Abstract
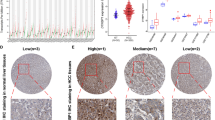
Hepatocellular carcinoma (HCC) is one of the leading causes of cancer death worldwide. Consequently, it is essential to identify biomarkers for treatment response and the prognosis prediction. We investigated whether ABL1 can function as a biomarker or a drug target for HCC. We assessed the ABL1 expression, genetic alterations and patients’ survival from LinkedOmics, GEO, TCGA and Human Protein Atlas. We analyzed PPI, GO and KEGG pathways. GSEA was analyzed for functional comparison. The current drugs targeting ABL1 were statistically analyzed using DRUGSURV and DGIdb database. We found ABL1 is overexpressed in HCC and its higher expression reduces survival probability. Genetic changes of ABL1 are not frequent. We screened out 25 differentially expressed genes correlated with ABL1. The top functions of ABL1 are biological regulation, metabolic process, protein-containing, and protein binding. KEGG pathways showed that ABL1 and correlated with ABL1 significantly genes markedly enriched in the ErbB signaling pathway, and pathways in cancer. We counted the existing drugs targeting ABL1, which indicates that inhibiting ABL1 expression may improve the survival probability of HCC. In conclusion, ABL1 plays a crucial role in the development and progression of this cancerization and is a potential drug target.










Similar content being viewed by others
Data availability
All data generated or analyzed during this study are included in this published article files.
Abbreviations
- ABL1:
-
ABL proto-oncogene 1 non-receptor tyrosine kinase
- EMT:
-
Epithelial-mesenchymal transformation
- GEO:
-
Gene expression omnibus
- GO:
-
Gene ontology
- GSEA:
-
Gene set enrichment analysis
- HCC:
-
Hepatocellular carcinoma
- HPA:
-
Human protein atlas
- KEGG:
-
Kyoto encyclopedia of genes and genomes
- PPI:
-
Protein–protein interaction
- TCGA:
-
The cancer genome atlas
References
Farazi PA, DePinho RA. Hepatocellular carcinoma pathogenesis: from genes to environment. Nat Rev Cancer. 2006;6(9):674–87. https://doi.org/10.1038/nrc1934].
Greten TF, Lai CW, Li G, Staveley-O’Carroll KF. Targeted and immune-based therapies for hepatocellular carcinoma. Gastroenterology. 2019;156(2):510–24. https://doi.org/10.1053/j.gastro.2018.09.051].
Bruix J, Gores GJ, Mazzaferro V. Hepatocellular carcinoma: clinical frontiers and perspectives. Gut. 2014;63(5):844–55. https://doi.org/10.1136/gutjnl-2013-306627].
Zhu X, Chen MS, Tian LW, Li DP, Xu PL, Lin MC, Xie D, Kung HF. Single nucleotide polymorphism of rs430397 in the fifth intron of GRP78 gene and clinical relevance of primary hepatocellular carcinoma in Han Chinese: risk and prognosis. Int J Cancer. 2009;125(6):1352–7. https://doi.org/10.1002/ijc.24487].
Vilchez V, Turcios L, Marti F, Gedaly R. Targeting Wnt/beta-catenin pathway in hepatocellular carcinoma treatment. World J Gastroenterol. 2016;22(2):823–32. https://doi.org/10.3748/wjg.v22.i2.823].
Zou Z, Tao T, Li H, Zhu X. mTOR signaling pathway and mTOR inhibitors in cancer: progress and challenges. Cell Biosci. 2020;10:31. https://doi.org/10.1186/s13578-020-00396-1].
Greuber EK, Smith-Pearson P, Wang J, Pendergast AM. Role of ABL family kinases in cancer: from leukaemia to solid tumours. Nat Rev Cancer. 2013;13(8):559–71. https://doi.org/10.1038/nrc3563].
Colicelli J. ABL tyrosine kinases: evolution of function, regulation, and specificity. Sci Signal. 2010;3(139):re6. https://doi.org/10.1126/scisignal.3139re6].
Abelson HT, Rabstein LS. Lymphosarcoma: virus-induced thymic-independent disease in mice. Cancer Res. 1970;30(8):2213–22.
