Abstract
Background
Osteosarcoma (OS) is a form of primary bone malignancy associated with poor prognostic outcomes. Recent work has highlighted vasculogenic mimicry (VM) as a key mechanism that supports aggressive tumor growth. The patterns of VM-associated gene expression in OS and the relationship between these genes and patient outcomes, however, have yet to be defined.
Methods
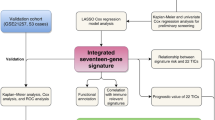
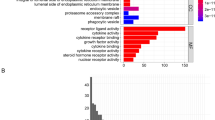
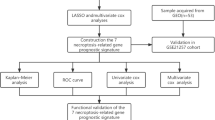
Here, 48 VM-related genes were systematically assessed to examine correlations between the expression of these genes and OS patient prognosis in the Therapeutically Applicable Research to Generate Effective Treatments (TARGET) cohort. Patients were classified into three OS subtypes. Differentially expressed genes for these three OS subtypes were then compared with hub genes detected in a weighted gene co-expression network analysis, leading to the identification of 163 overlapping genes that were subject to further biological activity analyses. A three-gene signature (CGREF1, CORT, and GALNT14) was ultimately constructed through a least absolute shrinkage and selection operator Cox regression analysis, and this signature was used to separate patients into low- and high-risk groups. The K–M survival analysis, receiver operating characteristic analysis, and decision curve analysis were adopted to evaluate the prognostic prediction performance of the signature. Furthermore, the expression patterns of three genes derived from the prognostic model were validated by quantitative real-time polymerase chain reaction (RT-qPCR).
Results
VM-associated gene expression patterns were successfully established, and three VM subtypes of OS that were associated with patient prognosis and copy number variants were defined. The developed three-gene signature was constructed, which served as independent prognostic markers and prediction factors for the clinicopathological features of OS. Finally, lastly, the signature may also have a guiding effect on the sensitivity of different chemotherapeutic drugs.
Conclusion
Overall, these analyses facilitated the development of a prognostic VM-associated gene signature capable of predicting OS patient outcomes. This signature may be of value for both studies of the mechanistic basis for VM and clinical decision-making in the context of OS patient management.









Similar content being viewed by others
Data availability
The datasets analyzed for this study can be found in the public database. All data generated or analyzed during this study are included as supplementary information file.
References
Ritter J, Bielack SS. Osteosarcoma. Ann Oncol. 2010;21(Suppl 7):vii320–5.
Kansara M, Teng MW, Smyth MJ, Thomas DM. Translational biology of osteosarcoma. Nat Rev Cancer. 2014;14:722–35.
Grimer RJ. Surgical options for children with osteosarcoma. Lancet Oncol. 2005;6:85–92.
Zils K, Bielack S, Wilhelm M, Werner M, Schwarz R, Windhager R, et al. Osteosarcoma of the mobile spine. Ann Oncol. 2013;24:2190–5.
Loh AH, Wu H, Bahrami A, Navid F, McCarville MB, Wang C, et al. Influence of bony resection margins and surgicopathological factors on outcomes in limb-sparing surgery for extremity osteosarcoma. Pediatr Blood Cancer. 2015;62:246–51.
Zhang W, Bado I, Wang H, Lo HC, Zhang XH. Bone metastasis: find your niche and fit in. Trends Cancer. 2019;5:95–110.
Maniotis AJ, Folberg R, Hess A, Seftor EA, Gardner LM, Pe’er J, et al. Vascular channel formation by human melanoma cells in vivo and in vitro: vasculogenic mimicry. Am J Pathol. 1999;155:739–52.
Shirakawa K, Kobayashi H, Sobajima J, Hashimoto D, Shimizu A, Wakasugi H. Inflammatory breast cancer: vasculogenic mimicry and its hemodynamics of an inflammatory breast cancer xenograft model. Breast Cancer Res. 2003;5:136–9.
Tang HS, Feng YJ, Yao LQ. Angiogenesis, vasculogenesis, and vasculogenic mimicry in ovarian cancer. Int J Gynecol Cancer. 2009;19:605–10.
Li M, Gu Y, Zhang Z, Zhang S, Zhang D, Saleem AF, et al. Vasculogenic mimicry: a new prognostic sign of gastric adenocarcinoma. Pathol Oncol Res. 2010;16:259–66.
Williamson SC, Metcalf RL, Trapani F, Mohan S, Antonello J, Abbott B, et al. Vasculogenic mimicry in small cell lung cancer. Nat Commun. 2016;7:13322.
Liu R, Yang K, Meng C, Zhang Z, Xu Y. Vasculogenic mimicry is a marker of poor prognosis in prostate cancer. Cancer Biol Ther. 2012;13:527–33.
Seftor RE, Hess AR, Seftor EA, Kirschmann DA, Hardy KM, Margaryan NV, et al. Tumor cell vasculogenic mimicry: from controversy to therapeutic promise. Am J Pathol. 2012;181:1115–25.
Kirschmann DA, Seftor EA, Hardy KM, Seftor RE, Hendrix MJ. Molecular pathways: vasculogenic mimicry in tumor cells: diagnostic and therapeutic implications. Clin Cancer Res. 2012;18:2726–32.
