Abstract
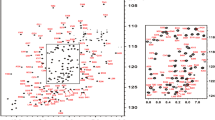
Hepatocyte nuclear factor 1β (HNF1β) is a transcription factor that plays a key role in the development and function of the liver, pancreas, and kidney. HNF1β plays a key role in early vertebrate development and the morphogenesis of these organs. In humans, heterozygous mutations in the HNF1B gene can result in organ dysplasia, making it the most common cause of developmental renal diseases, including renal cysts, renal malformations, and familial hypoplastic glomerular cystic kidney disease. Pathogenic variants in the HNF1B gene are known to cause various diseases, including maturity-onset diabetes of the young and developmental renal diseases. This study presents the backbone resonance assignments of HNF1β POUS and POUHD domains, which are highly conserved domains required for the recognition of double-stranded DNA. Our data will be useful for NMR studies to verify the altered structures and functions of mutant HNF1B proteins that can induce developmental renal diseases, including renal cysts, renal malformations, and familial hypoplastic glomerular cystic kidney disease. This study will provide the structural basis for future studies to elucidate the molecular mechanisms underlying how mutations in HNF1β cause diseases.



Similar content being viewed by others
Data availability
Assignments have been deposited at the Biological Magnetic Resonance Bank (BMRB) under ID 26339 and 26,340.
References
Bingham C, Bulman MP, Ellard S, Allen LI, Lipkin GW, Hoff WG, Woolf AS, Rizzoni G, Novelli G, Nicholls AJ, Hattersley AT (2001) Mutations in the hepatocyte nuclear factor-1beta gene are associated with familial hypoplastic glomerulocystic kidney disease. Am J Hum Genet 68:219–224. https://doi.org/10.1086/316945
Bockenhauer D, Jaureguiberry G (2016) HNF1B-associated clinical phenotypes: the kidney and beyond. Pediatr Nephrol 31:707–714. https://doi.org/10.1007/s00467-015-3142-2
Cubuk H, Yalcin Capan O (2021) A Review of functional characterization of single amino acid change mutations in HNF transcription Factors in MODY pathogenesis. Protein J 40:348–360. https://doi.org/10.1007/s10930-021-09991-8
Delaglio F, Grzesiek S, Vuister GW, Zhu G, Pfeifer J, Bax A (1995) NMRPipe: a multidimensional spectral processing system based on UNIX pipes. J Biomol NMR 6:277–293. https://doi.org/10.1007/BF00197809
El-Khairi R, Vallier L (2016) The role of hepatocyte nuclear factor 1beta in disease and development. Diabetes Obes Metab 18(Suppl 1):23–32. https://doi.org/10.1111/dom.12715
Fajans SS, Bell GI (2011) MODY: history, genetics, pathophysiology, and clinical decision making. Diabetes Care 34:1878–1884. https://doi.org/10.2337/dc11-0035
Heidet L, Decramer S, Pawtowski A, Moriniere V, Bandin F, Knebelmann B, Lebre AS, Faguer S, Guigonis V, Antignac C, Salomon R (2010) Spectrum of HNF1B mutations in a large cohort of patients who harbor renal diseases. Clin J Am Soc Nephrol 5:1079–1090. https://doi.org/10.2215/CJN.06810909
Herr W, Cleary MA (1995) The POU domain: versatility in transcriptional regulation by a flexible two-in-one DNA-binding domain. Genes Dev 9:1679–1693. https://doi.org/10.1101/gad.9.14.1679
Hyberts SG, Milbradt AG, Wagner AB, Arthanari H, Wagner G (2012) Application of iterative soft thresholding for fast reconstruction of NMR data non-uniformly sampled with multidimensional poisson gap scheduling. J Biomol NMR 52:315–327. https://doi.org/10.1007/s10858-012-9611-z