Wang F, Hou W, Chitsike L, Xu Y, Bettler C, Perera A, Bank T, Cotler SJ, Dhanarajan A, Denning MF, Ding X, Breslin P, Qiang W, Li J, Koleske AJ, Qiu W. ABL1, overexpressed in hepatocellular carcinomas, regulates expression of NOTCH1 and promotes development of liver tumors in mice. Gastroenterology. 2020;159(1):289–305. https://doi.org/10.1053/j.gastro.2020.03.013].
Chitsike L, Ding X, Breslin P, Qiu W. ABL1 is overexpressed and activated in hepatocellular carcinoma. J Cancer Tumor Int. 2017;6(3):1–8. https://doi.org/10.9734/jcti/2017/37131.
Van Etten RA. Cycling, stressed-out and nervous: cellular functions of c-Abl. Trends Cell Biol. 1999;9(5):179–86. https://doi.org/10.1016/s0962-8924(99)01549-4].
Yuan X, Wu H, Xu H, Xiong H, Chu Q, Yu S, Wu GS, Wu K. Notch signaling: an emerging therapeutic target for cancer treatment. Cancer Lett. 2015;369(1):20–7. https://doi.org/10.1016/j.canlet.2015.07.048].
Rhodes DR, Yu J, Shanker K, Deshpande N, Varambally R, Ghosh D, Barrette T, Pandey A, Chinnaiyan AM. ONCOMINE: a cancer microarray database and integrated data-mining platform. Neoplasia. 2004;6(1):1–6. https://doi.org/10.1016/s1476-5586(04)80047-2].
Rhodes DR, Kalyana-Sundaram S, Mahavisno V, Varambally R, Yu J, Briggs BB, Barrette TR, Anstet MJ, Kincead-Beal C, Kulkarni P, Varambally S, Ghosh D, Chinnaiyan AM. Oncomine 3.0: genes, pathways, and networks in a collection of 18,000 cancer gene expression profiles. Neoplasia. 2007;9(2):166–80. https://doi.org/10.1593/neo.07112].
Chen X, Cheung ST, So S, Fan ST, Barry C, Higgins J, Lai KM, Ji J, Dudoit S, Ng IO, Van De Rijn M, Botstein D, Brown PO. Gene expression patterns in human liver cancers. Mol Biol Cell. 2002;13(6):1929–39. https://doi.org/10.1091/mbc.02-02-0023].
Roessler S, Jia HL, Budhu A, Forgues M, Ye QH, Lee JS, Thorgeirsson SS, Sun Z, Tang ZY, Qin LX, Wang XW. A unique metastasis gene signature enables prediction of tumor relapse in early-stage hepatocellular carcinoma patients. Cancer Res. 2010;70(24):10202–12. https://doi.org/10.1158/0008-5472.CAN-10-2607].
Wurmbach E, Chen YB, Khitrov G, Zhang W, Roayaie S, Schwartz M, Fiel I, Thung S, Mazzaferro V, Bruix J, Bottinger E, Friedman S, Waxman S, Llovet JM. Genome-wide molecular profiles of HCV-induced dysplasia and hepatocellular carcinoma. Hepatology. 2007;45(4):938–47. https://doi.org/10.1002/hep.21622].
Mas VR, Maluf DG, Archer KJ, Yanek K, Kong X, Kulik L, Freise CE, Olthoff KM, Ghobrial RM, McIver P, Fisher R. Genes involved in viral carcinogenesis and tumor initiation in hepatitis C virus-induced hepatocellular carcinoma. Mol Med. 2009;15(3–4):85–94. https://doi.org/10.2119/molmed.2008.00110].
Guichard C, Amaddeo G, Imbeaud S, Ladeiro Y, Pelletier L, Maad IB, Calderaro J, Bioulac-Sage P, Letexier M, Degos F, Clement B, Balabaud C, Chevet E, Laurent A, Couchy G, Letouze E, Calvo F, Zucman-Rossi J. Integrated analysis of somatic mutations and focal copy-number changes identifies key genes and pathways in hepatocellular carcinoma. Nat Genet. 2012;44(6):694–8. https://doi.org/10.1038/ng.2256].