Massimini M, Romanucci M, De Maria R, Della SL. An update on molecular pathways regulating vasculogenic mimicry in human osteosarcoma and their role in canine oncology. Front Vet Sci. 2021;8: 722432.
Livak KJ, Schmittgen TD. Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) Method. Methods. 2001;25:402–8.
Lee BT, Barber GP, Benet-Pagès A, Casper J, Clawson H, Diekhans M, et al. The UCSC Genome Browser database: 2022 update. Nucleic Acids Res. 2022;50:D1115–22.
GTEx Consortium. Human genomics. The Genotype-Tissue Expression (GTEx) pilot analysis: multitissue gene regulation in humans. Science. 2015;348:648–60.
Leek JT, Johnson WE, Parker HS, Jaffe AE, Storey JD. The sva package for removing batch effects and other unwanted variation in high-throughput experiments. Bioinformatics. 2012;28:882–3.
Hendrix MJ, Seftor EA, Hess AR, Seftor RE. Vasculogenic mimicry and tumour-cell plasticity: lessons from melanoma. Nat Rev Cancer. 2003;3:411–21.
Delgado-Bellido D, Serrano-Saenz S, Fernández-Cortés M, Oliver FJ. Vasculogenic mimicry signaling revisited: focus on non-vascular VE-cadherin. Mol Cancer. 2017;16:65.
Wei X, Chen Y, Jiang X, Peng M, Liu Y, Mo Y, et al. Mechanisms of vasculogenic mimicry in hypoxic tumor microenvironments. Mol Cancer. 2021;20:7.
Kawahara R, Niwa Y, Simizu S. Integrin β1 is an essential factor in vasculogenic mimicry of human cancer cells. Cancer Sci. 2018;109:2490–6.
Ritchie ME, Phipson B, Wu D, Hu Y, Law CW, Shi W, et al. limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 2015;43: e47.
Szklarczyk D, Gable AL, Nastou KC, Lyon D, Kirsch R, Pyysalo S, et al. The STRING database in 2021: customizable protein-protein networks, and functional characterization of user-uploaded gene/measurement sets. Nucleic Acids Res. 2021;49:D605–12.
Seiler M, Huang CC, Szalma S, Bhanot G. ConsensusCluster: a software tool for unsupervised cluster discovery in numerical data. OMICS. 2010;14:109–13.
Hänzelmann S, Castelo R, Guinney J. GSVA: gene set variation analysis for microarray and RNA-seq data. BMC Bioinformatics. 2013;14:7.
Rich JT, Neely JG, Paniello RC, Voelker CC, Nussenbaum B, Wang EW. A practical guide to understanding Kaplan-Meier curves. Otolaryngol Head Neck Surg. 2010;143:331–6.
Barbie DA, Tamayo P, Boehm JS, Kim SY, Moody SE, Dunn IF, et al. Systematic RNA interference reveals that oncogenic KRAS-driven cancers require TBK1. Nature. 2009;462:108–12.
Mermel CH, Schumacher SE, Hill B, Meyerson ML, Beroukhim R, Getz G. GISTIC2.0 facilitates sensitive and confident localization of the targets of focal somatic copy-number alteration in human cancers. Genome Biol. 2011;12:R41.
Mayakonda A, Lin DC, Assenov Y, Plass C, Koeffler HP. Maftools: efficient and comprehensive analysis of somatic variants in cancer. Genome Res. 2018;28:1747–56.
Langfelder P, Horvath S. WGCNA: an R package for weighted correlation network analysis. BMC Bioinformatics. 2008;9:559.
Yu G, Wang LG, Han Y, He QY. clusterProfiler: an R package for comparing biological themes among gene clusters. OMICS. 2012;16:284–7.
Friedman J, Hastie T, Tibshirani R. Regularization paths for generalized linear models via coordinate descent. J Stat Softw. 2010;33:1–22.
Park SY. Nomogram: an analogue tool to deliver digital knowledge. J Thorac Cardiovasc Surg. 2018;155:1793.
Vickers AJ, Elkin EB. Decision curve analysis: a novel method for evaluating prediction models. Med Decis Making. 2006;26:565–74.
Geeleher P, Cox N, Huang RS. pRRophetic: an R package for prediction of clinical chemotherapeutic response from tumor gene expression levels. PLoS ONE. 2014;9: e107468.
Qiao L, Liang N, Zhang J, Xie J, Liu F, Xu D, et al. Advanced research on vasculogenic mimicry in cancer. J Cell Mol Med. 2015;19:315–26.
Deng W, Wang L, Xiong Y, Li J, Wang Y, Shi T, et al. The novel secretory protein CGREF1 inhibits the activation of AP-1 transcriptional activity and cell proliferation. Int J Biochem Cell Biol. 2015;65:32–9.
