Abstract
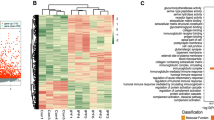
Idiopathic pulmonary fibrosis (IPF) carries a high mortality rate and has a poor prognosis. The pathogenesis of pulmonary fibrosis (PF) is highly related to dysregulation of multiple RNAs. This study aims to identify and validate dysregulated RNAs that exhibited dynamic alterations in response to bleomycin (BLM)-induced PF. The results will provide therapeutic targets for patients suffering from IPF. Whole transcriptomic profiles of BLM-induced PF were obtained through high-throughput RNA sequencing. miRNA profiling was downloaded from GSE45789 database in the Gene Expression Omnibus (GEO). We identified the differentially expressed RNAs (DERNAs) that exhibited dynamic alterations in response to BLM-induced PF. Subsequently, Gene Ontology (GO) and Kyoto Encyclopedia of Genes and Genome (KEGG) pathway enrichment analysis were conducted to discovery regulatory processes of PF. Weighted gene co-expression network analysis (WGCNA), protein–protein interaction (PPI) analysis, and co-expression analysis were performed to identify key genes and pathogenic pattern during the progression of PF. MiRanda, miRcode, and TargetScan were utilized to predict target relationships in the potential competing endogenous RNA (ceRNA) network. The results were verified by qRT-PCR analysis. In the context of BLM-induced PF, this study identified a total of 167 differentially expressed messenger RNAs (DEmRNAs), 115 differentially expressed long non-coding RNAs (DElncRNAs), 45 differentially expressed circular RNAs (DEcircRNAs), and 87 differentially expressed microRNAs (DEmiRNAs). These RNA molecules showed dynamic alterations in response to BLM-induced PF. These DEmRNAs exhibited a predominant association with the biological processes pertaining to the organization of extracellular matrix. A regulatory network was built in PF, encompassing 31 DEmRNAs, 18 DE lncRNAs, 13 DEcircRNAs, and 13 DEmiRNAs. Several DERNA molecules were subjected to validate using additional BLM-induced PF model. The outcomes of this validation process shown a strong correlation with the results obtained from RNA sequencing analysis. The GSE213001 dataset was utilized to validate the expression levels and diagnostic efficacy of four specific hub mRNAs (CCDC80, CLU, COL5A1, and COL6A3) in individuals diagnosed with PF. In this study, we identified and validated several RNA molecules that exhibited dynamic alternations in response to BLM-induced PF. These dysregulated RNAs participated in the pathogenesis of PF and can be used as therapeutic targets for early-stage IPF. Although more work must be done to confirm the results, our study may provide directions for future studies.









Similar content being viewed by others
Data Availability
NGS data are available from the NCBI Sequence Read Archive database (BioProject: PRJNA962191).
Abbreviations
- IPF:
-
Idiopathic pulmonary fibrosis
- ILD:
-
Interstitial lung disease
- GEO:
-
Gene Expression Omnibus
- DERNAs:
-
Differentially expressed RNAs
- CeRNA:
-
Competiting endogenous RNA
- GO:
-
Gene Ontology
- KEGG:
-
Kyoto Encyclopedia of Genes and Genome
- PPI:
-
Protein–protein interaction
- WGCNA:
-
Weighted gene co-expression network analysis
- ncRNAs:
-
Non-coding RNAs
- PCA:
-
Principal component analysis
- FMT:
-
Fibroblast-to-myofibroblast transition
- EMT:
-
Epithelial-mesenchymal transition
- BLM:
-
Bleomycin
- qRT-PCR:
-
Quantitative Real-Time PCR
- DEGs:
-
Differentially expressed genes
- ROC:
-
Receive operating characteristic
- AUCs:
-
The area under the curve
References
Raghu, G., Remy-Jardin, M., Richeldi, L., et al. (2022). Idiopathic pulmonary fibrosis (an Update) and progressive pulmonary fibrosis in adults: an official ATS/ERS/JRS/ALAT clinical practice guideline. American Journal of Respiratory and Critical Care Medicine, 205(9), e18–e47.