Chandrashekar DS, Bashel B, Balasubramanya SAH, Creighton CJ, Ponce-Rodriguez I, Chakravarthi B, Varambally S. UALCAN: a portal for facilitating tumor subgroup gene expression and survival analyses. Neoplasia. 2017;19(8):649–58. https://doi.org/10.1016/j.neo.2017.05.002].
Thul PJ, Lindskog C. The human protein atlas: a spatial map of the human proteome. Protein Sci. 2018;27(1):233–44. https://doi.org/10.1002/pro.3307].
Gao J, Aksoy BA, Dogrusoz U, Dresdner G, Gross B, Sumer SO, Sun Y, Jacobsen A, Sinha R, Larsson E, Cerami E, Sander C, Schultz N. Integrative analysis of complex cancer genomics and clinical profiles using the cBioPortal. Sci Signal. 2013. https://doi.org/10.1126/scisignal.2004088].
Szklarczyk D, Morris JH, Cook H, Kuhn M, Wyder S, Simonovic M, Santos A, Doncheva NT, Roth A, Bork P, Jensen LJ, von Mering C. The STRING database in 2017: quality-controlled protein-protein association networks, made broadly accessible. Nucleic Acids Res. 2017;45(D1):D362–8. https://doi.org/10.1093/nar/gkw937].
Vasaikar SV, Straub P, Wang J, Zhang B. LinkedOmics: analyzing multi-omics data within and across 32 cancer types. Nucleic Acids Res. 2018;46(D1):D956–63. https://doi.org/10.1093/nar/gkx1090].
Wang J, Vasaikar S, Shi Z, Greer M, Zhang B. WebGestalt 2017: a more comprehensive, powerful, flexible and interactive gene set enrichment analysis toolkit. Nucleic Acids Res. 2017;45(W1):W130–7. https://doi.org/10.1093/nar/gkx356].
Liao Y, Wang J, Jaehnig EJ, Shi Z, Zhang B. WebGestalt 2019: gene set analysis toolkit with revamped UIs and APIs. Nucleic Acids Res. 2019;47(W1):W199–205. https://doi.org/10.1093/nar/gkz401].
Xie C, Mao X, Huang J, Ding Y, Wu J, Dong S, Kong L, Gao G, Li CY, Wei L. KOBAS 2.0: a web server for annotation and identification of enriched pathways and diseases. Nucleic Acids Res. 2011. https://doi.org/10.1093/nar/gkr483].
Wu J, Mao X, Cai T, Luo J, Wei L. KOBAS server: a web-based platform for automated annotation and pathway identification. Nucleic Acids Res. 2006. https://doi.org/10.1093/nar/gkl167].
Amelio I, Gostev M, Knight RA, Willis AE, Melino G, Antonov AV. DRUGSURV: a resource for repositioning of approved and experimental drugs in oncology based on patient survival information. Cell Death Dis. 2014;5: e1051. https://doi.org/10.1038/cddis.2014.9].
Li R, Knight JF, Park M, Pendergast AM. Abl kinases regulate HGF/Met signaling required for epithelial cell scattering, tubulogenesis and motility. PLoS ONE. 2015;10(5): e0124960. https://doi.org/10.1371/journal.pone.0124960].
Smith-Pearson PS, Greuber EK, Yogalingam G, Pendergast AM. Abl kinases are required for invadopodia formation and chemokine-induced invasion. J Biol Chem. 2010;285(51):40201–11. https://doi.org/10.1074/jbc.M110.147330].
Bradley WD, Koleske AJ. Regulation of cell migration and morphogenesis by Abl-family kinases: emerging mechanisms and physiological contexts. J Cell Sci. 2009;122(Pt 19):3441–54. https://doi.org/10.1242/jcs.039859].
Feigin ME, Muthuswamy SK. Polarity proteins regulate mammalian cell-cell junctions and cancer pathogenesis. Curr Opin Cell Biol. 2009;21(5):694–700. https://doi.org/10.1016/j.ceb.2009.07.003].
Lee M, Vasioukhin V. Cell polarity and cancer–cell and tissue polarity as a non-canonical tumor suppressor. J Cell Sci. 2008;121(Pt 8):1141–50. https://doi.org/10.1242/jcs.016634].
