Abstract
In mammals, photoreceptor loss causes permanent blindness, but in zebrafish (Danio rerio), photoreceptor loss reprograms Müller glia to function as stem cells, producing progenitors that regenerate photoreceptors. MicroRNAs (miRNAs) regulate CNS neurogenesis, but the roles of miRNAs in injury-induced neuronal regeneration are largely unknown. In the embryonic zebrafish retina, miR-18a regulates photoreceptor differentiation. The purpose of the current study was to determine, in zebrafish, the function of miR-18a during injury-induced photoreceptor regeneration. RT-qPCR, in situ hybridization, and immunohistochemistry showed that miR-18a expression increases throughout the retina between 1 and 5 days post-injury (dpi). To test miR-18a function during photoreceptor regeneration, we used homozygous miR-18a mutants (miR-18ami5012), and knocked down miR-18a with morpholino oligonucleotides. During photoreceptor regeneration, miR-18ami5012 retinas have fewer mature photoreceptors than WT at 7 and 10 dpi, but there is no difference at 14 dpi, indicating that photoreceptor regeneration is delayed. Labeling dividing cells with 5-bromo-2′-deoxyuridine (BrdU) showed that at 7 and 10 dpi, there are excess dividing progenitors in both mutants and morphants, indicating that miR-18a negatively regulates injury-induced proliferation. Tracing 5-ethynyl-2′-deoxyuridine (EdU) and BrdU-labeled cells showed that in miR-18ami5012 retinas excess progenitors migrate to other retinal layers in addition to the photoreceptor layer. Inflammation is critical for photoreceptor regeneration, and RT-qPCR showed that in miR-18ami5012 retinas, inflammatory gene expression and microglia activation are prolonged. Suppressing inflammation with dexamethasone rescues the miR-18ami5012 phenotype. Together, these data show that in the injured zebrafish retina, disruption of miR-18a alters proliferation, inflammation, the microglia/macrophage response, and the timing of photoreceptor regeneration.











Similar content being viewed by others
Data availability
All data and miR-18ami5012 fish are freely available upon request.
References
Hitchcock P, Ochocinska M, Sieh A, Otteson D (2004) Persistent and injury-induced neurogenesis in the vertebrate retina. Prog Retin Eye Res 23(2):183–194. https://doi.org/10.1016/j.preteyeres.2004.01.001
Bernardos RL, Barthel LK, Meyers JR, Raymond PA (2007) Late-stage neuronal progenitors in the retina are radial Müller glia that function as retinal stem cells. J Neurosci 27(26):7028–7040. https://doi.org/10.1523/JNEUROSCI.1624-07.2007
Goldman D (2014) Müller glial cell reprogramming and retina regeneration. Nat Chem Biol:1–12. https://doi.org/10.1038/nrn3723
Lenkowski JR, Raymond PA (2014) Müller glia: stem cells for generation and regeneration of retinal neurons in teleost fish. Prog Retin Eye Res 40C:94–123. https://doi.org/10.1016/j.preteyeres.2013.12.007
Brockerhoff SE, Fadool JM (2011) Genetics of photoreceptor degeneration and regeneration in zebrafish. Cell Mol Life Sci 68(4):651–659. https://doi.org/10.1007/s00018-010-0563-8
Pollak J, Wilken MS, Ueki Y, Cox KE, Sullivan JM, Taylor RJ, Levine EM, Reh TA (2013) ASCL1 reprograms mouse Muller glia into neurogenic retinal progenitors. Development 140(12):2619–2631. https://doi.org/10.1242/dev.091355
Jorstad NL, Wilken MS, Grimes WN, Wohl SG, VandenBosch LS, Yoshimatsu T, Wong RO, Rieke F, Reh TA (2017) Stimulation of functional neuronal regeneration from Müller glia in adult mice. Nature 548(7665):103–107. https://doi.org/10.1038/nature23283