Spagnolo, P., Kropski, J. A., Jones, M. G., et al. (2021). Idiopathic pulmonary fibrosis: Disease mechanisms and drug development. Pharmacology & therpeutics, 222, 107798.
Thannickal, V. J., Toews, G. B., White, E. S., Lynch, J. P., 3rd., & Martinez, F. J. (2004). Mechanisms of pulmonary fibrosis. Annual Review of Medicine, 55, 395–417.
Ren, Z., Pan, X., Li, J., et al. (2023). G protein coupled receptor 41 regulates fibroblast activation in pulmonary fibrosis via Gα(i/o) and downstream Smad2/3 and ERK1/2 phosphorylation. Pharmacological Research, 191, 106754.
Morse, C., Tabib, T., Sembrat, J., et al. (2019). Proliferating SPP1/MERTK-expressing macrophages in idiopathic pulmonary fibrosis. The European Respiratory Journal, 54(2), 1802441.
Liu, S. S., Liu, C., Lv, X. X., et al. (2021). The chemokine CCL1 triggers an AMFR-SPRY1 pathway that promotes differentiation of lung fibroblasts into myofibroblasts and drives pulmonary fibrosis. Immunity, 54(9), 2042–56.e8.
Justet, A., Ghanem, M., Boghanim, T., et al. (2022). FGF19 is downregulated in idiopathic pulmonary fibrosis and inhibits lung fibrosis in mice. American Journal of Respiratory Cell and Molecular Biology, 67(2), 173–187.
Kraven, L. M., Taylor, A. R., Molyneaux, P. L., et al. (2023). Cluster analysis of transcriptomic datasets to identify endotypes of idiopathic pulmonary fibrosis. Thorax, 78(6), 551–558.
Liu, H., Wang, B., Zhang, J., et al. (2017). A novel lnc-PCF promotes the proliferation of TGF-β1-activated epithelial cells by targeting miR-344a-5p to regulate map3k11 in pulmonary fibrosis. Cell Death & Disease, 8(10), e3137.
Cui, H., Banerjee, S., Guo, S., et al. (2019). Long noncoding RNA Malat1 regulates differential activation of macrophages and response to lung injury. JCI Insight. https://doi.org/10.1172/jci.insight.124522
Xu, Q., Cheng, D., Li, G., et al. (2021). CircHIPK3 regulates pulmonary fibrosis by facilitating glycolysis in miR-30a-3p/FOXK2-dependent manner. International Journal of Biological Sciences, 17(9), 2294–2307.
Wu, Q., Liu, Y., Xie, Y., Wei, S., & Liu, Y. (2021). Identification of potential ceRNA network and patterns of immune cell infiltration in systemic Sclerosis-associated interstitial lung disease. Frontiers in Cell and Developmental Biology, 9, 622021.
Liu, Q., Chen, X., Kan, M., et al. (2021). Gypenoside XLIX loaded nanoparticles targeting therapy for renal fibrosis and its mechanism. European Journal of Pharmacology, 910, 174501.
Ashcroft, T., Simpson, J. M., & Timbrell, V. (1988). Simple method of estimating severity of pulmonary fibrosis on a numerical scale. Journal of Clinical Pathology, 41(4), 467–470.
Shi, F., Xu, H., Liu, C., et al. (2021). Whole-transcriptome sequencing reveals a vernalization-related ceRNA regulatory network in chinese cabbage (Brassica campestris L. ssp pekinensis). BMC Genomics, 22(1), 819.
Langfelder, P., & Horvath, S. (2008). WGCNA: An R package for weighted correlation network analysis. BMC Bioinformatics, 9, 559.
Niu, Y., Zhang, L., Qiu, H., et al. (2015). An improved method for detecting circulating microRNAs with S-Poly(T) Plus real-time PCR. Scientific Reports, 5, 15100.
Selman, M., & Pardo, A. (2002). Idiopathic pulmonary fibrosis: An epithelial/fibroblastic cross-talk disorder. Respiratory Research, 3(1), 3.
