT, Hayashi C, Pedreira MM, Pelli A (2014) Klebsiella pneumoniae as a main cause of infection in Nishikigoi Cyprinus carpio (carp) by inadequate handling. Braz J Vet Pathol 7:86–88
Ørskov I, Ørskov F (1984) Serotyping of Klebsiella. In Methods Microbiol 14:143–164
Osundiya OO, Oladele RO, Oduyebo OO (2013) Multiple antibiotic resistance (MAR) indices of Pseudomonas and Klebsiella species isolates in Lagos University Teaching Hospital. AJCEM 14:164–168
Park SB, Aoki T, Jung TS (2012) Pathogenesis of and strategies for preventing Edwardsiella tarda infection in fish. Vet Res 43:67–78
Patel SS, Chauhan HC, Patel AC, Shrimali MD, Patel KB, Prajapati BI (2017) Isolation and identification of Klebsiella pneumoniae from Sheep-Case Report. Int J Curr Microbiol App Sci 6:331–334
Podschun R, Ullmann U (1998) Klebsiella spp. as nosocomial pathogens: epidemiology, taxonomy, typing methods, and pathogenicity factors. Clin Microbiol 11:589–603
Pulkkinen K, Suomalainen LR, Read AF, Ebert D, Rintamäki P, Valtonen ET (2010) Intensive fish farming and the evolution of pathogen virulence: the case of columnaris disease in Finland. Proc R SocLond B BiolSci 277:593–600
Rhoads DD, Cox SB, Rees EJ, Sun Y, Wolcott RD (2012) Clinical identification of bacteria in human chronic wound infections: culturing vs. 16S ribosomal DNA sequencing. BMC Infect Dis 12:321–329
Rompre A, Servais P, Baudart J, De-Roubin MR, Laurent P (2002) Detection and enumeration of coliforms in drinking water: current methods and emerging approaches. J Microbiol Methods 49:31–54
Sambrook J, Fritsch EF, Maniatis T (1989) Molecular cloning: a laboratory manual, vol 62. Cold spring harbor laboratory press, New York
Sandhu R, Dahiya S, Sayal P (2016) Evaluation of multiple antibiotic resistance (MAR) index and doxycycline susceptibility of Acinetobacter species among inpatients. Indian J Microbiol Res 3:299–304
Sarter S, Nguyen HN, Hung LT, Lazard J, Montet D (2007) Antibiotic resistance in Gram-negative bacteria isolated from farmed catfish. Food Control 1811:1391–1396
Sharma SK, Mudgal NK, Sharma P, Shrngi BN (2015) Comparison of phenotypic characteristics and virulence traits of Klebsiella pneumoniae obtained from pneumonic and healthy camels (Camelus dromedarius). Adv Anim Vet Sci 3:116–122
Shon AS, Bajwa RP, Russo TA (2013) Hypervirulent (hypermucoviscous) Klebsiella pneumoniae: a new and dangerous breed. Virulence 4:107–118
Singh V, Chaudhary DK, Mani I (2012) Molecular characterization and modeling of secondary structure of 16S rRNA from Aeromonas veronii. Int J Appl Biol Pharm Technol 3:253–260
Siu LK, Fung CP, Chang FY, Lee N, Yeh KM, Koh TH, Ip M (2011) Molecular typing and virulence analysis among serotype K1 Klebsiella pneumoniae isolated from liver abscess patients and stool carriage from non-infectious subjects in Hong Kong, Singapore and Taiwan. JCM 49:3761–3765
Song J, Lee SC, Kang JW, Baek HJ, Suh JW (2004) Phylogenetic analysis of Streptomyces spp. isolated from potato scab lesions in Korea on the basis of 16S rRNA gene and 16S–23S rDNA internally transcribed spacer sequences. Int J Syst Evol Microbiol 54:203–209
Srinivasan R, Karaoz U, Volegova M, MacKichan J, Kato-Maeda M, Miller S, Nadarajan R, Brodie EL, Lynch SV (2015) Use of 16S rRNA gene for identification of a broad range of clinically relevant bacterial pathogens. PLoS One 10:e0117617
Starliper CE (2011) Bacterial coldwater disease of fishes caused by Flavobacterium psychrophilum. J Adv Res 2:97–108
Sugita H, Nakamura T, Tanaka K, Deguchi Y (1994) Identification of Aeromonas species isolated from freshwater fish with the microplate hybridization method. Appl Environ Microbiol 60:3036–3038