Yao K, Qiu S, Wang YV, Park SJH, Mohns EJ, Mehta B, Liu X, Chang B, Zenisek D, Crair MC, Demb JB, Chen B (2018) Restoration of vision after de novo genesis of rod photoreceptors in mammalian retinas. Nature 560(7719):484–488. https://doi.org/10.1038/s41586-018-0425-3
Hamon A, Roger JE, Yang X-J, Perron M (2016) Müller glial cell-dependent regeneration of the neural retina: an overview across vertebrate model systems. Dev Dyn 245(7):727–738. https://doi.org/10.1002/dvdy.24375
Wan J, Goldman D (2016) Retina regeneration in zebrafish. Curr Opin Genet Dev 40:41–47. https://doi.org/10.1016/j.gde.2016.05.009
Rajaram K, Harding RL, Bailey T, Patton JG, Hyde DR (2014) Dynamic miRNA expression patterns during retinal regeneration in zebrafish: reduced dicer or miRNA expression suppresses proliferation of Müller glia-derived neuronal progenitor cells. Dev Dyn 243(12):1591–1605. https://doi.org/10.1002/dvdy.24188
Rajaram K, Harding RL, Hyde DR, Patton JG (2014) miR-203 regulates progenitor cell proliferation during adult zebrafish retina regeneration. Dev Biol 392(2):393–403. https://doi.org/10.1016/j.ydbio.2014.05.005
Kaur S, Gupta S, Chaudhary M, Khursheed MA, Mitra S, Kurup AJ, Ramachandran R (2018) let-7 micro RNA-mediated regulation of Shh signaling and the gene regulatory network is essential for retina regeneration. Cell Rep 23(5):1409–1423. https://doi.org/10.1016/j.celrep.2018.04.002
Ramachandran R, Fausett BV, Goldman D (2010) Ascl1a regulates Müller glia dedifferentiation and retinal regeneration through a Lin-28-dependent, let-7 microRNA signalling pathway. Nat Cell Biol 12(11):1101–1107. https://doi.org/10.1038/ncb2115
Konar GJ, Ferguson C, Flickinger Z, Kent MR, Patton JG (2020) miRNAs and Müller Glia reprogramming during retina regeneration. Front Cell Dev Biol 8:632632. https://doi.org/10.3389/fcell.2020.632632
Plotnikova O, Baranova A, Skoblov M (2019) Comprehensive analysis of human microRNA-mRNA interactome. Front Genet 10:933. https://doi.org/10.3389/fgene.2019.00933
Nelson CM, Ackerman KM, O’Hayer P, Bailey TJ, Gorsuch RA, Hyde DR (2013) Tumor necrosis factor-alpha is produced by dying retinal neurons and is required for Muller glia proliferation during zebrafish retinal regeneration. J Neurosci 33(15):6524–6539. https://doi.org/10.1523/JNEUROSCI.3838-12.2013
Wan J, Zhao X-F, Vojtek A, Goldman D (2014) Retinal injury, growth factors, and cytokines converge on β-catenin and pStat3 signaling to stimulate retina regeneration. Cell Rep 9(1):285–297. https://doi.org/10.1016/j.celrep.2014.08.048
Zhao X-F, Wan J, Powell C, Ramachandran R, Myers MG, Goldman D (2014) Leptin and IL-6 family cytokines synergize to stimulate Müller glia reprogramming and retina regeneration. Cell Rep 9(1):272–284. https://doi.org/10.1016/j.celrep.2014.08.047
White DT, Sengupta S, Saxena MT, Xu Q, Hanes J, Ding D, Ji H, Mumm JS (2017) Immunomodulation-accelerated neuronal regeneration following selective rod photoreceptor cell ablation in the zebrafish retina. Proc Natl Acad Sci USA 114(18):E3719–E3728. https://doi.org/10.1073/pnas.1617721114
Silva NJ, Nagashima M, Li J, Kakuk-Atkins L, Ashrafzadeh M, Hyde DR, Hitchcock PF (2020) Inflammation and matrix metalloproteinase 9 (Mmp-9) regulate photoreceptor regeneration in adult zebrafish. Glia 68(7):1445–1465. https://doi.org/10.1002/glia.23792
