Abstract
LEE-encoded effector EspF (EspF) is an effector protein part of enteropathogenic Escherichia coli’s (EPEC’s) arsenal for intestinal infection. This intrinsically disordered protein contains three highly conserved repeats which together compose over half of the protein’s complete amino acid sequence. EPEC uses EspF to hijack host proteins in order to promote infection. In the attack EspF is translocated, together with other effector proteins, to host cell via type III secretion system. Inside host EspF stimulates actin polymerization by interacting with Neural Wiskott-Aldrich syndrome protein (N-WASP), a regulator in actin polymerization machinery. It is presumed that EspF acts by disrupting the autoinhibitory state of N-WASP GTPase binding domain. In this NMR spectroscopy study, we report the 1H, 13C, and 15N resonance assignments for the complex formed by the first 47-residue repeat of EspF and N-WASP GTPase binding domain. These near-complete resonance assignments provide the basis for further studies which aim to characterize structure, interactions, and dynamics between these two proteins in solution.
Similar content being viewed by others
Biological context
An intrinsically disordered protein (IDP) is a functional polypeptide which cannot fold spontaneously into a stable three-dimensional conformation and extensive disorder is important for its function. Similar segments within a protein are called intrinsically disordered regions. IDPs are often described as important components of the cellular signalling machinery, especially in eukaryotes’ proteome where IDPs are abundant (van der Lee et al. 2014; Wright and Dyson 2015). Previous research has revealed that pathogens can produce protein mimics which target the components of the cellular signalling machinery. These mimics outcompete their model proteins and change the host metabolism making it favourable to the pathogen (Davey et al. 2011; Aitio et al. 2012; Tossavainen et al. 2016; Sámano-Sánchez and Gibson 2020).
LEE-encoded effector EspF (EspF) is one example of such a mimic, which is part of enteropathogenic Escherichia coli’s (EPEC’s) arsenal for intestinal infection. EspF is recognized as a type III effector protein and it is translocated from bacteria into host cells through a dedicated protein translocation apparatus, the type III secretion system. As a translocated effector, EspF interacts with cellular proteins and modifies their activity. EspF is also found in enterohaemorrhagic E. coli (EHEC) and Citrobacter rodentium. EPEC EspF is a 21 kDa IDP which contains three similar 40–47 amino acid proline-rich repeats at its C-terminus (McNamara and Donnenberg 1998; Alto et al. 2007). In this study, the first repeat (residues 73–119) was studied (Fig. 1a).
EspF repeats include amino acid sequences which mimic bona fide interaction sites in human proteins. The N-terminal proline-rich region has been shown to interact with the Src homology 3 (SH3) domain of sorting nexin 9 (SNX9). SNX9 is involved in clathrin-mediated endocytosis (Bendris and Schmid 2017). In the C-terminal half resides a putative Neural Wiskott-Aldrich syndrome protein (N-WASP) GTPase binding domain (GBD) binding motif (Marchès et al. 2006; Alto et al. 2007). N-WASP plays a role in actin polymerization machinery where it regulates actin filament branching and assembly through the activation of actin-related protein 2/3 (ARP2/3) complex. Alone in basal conditions N-WASP’s own autoinhibitory element, termed the C-helix, is bound to GBD domain’s hydrophobic core, keeping N-WASP inactive. For N-WASP activation, the autoinhibitory state is disrupted by the GTPase Cdc42 or other human signalling molecules. Binding of the GTPase Cdc42 to autoinhibited N-WASP leads to the release of the C-helix and exposes it for ARP2/3 complex interaction (Miki et al. 1998; Rohatgi et al. 1999; Sallee et al. 2008). It has been demonstrated that EHEC exploits this autoinhibitory interaction to promote pathogenesis. A related effector protein EspFU, expressed by EHEC only, can bind to N-WASP and to another Wiskott-Aldrich syndrome family protein WASP. In both interactions, EspFU mimics and outcompetes the C-helix in the hydrophobic core to activate the Wiskott-Aldrich syndrome family protein (Cheng et al. 2008; Sallee et al. 2008; Aitio et al. 2012). Because previous research has shown that EspF interacts with N-WASP, the same structural mechanism of C-helix mimicry and substitution has been implied for EspF and N-WASP (Alto et al. 2007; Garber et al. 2018).
Structures of EspFU in complex with N-WASP GBD and WASP GBD have been solved (Cheng et al. 2008; Aitio et al. 2012). To our knowledge, there is no published structural data on EspF in complex with N-WASP GDB. Here we report the near-complete 1H, 13C, and 15N resonance assignments for the complex formed by the first EspF repeat with N-WASP GBD. These initial studies enable further studies where the aim is to characterize structure, interactions, and dynamics between these two proteins in solution.
