Abstract
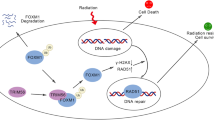
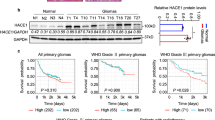
Glioblastoma (GBM) is the most common and lethal subtype of glioma, characterized by uncontrolled cancer cell proliferation, extensive infiltration, and therapeutic resistance. Ring finger protein 216 (RNF216) is a RING-type E3 ubiquitin ligase aberrantly expressed in multiple human cancers. Tumor protein 53 (p53) is a transcription factor that acts as a tumor suppressor. This study aimed to compare the RNF216 expression in GBM tissues and normal peritumoral tissues and to examine the effects of RNF216 overexpression/knockdown on tumorigenesis, radioresistance, and the p53 pathway in GBM. The results showed that RNF216 was overexpressed in GBM tissues and cell lines, and high RNF216 expression was related to a poor prognosis. RNF216 overexpression promoted GBM cell growth and inhibited apoptosis, while RNF216 knockdown impaired GBM cell growth and enhanced cell death. RNF216 was also highly expressed in recurrent GBM tissues compared with paired primary tumors. The upregulation of RNF216 not only facilitated GBM cell growth but also protected cells against X-ray irradiation-induced apoptosis and DNA damage, while RNF216 knockdown exerted opposite effects. Moreover, the implantation of GBM cells with RNF216 silencing suppressed tumorigenesis and increased radiosensitivity of mice bearing GBM xenografts. Further analysis revealed that RNF216 overexpression reduced the stability of p53 protein via ubiquitination and negatively regulated the p53 pathway, while RNF216 knockdown preserved the p53 protein. In conclusion, RNF216 effectively attenuated radiation-induced apoptosis and DNA damage in GBM via inducing ubiquitination-mediated degradation of p53. These findings suggest the potential therapeutic use of RNF216 inhibition for tumorigenesis and therapeutic resistance in GBM.







Similar content being viewed by others
Data Availability
All data generated or analyzed during this study are included in this published article.
References
Davis ME (2018) Epidemiology and overview of gliomas. Semin Oncol Nurs 34:420–429. https://doi.org/10.1016/j.soncn.2018.10.001
Ostrom QT, Gittleman H, Stetson L, Virk SM, Barnholtz-Sloan JS (2015) Epidemiology of gliomas. Cancer Treat Res 163:1–14. https://doi.org/10.1007/978-3-319-12048-5_1
DeCordova S, Shastri A, Tsolaki AG et al (2020) Molecular heterogeneity and immunosuppressive microenvironment in glioblastoma. Front Immunol 11:1402. https://doi.org/10.3389/fimmu.2020.01402
Hambardzumyan D, Bergers G (2015) Glioblastoma: defining tumor niches. Trends in cancer 1:252–265. https://doi.org/10.1016/j.trecan.2015.10.009
Davis ME (2016) Glioblastoma: overview of disease and treatment. Clin J Oncol Nurs 20:S2-8. https://doi.org/10.1188/16.cjon.s1.2-8
Wilson TA, Karajannis MA, Harter DH (2014) Glioblastoma multiforme: state of the art and future therapeutics. Surg Neurol Int 5:64. https://doi.org/10.4103/2152-7806.132138
McLendon RE, Halperin EC (2003) Is the long-term survival of patients with intracranial glioblastoma multiforme overstated? Cancer 98:1745–1748. https://doi.org/10.1002/cncr.11666
Melnick AF, Gao Y, Liu J et al (2019) RNF216 is essential for spermatogenesis and male fertility†. Biol Reprod 100:1132–1134. https://doi.org/10.1093/biolre/ioz006
Chuang TH, Ulevitch RJ (2004) Triad3A, an E3 ubiquitin-protein ligase regulating Toll-like receptors. Nat Immunol 5:495–502. https://doi.org/10.1038/ni1066
Fearns C, Pan Q, Mathison JC, Chuang TH (2006) Triad3A regulates ubiquitination and proteasomal degradation of RIP1 following disruption of Hsp90 binding. J Biol Chem 281:34592–34600. https://doi.org/10.1074/jbc.M604019200
Wang H, Wang Y, Qian L et al (2016) RNF216 contributes to proliferation and migration of colorectal cancer via suppressing BECN1-dependent autophagy. Oncotarget 7:51174–83. https://doi.org/10.18632/oncotarget.9433
Tu J, Chen W, Meng M et al (2020) RNF216 promotes occurrence and progression of cholangiocarcinoma via regulation of the DIAPH3 ubiquitination. Res Sq. https://doi.org/10.21203/rs.3.rs-111960/v1
Sullivan KD, Galbraith MD, Andrysik Z, Espinosa JM (2018) Mechanisms of transcriptional regulation by p53. Cell Death Differ 25:133–143. https://doi.org/10.1038/cdd.2017.174
Olivier M, Hollstein M, Hainaut P (2010) TP53 mutations in human cancers: origins, consequences, and clinical use. Cold Spring Harb Perspect Biol 2:a001008. https://doi.org/10.1101/cshperspect.a001008
Bode AM, Dong Z (2004) Post-translational modification of p53 in tumorigenesis. Nat Rev Cancer 4:793–805. https://doi.org/10.1038/nrc1455
Zou S, Zhu Y, Wang B et al (2017) The ubiquitin ligase COP1 promotes glioma cell proliferation by preferentially downregulating tumor suppressor p53. Mol Neurobiol 54:5008–5016. https://doi.org/10.1007/s12035-016-0033-x
Zhang J, Zhang C, Cui J et al (2017) TRIM45 functions as a tumor suppressor in the brain via its E3 ligase activity by stabilizing p53 through K63-linked ubiquitination. Cell Death Dis 8:e2831. https://doi.org/10.1038/cddis.2017.149
National Research Council Committee for the Update of the Guide for the C, Use of Laboratory A. The National Academies Collection: Reports funded by National Institutes of Health. In: th, editor. Guide for the Care and Use of Laboratory Animals. Washington (DC): National Academies Press (US) National Academy of Sciences.; 2011.
