Abstract
Background
Hepatocellular carcinoma (HCC) is one of the most challenging malignancies, with high morbidity and mortality rates. The transforming growth factor-β (TGF-β) pathway plays a dual role in HCC, acting as both tumor suppressor and promoter. A thorough understanding of the mechanisms underlying its opposing functions is important. The growth suppressive effects of TGF-β remain largely unknown for mesenchymal HCC cells. Using a systematic approach, here we assess the cytostatic TGF-β responses and intracellular transduction of the canonical TGF-β/Smad signaling cascade in mesenchymal-like HCC cell lines.
Methods
Nine mesenchymal-like HCC cell lines, including SNU182, SNU387, SNU398, SNU423, SNU449, SNU475, Mahlavu, Focus, and Sk-Hep1, were used in this study. The cytostatic effects of TGF-β were evaluated by cell cycle analysis, BrdU labeling, and SA-β-Gal assay. RT-PCR and western blot analysis were utilized to determine the mRNA and protein expression levels of TGF-β signaling components and cytostatic genes. Immunoperoxidase staining and luciferase reporter assays were performed to comprehend the transduction of the canonical TGF-β pathway.
Results
We report that mesenchymal-like HCC cell lines are resistant to TGF-β-induced growth suppression. The vast majority of cell lines have an active canonical signaling from the cell membrane to the nucleus. Three cell lines had lost the expression of cytostatic effector genes.
Conclusion
Our findings reveal that cytostatic TGF-β responses have been selectively lost in mesenchymal-like HCC cell lines. Notably, their lack of responsiveness was not associated with a widespread impairment of TGF-β signaling cascade. These cell lines may serve as valuable models for studying the molecular mechanisms underlying the loss of TGF-β-mediated cytostasis during hepatocarcinogenesis.






Similar content being viewed by others
References
Globocan. “WHO Liver Cancer Fact Sheet”. 2020.
Llovet JM et al. “Hepatocellular carcinoma,” Nat Rev Dis Primer. 2021;7(1). https://doi.org/10.1038/s41572-020-00240-3.
Villanueva A. “Hepatocellular carcinoma,” N Engl J Med. 2019;1450–1462. https://doi.org/10.1056/NEJMra1713263.
Llovet JM, Zucman-rossi J, Pikarsky E, Sangro B, Sherman M, Gores G. “Hepatocellular carcinoma,” Nat. Rev. Dis. Primer. 2016;2. https://doi.org/10.1038/nrdp.2016.18.
Zucman-rossi J, Villanueva A, Nault J, Llovet JM. Genetic landscape and biomarkers of hepatocellular carcinoma. YGAST. 2015;149(5):1226-1239.e4. https://doi.org/10.1053/j.gastro.2015.05.061.
Ozturk M, Arslan-Ergul A, Bagislar S, Senturk S, Yuzugullu H. Senescence and immortality in hepatocellular carcinoma. Cancer Lett. 2009;286(1):103–13. https://doi.org/10.1016/j.canlet.2008.10.048.
Yuzugullu H et al. Canonical Wnt signaling is antagonized by noncanonical Wnt5a in hepatocellular carcinoma cells. Mol Cancer. 2009;8:1–20. https://doi.org/10.1186/1476-4598-8-90.
Caruso S et al. “Analysis of liver cancer cell lines identifies agents with likely efficacy against hepatocellular carcinoma and markers of response,” Gastroenterology. 2019;1–17. https://doi.org/10.1053/j.gastro.2019.05.001.
Dituri F, Mancarella S, Cigliano A, Chieti A, Giannelli G. TGF-β as multifaceted orchestrator in HCC progression: signaling, EMT, immune microenvironment, and novel therapeutic perspectives. Semin Liver Dis. 2019;39(1):53–69. https://doi.org/10.1055/s-0038-1676121.
Weiss A, Attisano L. The TGFbeta superfamily signaling pathway. WIREs Dev Biol. 2013;2:47–63. https://doi.org/10.1002/wdev.86.
Derynck R, Zhang YE. Smad-dependent and Smad-independent pathways in TGF-β family signalling. Nature. 2003;425:577–84.
Zhang YE “Non-Smad signaling pathways of the TGF- b family,” Cold Spring Harb Perspect Biol. 2017;1–18.
Massagué J, Seoane J, Wotton D. Smad transcription factors. Genes Dev. 2005;19(23):2783–810. https://doi.org/10.1101/gad.1350705.
Arrese M, Cabrera D, Hernandez A, Astete L, Estrada L. “TGF- β and hepatocellular carcinoma : when a friend becomes an enemy,” Curr Protein Pept Sci. 2018;19. https://doi.org/10.2174/1389203718666171117112619.
Seoane J, Gomis RR. “TGF- b family signaling in tumor suppression and cancer progression,” Cold Spring Harb Perspect Biol. 2017;9. https://doi.org/10.1101/cshperspect.a022277.
Dzieran J et al. Comparative analysis of TGF-β / Smad signaling dependent cytostasis in human hepatocellular carcinoma cell lines. PLoS One. 2013;8(8):1–18. https://doi.org/10.1371/journal.pone.0072252.