Johnson BA, Blevins RA (1994) NMR View: a computer program for the visualization and analysis of NMR data. J Biomol NMR 4:603–614. https://doi.org/10.1007/BF00404272
Kind L, Raasakka A, Molnes J, Aukrust I, Bjorkhaug L, Njolstad PR, Kursula P, Arnesen T (2022) Structural and biophysical characterization of transcription factor HNF-1A as a tool to study MODY3 diabetes variants. J Biol Chem 298:101803. https://doi.org/10.1016/j.jbc.2022.101803
Kobayashi N, Iwahara J, Koshiba S, Tomizawa T, Tochio N, Guntert P, Kigawa T, Yokoyama S (2007) KUJIRA, a package of integrated modules for systematic and interactive analysis of NMR data directed to high-throughput NMR structure studies. J Biomol NMR 39:31–52. https://doi.org/10.1007/s10858-007-9175-5
Lu P, Li Y, Gorman A, Chi YI (2006) Crystallization of hepatocyte nuclear factor 1beta in complex with DNA. Acta Crystallogr Sect F Struct Biol Cryst Commun 62:525–529. https://doi.org/10.1107/S1744309106015168
Lu P, Rha GB, Chi YI (2007) Structural basis of disease-causing mutations in hepatocyte nuclear factor 1beta. Biochemistry 46:12071–12080. https://doi.org/10.1021/bi7010527
Madariaga L, Moriniere V, Jeanpierre C, Bouvier R, Loget P, Martinovic J, Dechelotte P, Leporrier N, Thauvin-Robinet C, Jensen UB, Gaillard D, Mathieu M, Turlin B, Attie-Bitach T, Salomon R, Gubler MC, Antignac C, Heidet L (2013) Severe prenatal renal anomalies associated with mutations in HNF1B or PAX2 genes. Clin J Am Soc Nephrol 8:1179–1187. https://doi.org/10.2215/CJN.10221012
Peixoto-Barbosa R, Reis AF, Giuffrida FMA (2020) Update on clinical screening of maturity-onset diabetes of the young (MODY). Diabetol Metab Syndr 12:50. https://doi.org/10.1186/s13098-020-00557-9
Ryan AK, Rosenfeld MG (1997) POU domain family values: flexibility, partnerships, and developmental codes. Genes Dev 11:1207–1225. https://doi.org/10.1101/gad.11.10.1207
Shen Y, Bax A (2015) Protein structural information derived from NMR chemical shift with the neural network program TALOS-N. Methods Mol Biol 1260:17–32. https://doi.org/10.1007/978-1-4939-2239-0_2
Stenson PD, Mort M, Ball EV, Chapman M, Evans K, Azevedo L, Hayden M, Heywood S, Millar DS, Phillips AD, Cooper DN (2020) The human gene mutation database (HGMD((R))): optimizing its use in a clinical diagnostic or research setting. Hum Genet 139:1197–1207. https://doi.org/10.1007/s00439-020-02199-3
Thomas R, Sanna-Cherchi S, Warady BA, Furth SL, Kaskel FJ, Gharavi AG (2011) HNF1B and PAX2 mutations are a common cause of renal hypodysplasia in the CKiD cohort. Pediatr Nephrol 26:897–903. https://doi.org/10.1007/s00467-011-1826-9
Ying J, Delaglio F, Torchia DA, Bax A (2017) Sparse multidimensional iterative lineshape-enhanced (SMILE) reconstruction of both non-uniformly sampled and conventional NMR data. J Biomol NMR 68:101–118. https://doi.org/10.1007/s10858-016-0072-7
Funding
This study was supported by MEXT/JSPS KAKENHI (23K05720 to T.K.) and the Research Development Fund of Yokohama City University to T.K.
Author information
Authors and Affiliations
Contributions
S.H. and T.K. wrote the main manuscript text and prepared all figures and table. All authors reviewed the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Ethical approval
Not applicable.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Hokazono, S., Imagawa, E., Hirano, D. et al. 1H, 13C and 15N backbone resonance assignments of hepatocyte nuclear factor-1-beta (HNF1β) POUS and POUHD. Biomol NMR Assign 18, 59–63 (2024). https://doi.org/10.1007/s12104-024-10168-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12104-024-10168-4