Kang X, Xu E, Wang X, Qian L, Yang Z, Yu H, et al. Tenascin-c knockdown suppresses vasculogenic mimicry of gastric cancer by inhibiting ERK- triggered EMT. Cell Death Dis. 2021;12:890.
de Lecea L, Ruiz-Lozano P, Danielson PE, Peelle-Kirley J, Foye PE, Frankel WN, et al. Cloning, mRNA expression, and chromosomal mapping of mouse and human preprocortistatin. Genomics. 1997;42:499–506.
de Lecea L, del Rio JA, Criado JR, Alcántara S, Morales M, Danielson PE, et al. Cortistatin is expressed in a distinct subset of cortical interneurons. J Neurosci. 1997;17:5868–80.
de Lecea L, Criado JR, Prospero-Garcia O, Gautvik KM, Schweitzer P, Danielson PE, et al. A cortical neuropeptide with neuronal depressant and sleep-modulating properties. Nature. 1996;381:242–5.
Cassoni P, Muccioli G, Marrocco T, Volante M, Allia E, Ghigo E, et al. Cortistatin-14 inhibits cell proliferation of human thyroid carcinoma cell lines of both follicular and parafollicular origin. J Endocrinol Invest. 2002;25:362–8.
Cassoni P, Allia E, Marrocco T, Ghè C, Ghigo E, Muccioli G, et al. Ghrelin and cortistatin in lung cancer: expression of peptides and related receptors in human primary tumors and in vitro effect on the H345 small cell carcinoma cell line. J Endocrinol Invest. 2006;29:781–90.
Li M, Yan S, Fisher WE, Chen C, Yao Q. New roles of a neuropeptide cortistatin in the immune system and cancer. World J Surg. 2005;29:354–6.
Tarp MA, Clausen H. Mucin-type O-glycosylation and its potential use in drug and vaccine development. Biochim Biophys Acta. 2008;1780:546–63.
Ten Hagen KG, Fritz TA, Tabak LA. All in the family: the UDP-GalNAc:polypeptide N-acetylgalactosaminyltransferases. Glycobiology. 2003;13:1r–16r.
Song KH, Park MS, Nandu TS, Gadad S, Kim SC, Kim MY. GALNT14 promotes lung-specific breast cancer metastasis by modulating self-renewal and interaction with the lung microenvironment. Nat Commun. 2016;7:13796.
Yang J, Li G, Zhang K. MiR-125a regulates ovarian cancer proliferation and invasion by repressing GALNT14 expression. Biomed Pharmacother. 2016;80:381–7.
Hattinger CM, Fanelli M, Tavanti E, Vella S, Ferrari S, Picci P, et al. Advances in emerging drugs for osteosarcoma. Expert Opin Emerg Drugs. 2015;20:495–514.
Koshkina NV, Kleinerman ES, Waidrep C, Jia SF, Worth LL, Gilbert BE, et al. 9-Nitrocamptothecin liposome aerosol treatment of melanoma and osteosarcoma lung metastases in mice. Clin Cancer Res. 2000;6:2876–80.
Lu K, Wang X, Chen Y, Liang D, Luo H, Long L, et al. Identification of two potential glycogen synthase kinase 3β inhibitors for the treatment of osteosarcoma. Acta Biochim Biophys Sin (Shanghai). 2018;50:456–64.
Nishimura H, Nakamura O, Yamagami Y, Mori M, Horie R, Fukuoka N, et al. GSK-3 inhibitor inhibits cell proliferation and induces apoptosis in human osteosarcoma cells. Oncol Rep. 2016;35:2348–54.
Fu Z, Deng B, Liao Y, Shan L, Yin F, Wang Z, et al. The anti-tumor effect of shikonin on osteosarcoma by inducing RIP1 and RIP3 dependent necroptosis. BMC Cancer. 2013;13:580.
Brounais B, Chipoy C, Mori K, Charrier C, Battaglia S, Pilet P, et al. Oncostatin M induces bone loss and sensitizes rat osteosarcoma to the antitumor effect of Midostaurin in vivo. Clin Cancer Res. 2008;14:5400–9.
Acknowledgements
The authors gratefully acknowledge contributions from the GTEx, TARGET, and STRING databases for free use and the reviewers for their helpful comments on this study.
Funding
Sponsorship for this study was funded by grants from the National Natural Science Foundation of China (No. 81802204), China Postdoctoral Science Foundation (2020M671453).
Author information
Authors and Affiliations
Contributions
Conceptualization, L.Y., R.L, and B.W.; methodology, L.Y., D.L., and R.L; software, R.L; validation, Z.L., Y.Z., and B.W.; formal analysis, L.Y. and R.L.; data curation, R.L. and D.L.; writing—original draft preparation, L.Y. and R.L.; writing—review and editing, Y.Z. and B.W.; visualization, L.Y. and R.L.; supervision, Z.L. and B.W.; project administration, B.W.; funding acquisition, B.W.
Corresponding authors
Ethics declarations
Conflict of interest
All named authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Ethical approval
We performed this research though publicly available datasets. There was no ethical approval required.
Consent for publication
All authors have read and agreed to all the contents for publication.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Yan, L., Li, R., Li, D. et al. Development of a novel vasculogenic mimicry-associated gene signature for the prognostic assessment of osteosarcoma patients. Clin Transl Oncol 25, 3501–3518 (2023). https://doi.org/10.1007/s12094-023-03218-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12094-023-03218-1