Takyi R, Nunoo FK, Ziddah P, Oddoye J (2012) Occurrence of bacterial infection in two commonly cultured fish species on two fish farms in southern Ghana. World J Biol Res 5:81–92
Tamura K, Stecher G, Peterson D, Filipski A, Kumar S (2013) MEGA6: molecular evolutionary genetics analysis version 6.0. Mol Biol Evol 30:2725–2729
Turton JF, Englender H, Gabriel SN, Turton SE, Kaufmann ME, Pitt TL (2007) Genetically similar isolates of Klebsiella pneumoniae serotype K1 causing liver abscesses in three continents. J Med Microbiol 56:593–597
Woo PC, Teng JL, Wu JK, Leung FP, Tse H, Fung AM, Lau SK, Yuen KY (2009) Guidelines for interpretation of 16S rRNA gene sequence-based results for identification of medically important aerobic Gram-positive bacteria. J Med Microbiol 58:1030–1036
World Bank (2014) Reducing disease risk in aquaculture. World Bank Report number 88257-GLB, Agriculture and Environmental Services Discussion Paper 09. http://documents.worldbank.org/curated/en/110681468054563438/pdf/882570REPLACEM00NAME0Reantaso0Melba.pdf
Wyatt LE, Nickelson R, Vanderzant CA (1979) Edwardsiella tarda in freshwater catfish and their environment. Appl Environ Microbiol 38:710–714
Yeh KM, Lin JC, Yin FY, Fung CP, Hung HC, Siu LK, Chang FY (2010) Revisiting the importance of virulence determinant mag A and its surrounding genes in Klebsiella pneumoniae causing pyogenic liver abscesses: exact role in serotype K1 capsule formation. J Infect Dis 201:1259–1267
Younis G, Awad A, El-Gamal A, Hosni R (2016) Virulence properties and antimicrobial susceptibility profiles of Klebsiella species recovered from clinically diseased broiler chicken. AdvAnim Vet Sci 4:536–542
Yu WL, Ko WC, Cheng KC, Lee HC, Ke DS, Lee CC, Fung CP, Chuang YC (2006) Association between rmpA and magA genes and clinical syndromes caused by Klebsiella pneumoniae in Taiwan. Clin Infect Dis 42:1351–1358
Zamani A, Mashouf RY, Namvar AM, Alikhani MY (2013) Detection of magA Gene in Klebsiella spp. isolated from clinical samples detection of magA. Iran J Basic Med Sci 16:173–176
Acknowledgments
The authors are thankful to Dr. Joykrushna Jena, Deputy Director General (Fishery Science), Indian Council of Agricultural Research, New Delhi, for his support and guidance. The authors are also thankful to Mr. Asim Kumar Jana for laboratory assistance.
Funding
This work was funded by the National Fisheries Development Board (NFDB) (G/Nat. Surveillance/2013 dated 16.08.2013), Hyderabad, under the “National Surveillance Program for Aquatic Animal Diseases (NSPAAD)” Project.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Ethical approval
The fish samples for this study were collected and handled in accordance with the ICAR-Central Inland Fisheries Research Institute (CIFRI), Kolkata, India ethical guidelines, and regulations. Further, the experimental protocols for this study were approved by the Institute Research Committee of ICAR-CIFRI (Project Code: OXXO3049).
Conflict of interest
The authors have declared no conflict of interest.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
ESM 1
(DOCX 184 kb)
Rights and permissions
About this article
Cite this article
Das, A., Behera, B.K., Acharya, S. et al. Genetic diversity and multiple antibiotic resistance index study of bacterial pathogen, Klebsiella pneumoniae strains isolated from diseased Indian major carps. Folia Microbiol 64, 875–887 (2019). https://doi.org/10.1007/s12223-019-00701-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12223-019-00701-7