Hormuzdi, S. G., Strandjord, T. P., Madtes, D. K., & Bornstein, P. (1999). Mice with a targeted intronic deletion in the Col1a1 gene respond to bleomycin-induced pulmonary fibrosis with increased expression of the mutant allele. Matrix Biology : Journal of the International Society for Matrix Biology, 18(3), 287–294.
Kunugi, S., Fukuda, Y., Ishizaki, M., & Yamanaka, N. (2001). Role of MMP-2 in alveolar epithelial cell repair after bleomycin administration in rabbits. Laboratory investigation: A Journal of Technical Methods and Pathology, 81(9), 1309–1318.
Hung, C. F., Rohani, M. G., Lee, S. S., Chen, P., & Schnapp, L. M. (2013). Role of IGF-1 pathway in lung fibroblast activation. Respiratory Research, 14(1), 102.
Dong, Y., Geng, Y., Li, L., et al. (2015). Blocking follistatin-like 1 attenuates bleomycin-induced pulmonary fibrosis in mice. The Journal of Experimental Medicine, 212(2), 235–252.
Son, E. S., Fei, X., Yoon, J. H., et al. (2021). Comparison of Pharmacokinetics and Anti-Pulmonary Fibrosis-Related Effects of Sulforaphane and Sulforaphane N-acetylcysteine. Pharmaceutics, 13(7), 958.
Parra, E. R., Teodoro, W. R., de Morais, J., et al. (2010). Increased mRNA expression of collagen V gene in pulmonary fibrosis of systemic sclerosis. European Journal of Clinical Investigation, 40(2), 110–120.
Jessen, H., Hoyer, N., Prior, T. S., et al. (2021). Turnover of type I and III collagen predicts progression of idiopathic pulmonary fibrosis. Respiratory Research, 22(1), 205.
Hamaguchi, Y., Fujimoto, M., Matsushita, T., Hasegawa, M., Takehara, K., & Sato, S. (2008). Elevated serum insulin-like growth factor (IGF-1) and IGF binding protein-3 levels in patients with systemic sclerosis: Possible role in development of fibrosis. The Journal of Rheumatology, 35(12), 2363–2371.
Snyder-Talkington, B. N., Dong, C., Sargent, L. M., et al. (2016). mRNAs and miRNAs in whole blood associated with lung hyperplasia, fibrosis, and bronchiolo-alveolar adenoma and adenocarcinoma after multi-walled carbon nanotube inhalation exposure in mice. Journal of Applied Toxicology : JAT, 36(1), 161–174.
Makiguchi, T., Yamada, M., Yoshioka, Y., et al. (2016). Serum extracellular vesicular miR-21-5p is a predictor of the prognosis in idiopathic pulmonary fibrosis. Respiratory Research, 17(1), 110.
Jia, R., Li, T., & Wang, N. (2021). Long noncoding RNA HOTAIR functions as ceRNA to regulate MMP2 in paraquat induced lung epithelial-mesenchymal transition. Toxicology, 461, 152891.
Jeon, S., Jin, H., Kim, J. M., et al. (2023). The miR-15b-Smurf2-HSP27 axis promotes pulmonary fibrosis. Journal of Biomedical Science, 30(1), 2.
Lei, X., He, N., Zhu, L., et al. (2021). Mesenchymal stem cell-derived extracellular vesicles attenuate radiation-induced lung injury via miRNA-214-3p. Antioxidants & Redox Signaling, 35(11), 849–862.
Salmena, L., Poliseno, L., Tay, Y., Kats, L., & Pandolfi pp. (2011). A ceRNA hypothesis: the Rosetta Stone of a hidden RNA language? Cell, 146(3), 353–358.
Fatica, A., & Bozzoni, I. (2014). Long non-coding RNAs: New players in cell differentiation and development. Nature Reviews Genetics, 15(1), 7–21.
Tay, Y., Rinn, J., & Pandolfi pp. (2014). The multilayered complexity of ceRNA crosstalk and competition. Nature, 505(7483), 344–352.
Tay, Y., Kats, L., Salmena, L., et al. (2011). Coding-independent regulation of the tumor suppressor PTEN by competing endogenous mRNAs. Cell, 147(2), 344–357.
