Abstract
An endo-β-1,4-mannanase encoding gene, man5, was cloned from Bispora antennata CBS 126.38, which was isolated from a beech stump. The cDNA of man5 consists of 1,299 base pairs and encodes a 432-amino-acid protein with a theoretical molecular mass of 46.6 kDa. Deduced MAN5 exhibited the highest amino acid sequence identity of 58% to a β-mannanase of glycoside hydrolase family 5 from Aspergillus aculeatus. Recombinant MAN5 was expressed in Pichia pastoris and purified to electrophoretic homogeneity. The specific activity of the final preparation towards locust bean gum was 289 U mg−1. MAN5 showed optimal activity at pH 6.0 and 70 °C and had good adaptation and stability over a broad range of pH values. The enzyme showed more than 60% of peak activity at pH 3.0–8.0 and retained more than 80% of activity after incubation at 37 °C for 1 h in both acid and alkaline conditions (pH 4.0–11.0). The K m and V max values were 1.33 mg ml−1 and 444 μmol min−1 mg−1 and 1.17 mg ml−1 and 196 μmol min−1 mg−1 for locust bean gum and konjac flour, respectively. Of all tested metal ions and chemical reagents, Co2+, Ni2+, and β-mercaptoethanol enhanced the enzyme activity at 1 mM, whereas other chemicals had no effect on or partially inhibited the enzyme activity. MAN5 was highly resistant to acidic and neutral proteases (trypsin, α-chymotrypsin, collagenase, subtilisin A, and proteinase K). By virtue of the favorable properties of MAN5, it is possible to apply this enzyme in the paper and food industries.



Similar content being viewed by others
References
Suurnäkki, A., Heijnesson, A., Buchert, J., Tenkanan, M., Viikari, L., & Westermark, V. (1996). Chemical characterization of the surface layers of unbleached pine and birch kraft pulp fibres. Journal of Pulp and Paper Science, 22, 78–83.
Kuhad, R. C., Singh, A., & Eriksson, K. E. (1997). Microorganisms and enzymes involved in the degradation of plant fiber cell walls. Advances in Biochemical Engineering/Biotechnology, 57, 45–125.
McCleary, B. V. (1988). Purification of (1 → 3), (1 → 4)-β-D-glucan from barley flour. Methods in Enzymology, 160, 596–610.
Puls, J. (1997). Chemistry and biochemistry of hemicelluloses: Relationship between hemicellulose structure and enzymes required for hydrolysis. Macromolecular Symposia, 120, 183–196.
Zhang, M., Chen, X., Zhang, Z., Sun, C., Chen, L., He, H., et al. (2009). Purification and functional characterization of endo-β-mannanase MAN5 and its application in oligosaccharide production from konjac flour. Applied Microbiology and Biotechnology, 83, 865–873.
Dhawan, S., & Kaur, J. (2007). Microbial mannanases: An overview of production and applications. Critical Reviews in Biotechnology, 27, 197–216.
Xu, B., Sellos, D., & Janson, J. C. (2002). Cloning and expression in Pichia pastoris of a blue mussel (Mytilus edulis) β-mannanase gene. European Journal of Biochemistry, 269, 1753–1760.
Zahura, U. A., Rahman, M. M., Inoue, A., Tanaka, H., & Ojima, T. (2011). An endo-β-1,4-mannanase, AkMan, from the common sea hare Aplysia kurodai. Comparative Biochemistry and Physiology. Part B, Biochemistry & Molecular Biology, 159, 227–235.
Moreira, L. R., & Filho, E. X. (2008). An overview of mannan structure and mannan-degrading enzyme systems. An overview of mannan structure and mannan-degrading enzyme systems. Applied Microbiology and Biotechnology, 79, 165–178.
Shi, P., Yuan, T., Zhao, J., Huang, H., Luo, H., Meng, K., et al. (2010). Genetic and biochemical characterization of a protease-resistant mesophilic β-mannanase from Streptomyces sp. S27. Journal of Industrial Microbiology and Biotechnology, 38, 451–458.
Henrissat, B. (1991). A classification of glycosyl hydrolases based on amino acid sequence similarities. Biochemistry Journal, 280, 309–316.