Methods and experiments
Protein expression and purification
Residues 73–119 were selected to represent the first repeat from EspF protein (UniProtKB B7UM88) and gene (synthetic, GenScript Inc., USA) encoding these residues was cloned to pET15b vector (Novagen) into the NdeI and XhoI sites. The protein construct carried an N-terminal His-Tag, followed by GB1 fusion protein and TEV protease (from Tobacco Etch Virus) cleavage site which was connected to EspF. Residues 207–270 were selected to represent GBD from N-WASP protein (UniProtKB O00401) and gene (synthetic, GenScript Inc., USA) encoding these residues was cloned to pET15b vector (Novagen) into the NdeI and XhoI sites. The protein construct carried an N-terminal His-Tag.
Production and purification of EspF repeat was carried out as described before (Karjalainen et al. 2020), whereas production of N-WASP GBD was done as described in Aitio et al. 2012. In the production of EspF repeat or N-WASP GBD, protein constructs expressing BL21(DE3) cells were grown in M9 minimal medium, supplemented with 1 g/l of 15NH4Cl and 2 g/l of 13C-d-glucose as the sole nitrogen and carbon sources, or in LB medium for obtaining unlabelled EspF repeat or N-WASP GBD. To purify N-WASP GBD, recovered clarified supernatant of His-Tagged N-WASP GBD protein was applied to the 1-ml His GraviTrap column (GE Healthcare) according to the manufacturer’s instructions. Imidazole was used to collect the N-WASP GBD protein and imidazole was removed from eluted proteins by PD-10 (GE Healthcare) before thrombin cleavage. The cleaved N-WASP GBD was concentrated and applied into the Superdex 75 (16/60). The columns were equilibrated with 20 mM sodium phosphate, pH 6.5, 50 mM NaCl buffer. Elution fractions containing purified proteins were pooled and concentrated by Vivaspin 2 (Sartorius Stedim). The gel filtration was performed by using ÄKTA Purifier FLPC purification system (GE Healthcare).
NMR spectroscopy
All NMR spectra of the complex formed by EspF repeat and N-WASP GBD were acquired at 298 K using a Bruker Avance III HD 800 MHz NMR spectrometer equipped with a helium cooled TCI 1H/13C/15N cryoprobe. For the resonance assignments of the binary complex formed by EspF repeat and N-WASP GBD two samples were used. One was composed of 15N, 13C labelled EspF repeat in complex with unlabelled N-WASP GBD, the other was composed of 15N, 13C labelled N-WASP GBD in complex with unlabelled EspF repeat. Ratios between two proteins were 1:1.2 and the labelled protein was always saturated with the unlabelled protein. Protein sample concentrations varied between 0.3 and 1.0 mM and samples were loaded into 5 mm Shigemi NMR tubes. Proteins were in 4 %/96% D2O/H2O, 20 mM sodium phosphate, 50 mM NaCl, pH 6.5 NMR buffer. Chemical shifts were referenced to external 2,2-dimethyl-2-silapentane-5-sulfonic acid (DSS). Resonance assignment was carried out with the following set of experiments: 1H, 15N HSQC, constant time 1H,13C HSQC for aliphatic and aromatic regions (Cavanagh 2007), HNCACB (Grzesiek and Bax 1992a), HN(CO)CACB (Grzesiek and Bax 1992b), HBHA(CO)NH, H(CC)(CO)NH, (H)CC(CO)NH, 1H, 15N NOESY-HSQC, 1H, 13C NOESY-HSQC for aliphatic and aromatic regions (Sattler et al. 1999), (HB)CB(CGCD)HD, (HB)CB(CGCDCE)HE (Yamazaki et al. 1993; Sattler et al. 1999), HC(C)H-COSY (Kay et al. 1993), DE-MQ-(H)CCmHm-TOCSY (Permi et al. 2004), 4D (HACA)CONCAHA (Tossavainen et al. 2020), and CON (Bermel et al. 2006). NMR data were processed with TopSpin 3.5 (Bruker Corporation) and analysed with CcpNmr Analysis 2.4.2 (Vranken et al. 2005). The missing proline assignments of the N-terminal proline-rich region of EspF repeat (residues 73, 76, and 78–80) were supplemented from assignments of the free form (unpublished data).