Hao Y, Baker D, Ten Dijke P. “TGF-β-mediated epithelial-mesenchymal transition and cancer metastasis,” Int J Mol Sci. 2019;20(11). https://doi.org/10.3390/ijms20112767.
Fabregat I, Caballero-Díaz D. “Transforming growth factor-β-induced cell plasticity in liver fibrosis and hepatocarcinogenesis,” Front Oncol. 2018;8. https://doi.org/10.3389/fonc.2018.00357.
Senturk S, Mumcuoglu M, Gursoy-Yuzugullu O, Cingoz B, Akcali KC, Ozturk M. Transforming growth factor-beta induces senescence in hepatocellular carcinoma cells and inhibits tumor growth. Hepatology. 2010;52(3):966–74. https://doi.org/10.1002/hep.23769.
Coulouarn C, Factor VM, Thorgeirsson SS. Transforming growth factor-β gene expression signature in mouse hepatocytes predicts clinical outcome in human cancer. Hepatology. 2008;47(6):2059–67. https://doi.org/10.1038/jid.2014.371.
Huh N, Utakoji T. Production of HBs-antigen by two new human hepatoma cell lines and its enhancement by dexamethasone. Gan. 1981;72(1):178–9.
Park JG et al. Characterization of cell lines established from human hepatocellular carcinoma. Int J Cancer. 1995;62(3):276–82. https://doi.org/10.1002/ijc.2910620308.
Hirschfield H et al. In vitro modeling of hepatocellular carcinoma molecular subtypes for anti-cancer drug assessment. Exp Mol Med. 2018;50(1): e419. https://doi.org/10.1038/emm.2017.164.
Heffelfinger SC, Hawkins HH, Barrish J, Taylor L, Darlington GJ. “SK HEP-1: a human cell line of endothelial origin”, Vitro Cell. Dev Biol J Tissue Cult Assoc. 1992;28A(2):136–42. https://doi.org/10.1007/BF02631017.
Alexander JJ. “Human hepatoma cell lines,” in Neoplasms of the liver, K. Okuda and K. G. Ishak, Eds. Tokyo: Springer Japan. 1987;47–56. https://doi.org/10.1007/978-4-431-68349-0_4.
He L, Isselbacher KJ, Wands JR, Goodman HM, Shih C, Quaroni A. Establishment and characterization of a new human hepatocellular carcinoma cell line. Vitro. 1984;20(6):493–504. https://doi.org/10.1007/BF02619623.
Inagaki M, Moustakas A, Lin HY, Lodish HF, Carr BI. Growth inhibition by transforming growth factor beta (TGF-beta) type I is restored in TGF-beta-resistant hepatoma cells after expression of TGF-beta receptor type II cDNA. Proc Natl Acad Sci USA. 1993;90(11):5359–63. https://doi.org/10.1073/pnas.90.11.5359.
Cerami E et al. The cBio Cancer Genomics Portal: an open platform for exploring multidimensional cancer genomics data. Cancer Discov. 2012;2(5):401–4. https://doi.org/10.1158/2159-8290.CD-12-0095.
Tate JG et al. COSMIC: the Catalogue Of Somatic Mutations In Cancer. Nucleic Acids Res. 2019;47(D1):D941–7. https://doi.org/10.1093/nar/gky1015.
Barretina J et al. The Cancer Cell Line Encyclopedia enables predictive modelling of anticancer drug sensitivity. Nature. 2012;483(7391):603–7. https://doi.org/10.1038/nature11003.
Kitisin K et al. Disruption of transforming growth factor- b signaling through b -spectrin ELF leads to hepatocellular cancer through cyclin D1 activation. Oncogene. 2007;26:7103–10. https://doi.org/10.1038/sj.onc.1210513.
Baek MJ et al. “p16 is a major inactivation target in hepatocellular carcinoma”, Cancer no. Cdk. 2000;4:60–8.
Liew CT et al. High frequency of p16 INK4A gene alterations in hepatocellular carcinoma. Oncogene. 1999;18:789–95.
Tannapfel A et al. INK4a-ARF alterations and p53 mutations in hepatocellular carcinomas. Oncogene. 2001;20:7104–9.
Zang J et al. P16 gene hypermethylation and hepatocellular carcinoma : a systematic review and meta-analysis. World J Gastroenterol. 2011;17(25):3043–8. https://doi.org/10.3748/wjg.v17.i25.3043.
Yarchoan M et al. “Recent developments and therapeutic strategies against hepatocellular carcinoma”. 2019;79(17):4326–4331. https://doi.org/10.1158/0008-5472.CAN-19-0803.
Nault JC et al. Clinical impact of genomic diversity from early to advanced hepatocellular carcinoma. Hepatology. 2019;71:164–82. https://doi.org/10.1002/hep.30811.
Nagaraj NS, Datta PK. Targeting the transforming growth factor-β signaling pathway in human cancer. Expert Opin Investig Drugs. 2010;19(1):77–91. https://doi.org/10.1517/13543780903382609.