Zhao, Y., Zhang, Y., Cao, Y., Qi, J., Mao, L., Xue, Y., et al. (2011). Structural analysis of alkaline β-mannanase from alkaliphilic Bacillus sp. N16-5: Implications for adaptation to alkaline conditions. PLoS One, 6, 1–12.
Yoon, K. H., & Lim, B. L. (2007). Cloning and strong expression of a Bacillus subtilis WL-3 mannanase gene in B. subtilis. Journal of Microbiology and Biotechnology, 17, 1688–1694.
Chen, X., Cao, Y., Ding, Y., Lu, W., & Li, D. (2007). Cloning, functional expression and characterization of Aspergillus sulphureus β-mannanase in Pichia pastoris. Journal of Biotechnology, 128, 452–461.
Luo, H., Wang, Y., Wang, H., Yang, J., Yang, Y., Huang, H., et al. (2009). A novel highly acidic β-mannanase from the acidophilic fungus Bispora sp. MEY-1: gene cloning and overexpression in Pichia pastoris. Applied Microbiology and Biotechnology, 82, 423–461.
Duruksu, G., Ozturk, B., Biely, P., Bakir, U., & Ogel, Z. B. (2009). Cloning, expression and characterization of endo-β-1,4-mannanase from Aspergillus fumigatus in Aspergillus sojae and Pichia pastoris. Biotechnology Progress, 25, 271–276.
Zhao, J., Shi, P., Luo, H., Yang, P., Zhao, H., Bai, Y., et al. (2010). An acidophilic and acid-stable β-mannanase from Phialophora sp. P13 with high mannan hydrolysis activity under simulated gastric conditions. Journal of Agriculture and Food Chemistry, 58, 3184–3190.
Liu, Y. G., & Whittier, R. F. (1995). Thermal asymmetric interlaced PCR: automatable amplification and sequencing of insert end fragments from P1 and YAC clones for chromosome walking. Genomics, 25, 674–681.
Sabini, E., Schubert, H., Murshudov, G., Wilson, K. S., Siika-Ahob, M., & Penttilä, M. (2000). The three-dimensional structure of a Trichoderma reesei β-mannanase from glycoside hydrolase family 5. Acta Crystallographica, 56, 3–13.
Ramachandran, G. N., Ramakrishnan, C., & Sasisekharan, V. (1963). Stereochemistry of polypeptide chain configurations. Journal of Molecular Biology, 7, 95–99.
Miller, G. L. (1959). Use of dinitrosalicylic acid reagent for determination of reducing sugar. Analytical Chemistry, 31, 426–428.
Wang, H., Luo, H., Bai, Y., Wang, Y., Yang, P., Shi, P., et al. (2009). An acidophilic beta-galactosidase from Bispora sp. MEY-1 with high lactose hydrolytic activity under simulated gastric conditions. Journal of Agriculture and Food Chemistry, 57, 5535–5541.
Laemmli, U. K. (1970). Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature, 227, 680–685.
Yang, P., Shi, P., Wang, Y., Bai, Y., Meng, K., Luo, H., et al. (2007). Cloning and overexpression of a Paenibacillus beta-glucanase in Pichia pastoris: purification and characterization of the recombinant enzyme. Journal of Microbiology and Biotechnology, 17, 58–66.
Christgau, S., Kauppinen, S., Vind, J., & Kofod, L. V. (1994). Expression cloning, purification and characterization of a beta-1,4-mannanase from Aspergillus aculeatus. Biochemisty and Molecular Biology International, 33, 917–925.
Tang, C., Guo, J., Wu, M., Zhao, S., Shi, H., Li, J., et al. (2011). Cloning and bioinformatics analysis of a novel acidophilic β-mannanase gene, Auman5A, from Aspergillus samii YL-01-78. World Journal of Microbiology and Biotechnology. doi:10.1007/s11274-011-0775-6.
Bauer, S., Vasu, P., Persson, S., Mort, A. J., & Somerville, C. R. (2006). Development and application of a suite of polysaccharide-degrading enzymes for analyzing plant cell walls. Proceedings of the National Academy of Sciences of the United States of America, 103, 11417–11422.