Assignments and data deposition
The amino acid sequences of the first EspF repeat (hereafter EspF) and N-WASP GBD are presented in Fig. 1a. For EspF, bioinformatics tools were used to analyse the sequence and predict intrinsic disorder (IUPred2A (Mészáros et al. 2018), DISOPRED3 (Jones and Cozzetto 2015), disCop (Fan and Kurgan 2014), and PrDOS (Ishida and Kinoshita 2007)). IUPred2A prediction was selected as the representative prediction and the result shows disordered characteristics for EspF (Fig. 1b). The prediction gives a near 1 score for residues in the first half of EspF, indicating that these are very likely to reside in a disordered region. The second half of the sequence shows a significantly more ordered tendency and has values at both sides of the (dis)ordered cutoff value 0.5.
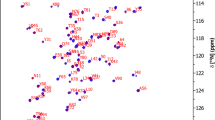
Because we are expecting a disorder-to-order transition for EspF upon binding to N-WASP GBD due to similarity to EspFU (Aitio et al. 2012), an additional ANCHOR2 prediction from IUPred2A is included (Fig. 1b). The prediction gives a close to 1 score for residues from 92P to 110S, indicating high probability of being part of a disordered binding region and a disorder-to-order transition upon binding to N-WASP GBD. Observations from the 1H, 15N-HSQC spectrum of bound EspF support this idea. The peaks of residues 93L to 114E have substantially shifted and become dispersed as compared to those in the spectrum of free EspF (unpublished data). The N-terminal half amide peaks, however, remain clustered in a narrow 1H region (Fig. 1d). These peaks have chemical shifts matching those of free EspF, which affirms the assumption that the N-terminal part is not involved in binding of N-WASP GBD.
For both proteins, extent of backbone assignments is shown in Fig. 1c. 1H, 15N-HSQC spectra highlighting the spectral quality together with the assignments are shown in Fig. 1d for EspF and in Fig. 1e for N-WASP GBD. For EspF, 15N 94%, 1HN 100 %, 13Cα 94 %, and 1Hα 100 % complete backbone assignments were attained. 13Cβ 93 % and 1Hβ 90 % were assigned. In total, side chain assignments were 84 % complete for achievable assignments. Missing assignments for prolines were supplemented from those of free EspF (unpublished data), assigned using 3D Hα-detected experiments (Mäntylahti et al. 2010, 2011). These included 15N assignments for prolines 73 and 76. Also, 15N, 13C’, and 1Hα assignments for prolines 78–80. As said, N-terminal residues do not participate in N-WASP GDB binding and proline shifts match those of the complex form.
15N 92 %, 1HN 92 %, 13Cα 100%, and 1Hα 100% complete backbone assignments for N-WASP GBD were obtained for residues 208–210, 212–252 and 254–270. Residues 207S, 211H, and 253K are missing backbone nitrogen and proton assignments. 13Cβ and 1Hβ were assigned completely. In total, side chain assignments were 92 % complete for achievable assignments. Both proteins had an N-terminal glycine residue as a cloning artefact and these are included in the reported chemical shifts.
a The amino acid sequences of EspF and N-WASP GBD used in this study. Underlined N-terminal glycine is not part of the native sequence. b Predicted disorder and disordered binding regions for EspF from IUPred2A and ANCHOR2. Both programs give a score between 0 and 1 to each residue which represents the probability for disorder (IUPred2A) or the probability for locating in a disordered binding region (ANCHOR2). A higher number corresponds to a higher probability. c Extent of backbone assignments of EspF and N-WASP GBD. d Assigned 1H, 15N-HSQC spectrum of 15N and 13C labelled EspF bound to unlabelled N-WASP. The shown contour level does not display 96I, but its peak is clearly visible at lower levels. The N-terminal half amide peaks are coloured orange to point out narrow 1H dispersion. e Assigned 1H, 15N-HSQC spectrum of 15N and 13C labelled N-WASP GBD bound to unlabelled EspF. Spectra were recorded at 800 MHz and 25 °C. The NH resonances are labelled with residue numbers and single letter amino acid codes. Asparagine δ and glutamine ε side chain resonances are connected by lines. Arginine and tryptophan side chain resonances are marked by asterisk
In this manuscript, we have reported nearly complete 1H, 13C, and 15N resonance assignments for the complex formed by the first EspF repeat with N-WASP GBD. These near-complete resonance assignments can be used in further studies where aim is to characterize structure, interactions, and dynamics between these two proteins in solution. The assigned 1H, 13C, and 15N chemical shifts have been deposited in the BioMagResBank (http://www.bmrb.wisc.edu/) database with the accession number 50548.