Zhang Z, Hou H, Yu S, Zhou C, Zhang X, Li N, Zhang S, Song K, Lu Y, Liu D, Lu H, Xu H (2020) Inflammation-induced mammalian target of rapamycin signaling is essential for retina regeneration. Glia 68(1):111–127. https://doi.org/10.1002/glia.23707
Iribarne M (2021) Inflammation induces zebrafish regeneration. Neural Regen Res 16(9):1693–1701. https://doi.org/10.4103/1673-5374.306059
Nagashima M, Hitchcock PF (2021) Inflammation regulates the multi-step process of retinal regeneration in zebrafish. Cells 10(4):783. https://doi.org/10.3390/cells10040783
Todd L, Finkbeiner C, Wong CK, Hooper MJ, Reh TA (2020) Microglia suppress Ascl1-induced retinal regeneration in mice. Cell Rep 33(11):108507. https://doi.org/10.1016/j.celrep.2020.108507
Tahamtan A, Teymoori-Rad M, Nakstad B, Salimi V (2018) Anti-Inflammatory microRNAs and their potential for inflammatory diseases treatment. Front Immunol 9:1377. https://doi.org/10.3389/fimmu.2018.01377
Roy S, Benz F, Luedde T, Roderburg C (2015) The role of miRNAs in the regulation of inflammatory processes during hepatofibrogenesis. Hepatobiliary Surg Nutr 4(1):24–33. https://doi.org/10.3978/j.issn.2304-3881.2015.01.05
Huntzinger E, Izaurralde E (2011) Gene silencing by microRNAs: contributions of translational repression and mRNA decay. Nature Publishing Group:1–12 https://doi.org/10.1038/nrg2936
Jonas S, Izaurralde E (2015) Towards a molecular understanding of microRNA-mediated gene silencing. Nat Rev Genet 16(7):421–433. https://doi.org/10.1038/nrg3965
Taylor SM, Giuffre E, Moseley P, Hitchcock PF (2019) The microRNA, miR-18a, regulates NeuroD and photoreceptor differentiation in the retina of zebrafish. Dev Neurobiol 279(27):28418–28492. https://doi.org/10.1002/dneu.22666
Bernardos RL, Raymond PA (2006) GFAP transgenic zebrafish. Gene Expr Patterns 6(8):1007–1013. https://doi.org/10.1016/j.modgep.2006.04.006
Avdesh A, Chen M, Martin-Iverson MT, Mondal A, Ong D, Rainey-Smith S, Taddei K, Lardelli M, Groth DM, Verdile G, Martins RN (2012) Regular care and maintenance of a zebrafish (Danio rerio) laboratory: an introduction. J Vis Exp 69:e4196. https://doi.org/10.3791/4196
Taylor S, Chen J, Luo J, Hitchcock P (2012) Light-induced photoreceptor degeneration in the retina of the zebrafish. Methods Mol Biol 884:247–254. https://doi.org/10.1007/978-1-61779-848-1_17
Luo J, Uribe RA, Hayton S, Calinescu A-A, Gross JM, Hitchcock PF (2012) Midkine-A functions upstream of Id2a to regulate cell cycle kinetics in the developing vertebrate retina. Neural Dev 7(1):33. https://doi.org/10.1186/1749-8104-7-33
Barthel LK, Raymond PA (1993) Subcellular localization of α-tubulin and opsin mRNA in the goldfish retina using digoxigenin-labeled cRNA probes detected by alkaline phosphatase and HRP histochemistry. J Neurosci Methods 50(2):145–152. https://doi.org/10.1016/0165-0270(93)90002-9
Hitchcock P, Kakuk-Atkins L (2004) The basic helix-loop-helix transcription factor neuroD is expressed in the rod lineage of the teleost retina. J Comp Neurol 477(1):108–117. https://doi.org/10.1002/cne.20244
David R, Wedlich D (2001) PCR-based RNA probes: a quick and sensitive method to improve whole mount embryo in situ hybridizations. Biotech 30 (4):769-775. E5750AEF-427A-404F-B8B5-8B198FE51962
Friedman LM, Dror AA, Mor E, Tenne T, Toren G, Satoh T, Biesemeier DJ, Shomron N, Fekete DM, Hornstein E, Avraham KB (2009) MicroRNAs are essential for development and function of inner ear hair cells in vertebrates. Proc Natl Acad Sci USA 106(19):7915–7920. https://doi.org/10.1073/pnas.0812446106