Song, J. M., Nam, K. W., Kang, S. G., Kim, C. G., Kwon, S. T., & Lee, Y. H. (2008). Molecular cloning and characterization of a novel cold-active β-1,4-D-mannanase from the Antarctic springtail, Cryptopygus antarcticus. Comparative Biochemistry and Physiology. Part B, Biochemistry & Molecular Biology, 151, 32–40.
Araujo, A., & Ward, O. P. (1990). Extracellular mannanases and galactanases from selected fungi. Journal of Industrial Microbiology and Biotechnology, 6, 171–178.
Kote, N. V., Patil, A. G., & Mulimani, V. H. (2009). Optimization of the production of thermostable endo-β-1,4 mannanases from a newly isolated Aspergillus niger gr and Aspergillus flavus gr. Applied Biochemistry and Biotechnology, 152, 213–223.
Puchart, V., Vrsanská, M., Svoboda, P., Pohl, J., Ogel, Z. B., & Biely, P. (2004). Purification and characterization of two forms of endo-β-1,4-mannanase from a thermotolerant fungus, Aspergillus fumigates IMI 385708. Biochimica et Biophysica Acta (BBA)—General Subjects, 1674, 239–250.
Arisan-Atac, I., Hodits, R., Kristufek, D., & Kubicek, C. P. (1993). Purification, and characterization of a β-mannanase of Trichoderma reesei C-30. Applied Microbiology and Biotechnology, 39, 58–62.
Ratanachomsri, U., Sriprang, R., Sornlek, W., Buaban, B., Champreda, V., Tanapongpipat, S., & Eurwilaichitr, L. (2006). Thermostable xylanase from Marasmius sp.: Purification and characterization. Journal of Biochemistry and Molecular Biology, 39, 105–110.
Cai, H., Shi, P., Huang, H., Luo, H., Bai, Y., Yang, P., et al. (2011). An acidic β-mannanase from Penicillium sp. C6: gene cloning and over-expression in Pichia pastoris. World Journal of Microbiology and Biotechnology. doi:10.1007/s11274-011-0759-6.
Naganagouda, K., Salimath, P. V., & Mulimani, V. H. (2009). Purification and characterization of endo-β-1,4 mannanase from Aspergillus niger gr for application in food processing industry. Journal of Microbiology and Biotechnology, 19, 1184–1190.
Takeda, N., Hirasawa, K., Uchimura, K., Nogi, Y., Hatada, Y., Usami, R., et al. (2004). Purification and enzymatic properties of a highly alkaline mannanase from alkaliphilic Bacillus sp. strain JAMB-750. Journal of Biological Macromolecules, 4, 67–74.
He, X., Liu, N., Li, W., Zhang, Z., Zhang, B., & Ma, Y. (2008). Inducible and constitutive expression of a novel thermostable alkaline β-mannanase from alkaliphilic Bacillus sp. N16-5 in Pichia pastoris and characterization of the recombinant enzyme. Enzyme and Microbial Technology, 43, 13–18.
Acknowledgements
This work was supported by the Earmarked Fund for the Key Program of Transgenic Plant Breeding (2009ZX08003-020B) and the Agricultural Science and Technology Conversion Funds (2009 GB23260444).
Author information
Authors and Affiliations
Corresponding authors
Electronic supplementary materials
Below is the link to the electronic supplementary material.
Fig. S1
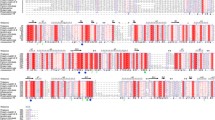
The cDNA nucleotide and deduced amino acid sequences of man5. The putative signal peptide is underlined. The stop codon is indicated with an asterisk (JPEG 98 kb)
Fig. S2
The tertiary structure of MAN5 predicted with Accelrys Discovery Studio software using the GH 5 β-mannanase (PDB: 1QNO) from T. reesei (41% identity) as the template. The putative catalytic residues are indicated in black (JPEG 64 kb)
Rights and permissions
About this article
Cite this article
Liu, Q., Yang, P., Luo, H. et al. A Novel endo-1,4-β-Mannanase from Bispora antennata with Good Adaptation and Stability over a Broad pH Range. Appl Biochem Biotechnol 166, 1442–1453 (2012). https://doi.org/10.1007/s12010-011-9537-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12010-011-9537-z