References
Aitio O, Hellman M, Skehan B, Kesti T, Leong JM, Saksela K, Permi P (2012) Enterohaemorrhagic Escherichia coli exploits a tryptophan switch to hijack host F-actin assembly. Structure 20:1692–1703. https://doi.org/10.1016/j.str.2012.07.015
Alto NM, Weflen AW, Rardin MJ, Yarar D, Lazar CS, Tonikian R, Koller A, Taylor SS, Boone C, Sidhu SS, Schmid SL, Hecht GA, Dixon JE (2007) The type III effector EspF coordinates membrane trafficking by the spatiotemporal activation of two eukaryotic signaling pathways. J Cell Biol 178:1265–1278. https://doi.org/10.1083/jcb.200705021
Bendris N, Schmid SL (2017) Endocytosis, metastasis and beyond: multiple facets of SNX9. Trends Cell Biol 27:189–200. https://doi.org/10.1016/j.tcb.2016.11.001
Bermel W, Bertini I, Felli IC, Kümmerle R, Pierattelli R (2006) Novel 13C direct detection experiments, including extension to the third dimension, to perform the complete assignment of proteins. J Magn Reson 178:56–64. https://doi.org/10.1016/j.jmr.2005.08.011
Cavanagh J (2007) Protein NMR spectroscopy: principles and practice. Academic Press, Amsterdam
Cheng H-C, Skehan BM, Campellone KG, Leong JM, Rosen MK (2008) Structural mechanism of WASP activation by the enterohaemorrhagic E. coli effector EspF U. Nature 454:1009–1013. https://doi.org/10.1038/nature07160
Davey NE, Travé G, Gibson TJ (2011) How viruses hijack cell regulation. Trends Biochem Sci 36:159–169. https://doi.org/10.1016/j.tibs.2010.10.002
Fan X, Kurgan L (2014) Accurate prediction of disorder in protein chains with a comprehensive and empirically designed consensus. J Biomol Struct Dyn 32:448–464. https://doi.org/10.1080/07391102.2013.775969
Garber JJ, Mallick EM, Scanlon KM, Turner JR, Donnenberg MS, Leong JM, Snapper SB (2018) Attaching-and-effacing pathogens exploit junction regulatory activities of N-WASP and SNX9 to disrupt the intestinal barrier. Cell Mol Gastroenterol Hepatol 5:273–288. https://doi.org/10.1016/j.jcmgh.2017.11.015
Grzesiek S, Bax A (1992a) An efficient experiment for sequential backbone assignment of medium-sized isotopically enriched proteins. J Magn Reson 99:201–207. https://doi.org/10.1016/0022-2364(92)90169-8
Grzesiek S, Bax A (1992b) Correlating backbone amide and side chain resonances in larger proteins by multiple relayed triple resonance NMR. J Am Chem Soc 114:6291–6293. https://doi.org/10.1021/ja00042a003
Ishida T, Kinoshita K (2007) PrDOS: prediction of disordered protein regions from amino acid sequence. Nucleic Acids Res 35:W460–W464. https://doi.org/10.1093/nar/gkm363
Jones DT, Cozzetto D (2015) DISOPRED3: precise disordered region predictions with annotated protein-binding activity. Bioinformatics 31:857–863. https://doi.org/10.1093/bioinformatics/btu744
Karjalainen M, Tossavainen H, Hellman M, Permi P (2020) HACANCOi: a new Hα-detected experiment for backbone resonance assignment of intrinsically disordered proteins. J Biomol NMR. https://doi.org/10.1007/s10858-020-00347-5
Kay LE, Xu GY, Singer AU, Muhandiram DR, Formankay JD (1993) A gradient-enhanced HCCH-TOCSY experiment for recording side-chain 1H and 13C correlations in H2O samples of proteins. J Magn Reson B 101:333–337. https://doi.org/10.1006/jmrb.1993.1053
Mäntylahti S, Hellman M, Permi P (2011) Extension of the HA-detection based approach: (HCA)CON(CA)H and (HCA)NCO(CA)H experiments for the main-chain assignment of intrinsically disordered proteins. J Biomol NMR 49:99–109. https://doi.org/10.1007/s10858-011-9470-z
Mäntylahti S, Aitio O, Hellman M, Permi P (2010) HA-detected experiments for the backbone assignment of intrinsically disordered proteins. J Biomol NMR 47:171–181. https://doi.org/10.1007/s10858-010-9421-0
Marchès O, Batchelor M, Shaw RK, Patel A, Cummings N, Nagai T, Sasakawa C, Carlsson SR, Lundmark R, Cougoule C, Caron E, Knutton S, Connerton I, Frankel G (2006) EspF of enteropathogenic Escherichia coli binds sorting nexin 9. J Bacteriol 188:3110–3115. https://doi.org/10.1128/JB.188.8.3110-3115.2006