Thummel R, Kassen SC, Montgomery JE, Enright JM, Hyde DR (2008) Inhibition of Müller glial cell division blocks regeneration of the light-damaged zebrafish retina. Dev Neurobiol 68(3):392–408. https://doi.org/10.1002/dneu.20596
Schneider CA, Rasband WS, Eliceiri KW (2012) NIH Image to ImageJ: 25 years of image analysis. Nat Meth 9(7):671–675. https://doi.org/10.1038/nmeth.2089
Nagashima M, Barthel LK, Raymond PA (2013) A self-renewing division of zebrafish Müller glial cells generates neuronal progenitors that require N-cadherin to regenerate retinal neurons. Development 140(22):4510–4521. https://doi.org/10.1242/dev.090738
Taylor SM, Alvarez-Delfin K, Saade CJ, Thomas JL, Thummel R, Fadool JM, Hitchcock PF (2015) The bHLH transcription factor NeuroD governs photoreceptor genesis and regeneration through delta-notch signaling. Invest Ophthalmol Vis Sci 56(12):7496–7515. https://doi.org/10.1167/iovs.15-17616
Martin JF, Poché RA (2019) Awakening the regenerative potential of the mammalian retina. Development 146 (23). https://doi.org/10.1242/dev.182642
Pesaresi M, Bonilla-Pons SA, Simonte G, Sanges D, Di Vicino U, Cosma MP (2018) Endogenous mobilization of bone-marrow cells into the murine retina induces fusion-mediated reprogramming of Müller glia cells. EBioMedicine 30:38–51. https://doi.org/10.1016/j.ebiom.2018.02.023
Sanges D, Romo N, Simonte G, Di Vicino U, Tahoces AD, Fernández E, Cosma MP (2013) Wnt/β-catenin signaling triggers neuron reprogramming and regeneration in the mouse retina. Cell Rep 4(2):271–286. https://doi.org/10.1016/j.celrep.2013.06.015
Sanges D, Simonte G, Di Vicino U, Romo N, Pinilla I, Nicolás M, Cosma MP (2016) Reprogramming Müller glia via in vivo cell fusion regenerates murine photoreceptors. J Clin Invest 126(8):3104–3116. https://doi.org/10.1172/JCI85193
Conner C, Ackerman KM, Lahne M, Hobgood JS, Hyde DR (2014) Repressing notch signaling and expressing TNFα are sufficient to mimic retinal regeneration by inducing Müller glial proliferation to generate committed progenitor cells. J Neurosci 34(43):14403–14419. https://doi.org/10.1523/JNEUROSCI.0498-14.2014
Mitchell DM, Lovel AG, Stenkamp DL (2018) Dynamic changes in microglial and macrophage characteristics during degeneration and regeneration of the zebrafish retina. J Neuroinflammation 15(1):120–163. https://doi.org/10.1186/s12974-018-1185-6
Liu L, Cai X, Liu E, Tian X, Tian C (2017) MicroRNA-18a promotes proliferation and metastasis in hepatocellular carcinoma via targeting KLF4. Oncotarget 8(40):68263–68269. https://doi.org/10.18632/oncotarget.19293
Shen K, Cao Z, Zhu R, You L, Zhang T (2019) The dual functional role of microRNA-18a (miR-18a) in cancer development. Clin Trans Med 8(1):1–13. https://doi.org/10.1186/s40169-019-0250-9
Jiang Y, Zhou J, Zhao J, Hou D, Zhang H, Li L, Zou D, Hu J, Zhang Y, Jing Z (2020) MiR-18a-downregulated RORA inhibits the proliferation and tumorigenesis of glioma using the TNF-α-mediated NF-κB signaling pathway. EBioMedicine 52:102651. https://doi.org/10.1016/j.ebiom.2020.102651
Bian S, Hong J, Li Q, Schebelle L, Pollock A, Knauss JL, Garg V, Sun T (2013) MicroRNA cluster miR-17-92 regulates neural stem cell expansion and transition to intermediate progenitors in the developing mouse neocortex. Cell Rep 3(5):1398–1406. https://doi.org/10.1016/j.celrep.2013.03.037