Haque S, Morris JC. Transforming growth factor-β: a therapeutic target for cancer. Hum Vaccines Immunother. 2017;13(8):1741–50. https://doi.org/10.1080/21645515.2017.1327107.
Caja L et al “Differential intracellular signalling induced by TGF- β in rat adult hepatocytes and hepatoma cells : implications in liver carcinogenesis”. 2007;19:683–694. https://doi.org/10.1016/j.cellsig.2006.09.002.
Sanchez A, Alberto MA, Benito M, Fabregat I. Apoptosis induced by transforming growth factor-β in fetal hepatocyte primary cultures. J Biol Chem. 1996;271(13):7416–22. https://doi.org/10.1074/jbc.271.13.7416.
Kim DH, Kim WD, Kim SK, Moon DH, Lee SJ. “TGF-β1-mediated repression of SLC7A11 drives vulnerability to GPX4 inhibition in hepatocellular carcinoma cells,” Cell Death Dis. 2007;11(5). https://doi.org/10.1038/s41419-020-2618-6.
Jong H et al. Attenuation of transforming growth factor β-induced growth inhibition in human hepatocellular carcinoma cell lines by cyclin D1 overexpression. Biochem Biophys Res Commun. 2002;292:383–9. https://doi.org/10.1006/bbrc.2002.6666.
Yakicier MC, Irmak MB, Romano A, Kew M, Ozturk M. Smad2 and Smad4 gene mutations in hepatocellular carcinoma. Oncogene. 1999;18:4879–83. https://doi.org/10.1038/sj.onc.1202866.
Di Guglielmo GM, Le Roy C, Goodfellow AF, Wrana JL. “Distinct endocytic pathways regulate TGF- β receptor signalling and turnover”. Nat Cell Biol. 2003;5. https://doi.org/10.1038/ncb975.
Mishra B et al. “Loss of cooperative function of transforming growth factor- b signaling proteins, smad3 with embryonic liver fodrin, a b -spectrin, in primary biliary cirrhosis”. 2004;(5)637–645. https://doi.org/10.1111/j.1478-3231.2004.0958.x.
Dijke P, Miyazono K, Heldin C. Signaling inputs converge on nuclear effectors in TGF-β signaling. Trends Biochem Sci. 2000;25(2):64–70. https://doi.org/10.1016/S0968-0004(99)01519-4.
Vidakovic AT et al. Context-specific effects of TGF-β/SMAD3 in cancer are modulated by the epigenome. Cell Rep. 2015;13:2480–90. https://doi.org/10.1016/j.celrep.2015.11.040.
Hill CS. “Transcriptional control by the SMADs,” Cold Spring Harb Perspect Biol. 2016;8. https://doi.org/10.1101/cshperspect.a022079.
Kojima T et al. Transforming growth factor-β induces epithelial to mesenchymal transition by down-regulation of claudin-1expression and the fence function in adult rat hepatocytes. Liver Int. 2008;28(4):534–45. https://doi.org/10.1111/j.1478-3231.2007.01631.x.
Ohashi S et al. Epidermal growth factor receptor and mutant p53 expand an esophageal cellular subpopulation capable of epithelial-to-mesenchymal transition through ZEB transcription factors. Cancer Res. 2010;70(10):4174–85. https://doi.org/10.1158/0008-5472.CAN-09-4614.
Postigo AA. Opposing functions of ZEB proteins in the regulation of the TGFβ/BMP signaling pathway. EMBO J. 2003;22(10):2443–52. https://doi.org/10.1093/emboj/cdg225.
Acun T, Oztas E, Yagci T, Yakicier MC. SIP1 is downregulated in hepatocellular carcinoma by promoter hypermethylation. BMC Cancer. 2011;11(1):223. https://doi.org/10.1186/1471-2407-11-223.
Acknowledgements
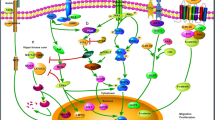
Serif Senturk is a recipient of the Young Scientists Award Program of the Turkish Academy of Sciences (TUBA GEBIP 2017) and the Science Academy’s Young Scientist Awards Program (BAGEP 2019). The schematic for the canonical TGF-β signaling cascade was created with BioRender.com. pSBE4-Luc was a gift from Bert Vogelstein (Addgene plasmid # 16495) and p3TP-Lux was a gift from Joan Massague (Addgene plasmid # 11767).
Author information
Authors and Affiliations
Contributions
MZG and MU performed cell cycle experiments as well as BrdU incorporation assay and analyzed the data. The remaining experiments were carried out by SS. SS and MO conceived and designed the study and analyzed the data. MZG, MU, and SS all made substantial contributions to the preparation of the figures and drafting the manuscript. The final manuscript was read and approved by all authors.
Corresponding author
Ethics declarations
Conflict of Interest
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Gungor, M.Z., Uysal, M., Ozturk, M. et al. Systematic Analysis of Cytostatic TGF-Beta Response in Mesenchymal-Like Hepatocellular Carcinoma Cell Lines. J Gastrointest Canc 52, 1320–1335 (2021). https://doi.org/10.1007/s12029-021-00704-z
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12029-021-00704-z