Wielenga VJ, van der Voort R, Taher TE, Smit L, Beuling EA, van Krimpen C, Spaargaren M, Pals ST. Expression of c-Met and heparan-sulfate proteoglycan forms of CD44 in colorectal cancer. Am J Pathol. 2000;157(5):1563–73. https://doi.org/10.1016/S0002-9440(10)64793-1].
Srinivasan D, Plattner R. Activation of Abl tyrosine kinases promotes invasion of aggressive breast cancer cells. Cancer Res. 2006;66(11):5648–55. https://doi.org/10.1158/0008-5472.CAN-06-0734].
Sourbier C, Ricketts CJ, Matsumoto S, Crooks DR, Liao PJ, Mannes PZ, Yang Y, Wei MH, Srivastava G, Ghosh S, Chen V, Vocke CD, Merino M, Srinivasan R, Krishna MC, Mitchell JB, Pendergast AM, Rouault TA, Neckers L, Linehan WM. Targeting ABL1-mediated oxidative stress adaptation in fumarate hydratase-deficient cancer. Cancer Cell. 2014;26(6):840–50. https://doi.org/10.1016/j.ccell.2014.10.005].
Ganguly SS, Fiore LS, Sims JT, Friend JW, Srinivasan D, Thacker MA, Cibull ML, Wang C, Novak M, Kaetzel DM, Plattner R. c-Abl and Arg are activated in human primary melanomas, promote melanoma cell invasion via distinct pathways, and drive metastatic progression. Oncogene. 2012;31(14):1804–16. https://doi.org/10.1038/onc.2011.361].
Heimbach JK, Kulik LM, Finn RS, Sirlin CB, Abecassis MM, Roberts LR, Zhu AX, Murad MH, Marrero JA. AASLD guidelines for the treatment of hepatocellular carcinoma. Hepatology. 2018;67(1):358–80. https://doi.org/10.1002/hep.29086].
Regad T, Targeting RTK. Signaling pathways in cancer. Cancers. 2015;7(3):1758–84. https://doi.org/10.3390/cancers7030860].
Al-Zoughbi W, Pichler M, Gorkiewicz G, Guertl-Lackner B, Haybaeck J, Jahn SW, Lackner C, Liegl-Atzwanger B, Popper H, Schauer S, Nusshold E, Kindt AS, Trajanoski Z, Speicher MR, Haemmerle G, Zimmermann R, Zechner R, Vesely PW, Hoefler G. Loss of adipose triglyceride lipase is associated with human cancer and induces mouse pulmonary neoplasia. Oncotarget. 2016;7(23):33832–40. https://doi.org/10.18632/oncotarget.9418].
Rani A, Murphy JJ. STAT5 in cancer and immunity. J Interferon Cytokine Res. 2016;36(4):226–37. https://doi.org/10.1089/jir.2015.0054].
Song G, Ouyang G, Bao S. The activation of Akt/PKB signaling pathway and cell survival. J Cell Mol Med. 2005;9(1):59–71. https://doi.org/10.1111/j.1582-4934.2005.tb00337.x].
Maik-Rachline G, Hacohen-Lev-Ran A, Seger R. Nuclear ERK: mechanism of translocation, substrates, and role in cancer. Int J Mol Sci. 2019. https://doi.org/10.3390/ijms20051194].
Funding
This work was supported partly by Guangzhou Science and Technology Project (202102080262); Guangdong Medical Science and Technology Research Fund (B2021205).
Author information
Authors and Affiliations
Contributions
XZ designed this study and supervised the research. FL, XC, LL and XZ conducted the main experiments, analyze data, and wrote the manuscript. ZX, ZC and XZ edited and discussed the manuscript. YZ, XZ, YY, YS and ML assisted the data analysis, edited and discussed the manuscript. ZH designed, supervised, edited and discussed the new version. XZ checked the statistical and bioinformatic accuracy as an expert in statistics and bioinformatics. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical approval and consent to participate
The work was approved by Guangdong Medical University Ethics committee. Informed consent forms are not required for patient data extracted from public databases.
Consent for publication
N/a.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Lan, F., Chen, X., Xiong, Z. et al. Comprehensive transcriptomic and co-expression analysis of ABL1 gene and molecularly targeted drugs in hepatocellular carcinoma based on multi-database mining. Med Oncol 39, 146 (2022). https://doi.org/10.1007/s12032-022-01730-y
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s12032-022-01730-y