Liu C, Chen M, Wang M, Pi W, Li N, Meng Q (2018) MiR-18a regulates myoblasts proliferation by targeting Fgf1. PLoS ONE 13(7):e0201551. https://doi.org/10.1371/journal.pone.0201551
Humphreys KJ, McKinnon RA, Michael MZ (2014) miR-18a inhibits CDC42 and plays a tumour suppressor role in colorectal cancer cells. PLoS ONE 9(11):e112288. https://doi.org/10.1371/journal.pone.0112288
Zhang N, Zhang H, Liu Y, Su P, Zhang J, Wang X, Sun M, Chen B, Zhao W, Wang L, Wang H, Moran MS, Haffty BG, Yang Q (2019) SREBP1, targeted by miR-18a-5p, modulates epithelial-mesenchymal transition in breast cancer via forming a co-repressor complex with snail and HDAC1/2. Cell Death Differ 26(5):843–859. https://doi.org/10.1038/s41418-018-0158-8
D’Orazi FD, Suzuki SC, Darling N, Wong RO, Yoshimatsu T (2020) Conditional and biased regeneration of cone photoreceptor types in the zebrafish retina. J Comp Neurol 528(17):2816–2830. https://doi.org/10.1002/cne.24933
Ranski AH, Kramer AC, Morgan GW, Perez JL, Thummel R (2018) Characterization of retinal regeneration in adult zebrafish following multiple rounds of phototoxic lesion. PeerJ 6:e5646. https://doi.org/10.7717/peerj.5646
Powell C, Cornblath E, Elsaeidi F, Wan J, Goldman D (2016) Zebrafish Müller glia-derived progenitors are multipotent, exhibit proliferative biases and regenerate excess neurons. Sci Rep 6:24851. https://doi.org/10.1038/srep24851
Belmadani A, Tran PB, Ren D, Miller RJ (2006) Chemokines regulate the migration of neural progenitors to sites of neuroinflammation. J Neurosci 26(12):3182–3191. https://doi.org/10.1523/JNEUROSCI.0156-06.2006
Barkho BZ, Munoz AE, Li X, Li L, Cunningham LA, Zhao X (2008) Endogenous matrix metalloproteinase (MMP)-3 and MMP-9 promote the differentiation and migration of adult neural progenitor cells in response to chemokines. Stem Cells 26(12):3139–3149. https://doi.org/10.1634/stemcells.2008-0519
Xia X, Wang Y, Zheng JC (2020) The microRNA-17 ~ 92 family as a key regulator of neurogenesis and potential regenerative therapeutics of neurological disorders. Stem Cell Rev Rep. https://doi.org/10.1007/s12015-020-10050-5
Ochocinska MJ, Hitchcock PF (2009) NeuroD regulates proliferation of photoreceptor progenitors in the retina of the zebrafish. Mech Dev 126(3–4):128–141. https://doi.org/10.1016/j.mod.2008.11.009
Rutar M, Natoli R, Chia RX, Valter K, Provis JM (2015) Chemokine-mediated inflammation in the degenerating retina is coordinated by Müller cells, activated microglia, and retinal pigment epithelium. J Neuroinflammation 12:8. https://doi.org/10.1186/s12974-014-0224-1
Zhang S, Zhang S, Gong W, Zhu G, Wang S, Wang Y, Halim M, Wang K, Zhou G, Liu Q (2018) Müller cell regulated microglial activation and migration in rats with N-methyl-N-nitrosourea-induced retinal degeneration. Front Neurosci 12:890. https://doi.org/10.3389/fnins.2018.00890
Natoli R, Fernando N, Madigan M, Chu-Tan JA, Valter K, Provis J, Rutar M (2017) Microglia-derived IL-1β promotes chemokine expression by Müller cells and RPE in focal retinal degeneration. Mol Neurodegener 12(1):31. https://doi.org/10.1186/s13024-017-0175-y
Errede M, Girolamo F, Rizzi M, Bertossi M, Roncali L, Virgintino D (2014) The contribution of CXCL12-expressing radial glia cells to neuro-vascular patterning during human cerebral cortex development. Front Neurosci 8:324. https://doi.org/10.3389/fnins.2014.00324