Taulli, R., Loretelli, C., & Pandolfi pp. (2013). From pseudo-ceRNAs to circ-ceRNAs: a tale of cross-talk and competition. Nature Structural & Molecular Biology, 20(5), 541–543.
Guil, S., & Esteller, M. (2015). RNA-RNA interactions in gene regulation: The coding and noncoding players. Trends in Biochemical Sciences, 40(5), 248–256.
Liu, Y., Lu, F. A., Wang, L., Wang, Y. F., & Wu, C. F. (2021). Long non-coding RNA NEAT1 promotes pulmonary fibrosis by regulating the microRNA-455-3p/SMAD3 axis. Molecular Medicine Reports. https://doi.org/10.3892/mmr.2021.11857
Thomson, D. W., & Dinger, M. E. (2016). Endogenous microRNA sponges: Evidence and controversy. Nature Reviews Genetics, 17(5), 272–283.
Denzler, R., Agarwal, V., Stefano, J., Bartel, D. P., & Stoffel, M. (2014). Assessing the ceRNA hypothesis with quantitative measurements of miRNA and target abundance. Molecular cell, 54(5), 766–776.
Li, J., Li, P., Zhang, G., Qin, P., Zhang, D., & Zhao, W. (2020). CircRNA TADA2A relieves idiopathic pulmonary fibrosis by inhibiting proliferation and activation of fibroblasts. Cell Death & Disease, 11(7), 553.
Fang, S., Guo, H., Cheng, Y., et al. (2018). circHECTD1 promotes the silica-induced pulmonary endothelial-mesenchymal transition via HECTD1. Cell Death & Disease, 9(3), 396.
Lin, W., Song, Y., Li, T., et al. (2023). Triptolide attenuates pulmonary fibrosis by inhibiting fibrotic extracellular matrix remodeling mediated by MMPs/LOX/integrin. Biomedicine & Pharmacotherapy Biomedecine & Pharmacotherapie, 166, 115394.
Zheng, S., Zhang, Y., Hou, Y., et al. (2023). Underlying Molecular Mechanism and Construction of a miRNA-Gene Network in Idiopathic Pulmonary Fibrosis by Bioinformatics. International Journal of Molecular Sciences, 24(17), 13305.
Zou, M., Zou, J., Hu, X., Zheng, W., Zhang, M., & Cheng, Z. (2021). Latent transforming growth Factor-β binding protein-2 regulates lung fibroblast-to-Myofibroblast differentiation in pulmonary fibrosis via NF-κB Signaling. Frontiers in Pharmacology, 12, 788714.
Jin, Y. K., Li, X. H., Wang, W., et al. (2018). Follistatin-like 1 promotes Bleomycin-induced pulmonary fibrosis through the transforming growth factor Beta 1/mitogen-activated protein kinase signaling pathway. Chinese Medical Journal, 131(16), 1917–1925.
Acknowledgements
Not applicable.
Funding
This work was supported by The National Natural Science Foundation of China (89202586), Joint Application Foundation Project of Kunming Medical University and Yunnan Provincial Science and Technology Department (202101AY070001-271 and 202001AY070001-289), and Open Project of Yunnan Provincial Clinical Medical Center for Respiratory System Diseases (2020LCZXKF-HX08, 2022LCZXKF-HX04).
Author information
Authors and Affiliations
Contributions
YHZ and DMG: designed the study. DYL and JZ: constructed the BLM-induced PF mice model. JW, SJL, LQ, GHF and KW: performed comprehensive bioinformatics analysis. DYL, DXS, and ZMH: performed the qRT-PCR validation. DYL: wrote the article. JW, DMG and YHZ: revised the article. All authors approved the final version of the manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no competing interests.
Ethical approval
All procedures comply with National Institutes of Health guidelines (NIH Publication, 8th edition, 2011), and were approved by the Institutional Animal Care and Use Committee (IACUC) of The First People’s Hospital of Yunnan Province.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Li, D., Wang, J., Zeng, J. et al. Identification and Validation of Genes Exhibiting Dynamic Alterations in Response to Bleomycin-Induced Pulmonary Fibrosis. Mol Biotechnol (2023). https://doi.org/10.1007/s12033-023-00943-4
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s12033-023-00943-4