Wang X, Liu X, Chen Y et al (2015) Histopathological findings in the peritumoral edema area of human glioma. Histol Histopathol 30:1101–9. https://doi.org/10.14670/hh-11-607
Louis DN, Perry A, Wesseling P et al (2021) The 2021 WHO Classification of Tumors of the Central Nervous System: a summary. Neuro Oncol 23:1231–1251. https://doi.org/10.1093/neuonc/noab106
Yuan JX, Munson JM (2017) Quantitative immunohistochemistry of the cellular microenvironment in patient glioblastoma resections. J Vis Exp: JoVE. https://doi.org/10.3791/56025
Irtenkauf SM, Sobiechowski S, Hasselbach LA et al (2017) Optimization of glioblastoma mouse orthotopic xenograft models for translational research. Comp Med 67:300–314
Rajaratnam V, Islam MM, Yang M, Slaby R, Ramirez HM, Mirza SP (2020) Glioblastoma: pathogenesis and current status of chemotherapy and other novel treatments. Cancers 12https://doi.org/10.3390/cancers12040937
Guan R, Cai S, Sun M, Xu M (2017) Upregulation of miR-520b promotes ovarian cancer growth. Oncol Lett 14:3155–3161. https://doi.org/10.3892/ol.2017.6552
Osuka S, Van Meir EG (2017) Overcoming therapeutic resistance in glioblastoma: the way forward. J Clin Investig 127:415–426. https://doi.org/10.1172/jci89587
Ali MY, Oliva CR, Noman ASM et al (2020) Radioresistance in glioblastoma and the development of radiosensitizers. Cancers 12https://doi.org/10.3390/cancers12092511
Golding SE, Morgan RN, Adams BR, Hawkins AJ, Povirk LF, Valerie K (2009) Pro-survival AKT and ERK signaling from EGFR and mutant EGFRvIII enhances DNA double-strand break repair in human glioma cells. Cancer Biol Ther 8:730–738. https://doi.org/10.4161/cbt.8.8.7927
King HO, Brend T, Payne HL et al (2017) RAD51 is a selective DNA repair target to radiosensitize glioma stem cells. Stem cell reports 8:125–139. https://doi.org/10.1016/j.stemcr.2016.12.005
Timme CR, Rath BH, O’Neill JW, Camphausen K, Tofilon PJ (2018) The DNA-PK inhibitor VX-984 enhances the radiosensitivity of glioblastoma cells grown in vitro and as orthotopic xenografts. Mol Cancer Ther 17:1207–1216. https://doi.org/10.1158/1535-7163.mct-17-1267
Scholz N, Kurian KM, Siebzehnrubl FA, Licchesi JDF (2020) Targeting the ubiquitin system in glioblastoma. Front Oncol 10:574011. https://doi.org/10.3389/fonc.2020.574011
Eitel JA, Bijangi-Vishehsaraei K, Saadatzadeh MR et al (2009) PTEN and p53 are required for hypoxia induced expression of maspin in glioblastoma cells. Cell Cycle (Georgetown, Tex) 8:896–901. https://doi.org/10.4161/cc.8.6.7899
Zhang Y, Dube C, Gibert M, Jr. et al (2018) The p53 pathway in glioblastoma. Cancers 10https://doi.org/10.3390/cancers10090297
Wang Z, Yu G, Liu Z et al (2018) Paeoniflorin inhibits glioblastoma growth in vivo and in vitro: a role for the Triad3A-dependent ubiquitin proteasome pathway in TLR4 degradation. Cancer Manag Res 10:887–897. https://doi.org/10.2147/cmar.s160292
Author information
Authors and Affiliations
Contributions
Songwang Xie and Zhen Hong designed the study and supervised the data collection; Yan Li and Junyong Wang analyzed the data and interpreted the data; Jian Wang, Shaoquan Li, and Yongchang Liu prepared the manuscript for publication and reviewed the draft of the manuscript. All authors have read and approved the manuscript.
Corresponding author
Ethics declarations
Ethics Approval
All procedures performed in studies involving human participants were in accordance with the standards upheld by the Ethics Committee of Cangzhou Central Hospital and with those of the 1964 Helsinki Declaration and its later amendments for ethical research involving human subjects. All animal experiments were approved by the Ethics Committee of Cangzhou Central Hospital for the use of animals and conducted in accordance with the National Institutes of Health Laboratory Animal Care and Use Guidelines.
Consent to Participate
Written informed consent was obtained from a legally authorized representative(s) for anonymized patient information to be published in this article.
Consent for Publication
Not applicable.
Competing Interests
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Supplementary file 1
The RNF216 expression and survival of GBM patients were analyzed using the CGGA dataset (http://www.cgga.org.cn/). (JPG 6109 KB)
Supplementary file 2
LN229 and A172 cells were treated with 10 µg/mL CHX for 0–60 min. The protein level of p53 at different time points was assessed by Western blot. (JPG 266 KB)
Rights and permissions
About this article
Cite this article
Xie, S., Hong, Z., Li, Y. et al. RNF216 Alleviates Radiation-Induced Apoptosis and DNA Damage Through Regulating Ubiquitination-Mediated Degradation of p53 in Glioblastoma. Mol Neurobiol 59, 4703–4717 (2022). https://doi.org/10.1007/s12035-022-02868-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12035-022-02868-6