Barzelay A, Weisthal Algor S, Niztan A, Katz S, Benhamou M, Nakdimon I, Azmon N, Gozlan S, Mezad-Koursh D, Neudorfer M, Goldstein M, Meilik B, Loewenstein A, Barak A (2018) Adipose-derived mesenchymal stem cells migrate and rescue RPE in the setting of oxidative stress. Stem Cells Int 2018:9682856. https://doi.org/10.1155/2018/9682856
Dokalis N, Prinz M (2019) Resolution of neuroinflammation: mechanisms and potential therapeutic option. Semin Immunopathol 41(6):699–709. https://doi.org/10.1007/s00281-019-00764-1
Fullerton JN, Gilroy DW (2016) Resolution of inflammation: a new therapeutic frontier. Nat Rev Drug Discov 15(8):551–567. https://doi.org/10.1038/nrd.2016.39
Mitchell DM, Sun C, Hunter SS, New DD, Stenkamp DL (2019) Regeneration associated transcriptional signature of retinal microglia and macrophages. Sci Rep 9(1):471768–4768. https://doi.org/10.1038/s41598-019-41298-8
Sifuentes CJ, Kim J-W, Swaroop A, Raymond PA (2016) Rapid, dynamic activation of Müller glial stem cell responses in zebrafish. Invest Ophthalmol Vis Sci 57(13):5148–5160. https://doi.org/10.1167/iovs.16-19973
Holtkamp GM, Kijlstra A, Peek R, de Vos AF (2001) Retinal pigment epithelium-immune system interactions: cytokine production and cytokine-induced changes. Prog Retin Eye Res 20(1):29–48. https://doi.org/10.1016/s1350-9462(00)00017-3
Kara N, Kent MR, Didiano D, Rajaram K, Zhao A, Summerbell ER, Patton JG (2019) The miR-216a-Dot1l regulatory axis is necessary and sufficient for Müller glia reprogramming during retina regeneration. Cell Rep 28(8):2037-2047.e2034. https://doi.org/10.1016/j.celrep.2019.07.061
Alam MM, O’Neill LA (2011) MicroRNAs and the resolution phase of inflammation in macrophages. Eur J Immunol 41(9):2482–2485. https://doi.org/10.1002/eji.201141740
Fulzele S, El-Sherbini A, Ahmad S, Sangani R, Matragoon S, El-Remessy A, Radhakrishnan R, Liou GI (2015) MicroRNA-146b-3p regulates retinal inflammation by suppressing adenosine deaminase-2 in diabetes. BioMed Res Int 2015. https://doi.org/10.1155/2015/846501
Fernando N, Wong JHC, Das S, Dietrich C, Aggio-Bruce R, Cioanca AV, Wooff Y, Chu-Tan JA, Schumann U, Ngo C, Essex RW, Dorian C, Robertson SA, Man SM, Provis J, Natoli R (2020) MicroRNA-223 regulates retinal function and inflammation in the healthy and degenerating retina. Front Cell Dev Biol. https://doi.org/10.3389/fcell.2020.00516
Aggio-Bruce R, Chu-Tan JA, Wooff Y, Cioanca AV, Schumann U, Natoli R (2021) Inhibition of microRNA-155 protects retinal function through attenuation of inflammation in retinal degeneration. Mol Neurobiol 58(2):835–854. https://doi.org/10.1007/s12035-020-02158-z
Kang S, Larbi D, Andrade M, Reardon S, Reh TA, Wohl SG (2020) A comparative analysis of reactive Müller glia gene expression after light damage and microRNA-depleted Müller glia-focus on microRNAs. Front Cell Dev Biol 8:620459. https://doi.org/10.3389/fcell.2020.620459
Rajman M, Schratt G (2017) MicroRNAs in neural development: from master regulators to fine-tuners. Development 144(13):2310–2322. https://doi.org/10.1242/dev.144337
Acknowledgements
The authors would like to thank James Hammond at UWF for facility support. We also thank Karen Gibbs, the RAE Office, and the Hal Marcus College of Science and Engineering for facilitating administrative, technical, and financial support.
Funding
This research was supported by the following grants: NIH 1R15EY031089-01 (SMT), NIH T32EY013934 (SMT), NIH F30EY031142 (ACK), NIH P30EY004068 (RT), NIH R21 EY031526 (RT), NIH P30EYO7003 (PFH), NIH R01EY07060 (PFH), and an unrestricted grant from the Research to Prevent Blindness, New York (RT and PFH).