McNamara BP, Donnenberg MS (1998) A novel proline-rich protein, EspF, is secreted from enteropathogenic Escherichia coli via the type III export pathway. FEMS Microbiol Lett 166:71–78. https://doi.org/10.1016/S0378-1097(98)00313-9
Mészáros B, Erdős G, Dosztányi Z (2018) IUPred2A: context-dependent prediction of protein disorder as a function of redox state and protein binding. Nucleic Acids Res 46:W329–W337. https://doi.org/10.1093/nar/gky384
Miki H, Sasaki T, Takai Y, Takenawa T (1998) Induction of filopodium formation by a WASP-related actin-depolymerizing protein N-WASP. Nature 391:93–96. https://doi.org/10.1038/34208
Permi P, Tossavainen H, Hellman M (2004) Efficient assignment of methyl resonances: enhanced sensitivity by gradient selection in a DE-MQ–(H)CCmHt m–TOCSY experiment. J Biomol NMR 30:275–282. https://doi.org/10.1007/s10858-004-3222-2
Rohatgi R, Ma L, Miki H, Lopez M, Kirchhausen T, Takenawa T, Kirschner MW (1999) The interaction between N-WASP and the Arp2/3 complex links Cdc42-dependent signals to actin assembly. Cell 97:221–231. https://doi.org/10.1016/S0092-8674(00)80732-1
Sallee NA, Rivera GM, Dueber JE, Vasilescu D, Mullins RD, Mayer BJ, Lim WA (2008) The pathogen protein EspFU hijacks actin polymerization using mimicry and multivalency. Nature 454:1005–1008. https://doi.org/10.1038/nature07170
Sámano-Sánchez H, Gibson TJ (2020) Mimicry of short linear motifs by bacterial pathogens: a drugging opportunity. Trends Biochem Sci 45:526–544. https://doi.org/10.1016/j.tibs.2020.03.003
Sattler M, Schleucher J, Griesinger C (1999) Heteronuclear multidimensional NMR experiments for the structure determination of proteins in solution employing pulsed field gradients. Prog Nucl Magn Reson Spectrosc 34:93–158. https://doi.org/10.1016/S0079-6565(98)00025-9
Tossavainen H, Aitio O, Hellman M, Saksela K, Permi P (2016) Structural basis of the high affinity interaction between the alphavirus nonstructural protein-3 (nsP3) and the SH3 domain of amphiphysin-2. J Biol Chem 291:16307–16317. https://doi.org/10.1074/jbc.M116.732412
Tossavainen H, Salovaara S, Hellman M, Ihalin R, Permi P (2020) Dispersion from Cα or NH: 4D experiments for backbone resonance assignment of intrinsically disordered proteins. J Biomol NMR 74:147–159. https://doi.org/10.1007/s10858-020-00299-w
van der Lee R, Buljan M, Lang B, Weatheritt RJ, Daughdrill GW, Dunker AK, Fuxreiter M, Gough J, Gsponer J, Jones DT, Kim PM, Kriwacki RW, Oldfield CJ, Pappu RV, Tompa P, Uversky VN, Wright PE, Babu MM (2014) Classification of intrinsically disordered regions and proteins. Chem Rev 114:6589–6631. https://doi.org/10.1021/cr400525m
Vranken WF, Boucher W, Stevens TJ, Fogh RH, Pajon A, Llinas M, Ulrich EL, Markley JL, Ionides J, Laue ED (2005) The CCPN data model for NMR spectroscopy: development of a software pipeline. Proteins Struct Funct Bioinform 59:687–696. https://doi.org/10.1002/prot.20449
Wright PE, Dyson HJ (2015) Intrinsically disordered proteins in cellular signalling and regulation. Nat Rev Mol Cell Biol 16:18–29. https://doi.org/10.1038/nrm3920
Yamazaki T, Forman-Kay JD, Kay LE (1993) Two-dimensional NMR experiments for correlating carbon-13.beta. and proton.delta./.epsilon. chemical shifts of aromatic residues in 13C-labeled proteins via scalar couplings. J Am Chem Soc 115:11054–11055. https://doi.org/10.1021/ja00076a099
Acknowledgements
This work is supported by the grant from the Academy of Finland (Number 288235 to Perttu Permi). We thank Laura Pitkänen for excellent technical assistance.
Funding
Open Access funding provided by University of Jyväskylä (JYU).
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Karjalainen, M., Hellman, M., Tossavainen, H. et al. 1H, 13C, and 15N NMR chemical shift assignment of the complex formed by the first EPEC EspF repeat and N-WASP GTPase binding domain. Biomol NMR Assign 15, 213–217 (2021). https://doi.org/10.1007/s12104-021-10008-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12104-021-10008-9