Author information
Authors and Affiliations
Contributions
All authors contributed to the study conception and design. Material preparation, data collection and analysis were performed by Evin Magner, Pamela Sandoval-Sanchez, Ashley Kramer, Ryan Thummel, and Scott M. Taylor. The first draft of the manuscript was written by Evin Magner, Pamela Sandoval-Sanchez, and Scott M. Taylor, and all authors commented on previous versions of the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Ethics Approval
All experimental procedures were approved by the University of West Florida Institutional Animal Care and Use Committee.
Consent to Participate
N/A
Consent for Publication
N/A
Informed Consent
N/A
Competing Interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information

ESM 1
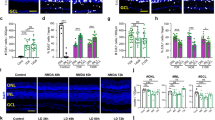
In situ hybridizations and quantification of mature cone and rod photoreceptors in WT and miR-18ami5012 retinas in uninjured retinas and at different numbers of days post-injury (dpi). (a-h) In situ hybridizations and (i, j) quantification for mature cones (arr3a) and rods (rho) in WT and miR-18ami5012 retinas in uninjured reinas (a, b, i, j) (cones—WT 123.8 ± 4.3 SD, miR-18ami5012 124.6 ± 3.4 SD cells/0.3 mm, p=0.81, n=3; rods—WT 291.7 ± 2.3 SD, miR-18ami5012 291.8 ± 13.8 SD cells/0.3 mm, p=0.99, n=3), at 7 dpi (c, d, i, j) (cones—WT 77.5 ± 4.5 SD, miR-18ami5012 32.3 ± 14.7 SD cells/0.3 mm, p=0.001, n=3; rods—WT 127.1 ± 72.1 SD, miR-18ami5012 98.5 ± 38.6 SD cells/0.3 mm, p=0.51, n=3), at 10 dpi (e, f, i, j) (cones—WT 107.6 ± 6.5 SD, miR-18ami5012 76.4 ± 2.5 SD cells/0.3 mm, p=0.001; rods—WT 275.6 ± 32.5 SD, miR-18ami5012 146.4 ± 46.7 SD cells/0.3 mm, p=0.017, n=3) and at 14 dpi (g-j) (cones—WT 120.3 ± 14.7 SD, miR-18ami5012 125.7 ± 11.4 SD cells/0.3 mm, p=0.462; rods—WT 261.3 ± 32.1 SD, miR-18ami5012 237.7 ± 28.8 SD cells/0.3 mm, p=0.241, n=3). Arrowheads show examples of labeled photoreceptors. Photoreceptor counts in retinal cross sections in the center of the lesioned area (cells per 0.3 mm of linear retina). Error bars represent standard deviation and asterisks indicate significant differences (Student’s t-test, p<0.05). Dotted lines on the graph connect the tops of the bars to show the trends. Abbreviations: RPE—retinal pigmented epithelium, ONL—outer nuclear layer, INL—inner nuclear layer; scale bar: 50 µm (PNG 5282 kb)

ESM 2
Efficiency of photolytic lesioning of photoreceptors in WT fish. (a, b) In situ hybridization for arr3a (cones) and rho (rods) in unlesioned retinas. (c, d) In situ hybridization for arr3a (cones) and rho (rods) in lesioned retinas at 3 dpi. (e, f) Quantification of mature cones (arr3a-labeled) and rods (rho-labeled) in retinal cross sections in unlesioned retinas and at 3 dpi (cones—unlesioned 88.4 ± 3.1 SD, 3 dpi 3.2 ± 5.0 SD cells/0.3 mm, p<0.0001, n=3; rods—unlesioned 291.7 ± 2.3 SD, 3 dpi 112.1 ± 76.5 SD cells/0.3 mm, p<0.015, n=3) . Cells were counted over 0.3 mm linear retina; error bars represent standard deviation and asterisks indicate significant differences (Student’s t-test, p<0.05); scale bar: 50 μm (PNG 6118 kb)
Rights and permissions
About this article
Cite this article
Magner, E., Sandoval-Sanchez, P., Kramer, A.C. et al. Disruption of miR-18a Alters Proliferation, Photoreceptor Replacement Kinetics, Inflammatory Signaling, and Microglia/Macrophage Numbers During Retinal Regeneration in Zebrafish. Mol Neurobiol 59, 2910–2931 (2022). https://doi.org/10.1007/s12035-022-02783-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12035-022-02783-w