Abstract
Glutamate dehydrogenase (GDH; EC 1.4.1.2) catalyzes the reductive amination of 2-oxoglutarate with ammonium and the oxidative deamination of glutamate. In some plants species, GDH is a hexamer and can be separated into seven isoenzymes that are composed of two distinct subunits: α and β. The large number of isoenzymes is the reason why GDH functions are still being intensively researched and widely studied. Until recently, the β subunit of GDH in triticale, a common Polish cereal, was thought to be encoded by the TsGDH1 gene, which can undergo posttranslational modifications and form a heterohexameric enzyme. Here, we report the cloning and molecular characterization of a second glutamate dehydrogenase gene—TsGDH2 (encoding α subunits). The TsGDH2 cDNA contains a 1236-bp open reading frame encoding a 411-amino-acid polypeptide with a calculated molecular mass of 44.5 kDa. To clarify the role of TsGDH2 in triticale, we used triticale GDH α-subunit cDNA to generate transgenic A. thaliana lines with increased and decreased GDH activity via alternation of α-subunit levels.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Glutamate dehydrogenase (GDH; EC 1.4.1.2) catalyzes reversible amination/deamination reactions in vitro, which can lead to either the synthesis or catabolism of glutamate. Because ammonium ions are one of the substrates of the amination reaction, GDH may be involved in the assimilation of ammonia. Therefore, GDH was previously considered the main ammonia (NH4 +)-assimilating enzyme in higher plants (Bullen 1956). However, since the discovery of glutamine synthetase and glutamate synthase (GS; EC 6.1.1.3 and NADH-GOGAT; EC 1.4.1.14; Fd-GOGAT; 1.4.7.1), the GS/GOGAT cycle has been recognized as the main route of ammonium assimilation/reassimilation (Lea and Mifflin 1974). Thereafter, the physiological functions of GDH have been the subject of extensive study. Many authors believe that NADH-GDH can act in vivo as an ammonium-assimilating enzyme only under certain conditions, for instance, in dark-stressed plants, under abiotic stress, and during the processes of natural plant aging and of seed formation (Grabowska et al. 2012; Kwinta and Kolik 2006; Skopelitis et al. 2006). Other authors have suggested that GDH plays a major role in amino acid catabolism during seed germination, plant senescence and nutrient remobilization (Aubert et al. 2001; Grabowska et al. 2011; Lehmann and Ratajczak 2008).
The functions of GDH are carried out by multiple isoforms of the enzyme, although this diversity is not always possible to explain. In higher plants, GDH functions as a homohexamer or heterohexamer consisting of two subunits, α and β, that differ minimally in terms of their mass and charge. The α and β polypeptides combine in different ratios to form seven possible NADH-GDH isoenzymes in A. thaliana, N. plumbaginifolia or V. vinifera (Fontaine et al. 2006; Loulakakis and Roubelakis-Angelakis 1991; Masclaux-Daubresse et al. 2002). Only one isoform was found to be present in triticale, whereas L. luteus contains fourteen isoforms of this enzyme (Kwinta et al. 2001; Ratajczak et al. 1986). The quantity and type of GDH isoenzymes in plants depends on the type of tissue, developmental stage and environmental growth conditions. NADH-dependent enzyme activity is usually higher in roots, developing seeds and ripening fruits compared with that in leaves (Singh and Srivastava 1982; Nanda et al. 1991; Kwinta et al. 2001). In A. thaliana, the most anodal isoenzyme, i.e., the homohexamer of α subunits, is active in both roots and floral stems, whereas the most cathodal isoenzyme, i.e., the homohexamer of β subunits, is active in leaves (Fontaine et al. 2006, 2013).
In most plant species, GDH is encoded by two separate nuclear genes: GDH1, also referred to as GDHB, encoding the β subunit, and GDH2, also called GDHA, encoding the α subunit (Melo-Oliveria et al. 1996; Restivo 2004; Turano et al. 1997). In some plant species, the α and β subunits may be encoded by more than one gene. For example, three genes encoding the α subunit were cloned in tomato plants (Ferraro et al. 2012), two genes encode the β subunit in rice (Qiu et al. 2009), and three genes encode the β subunit in L. luteus (Lehmann et al. 2011). Recent studies have shown that A. thaliana contains a third gene (GDH3) that encodes a γ subunit, which is expressed only in roots (Fontaine et al. 2012). The α, β and γ subunits can combine to form heterohexamers composed of the three subunits in different ratios (Fontaine et al. 2013). Such a large variety of isoenzymes provides opportunities for detailed analyses and in-depth examination of the physiological functions of individual GDH subunits. These studies are very often conducted on genetic mutants and transgenic plants that either contain additional GDH encoding genes or have these genes silenced in heterologous or homologous systems (Labboun et al. 2009; Purnell et al. 2005; Skopelitis et al. 2007; Tercé-Laforgue et al. 2013, 2015). One of the strategies to identify gene function is to perform comparative analyses of phenotypes obtained after the introduction of a gene construct that causes either gene silencing or overexpression. Purnell and Botella (2007) obtained transgenic tobacco plants (N. tabacum) overexpressing a gene derived from tomato (S. lycopersicum) encoding the β subunit. Analyses of the transformants demonstrated that GDH isoenzyme 1 (β-subunit homohexamer) exclusively catalyzes glutamic acid deamination. In contrast, Skopelitis et al. (2007) obtained tobacco plants with a gene construct derived from a vine plant (V. vinifera) encoding the α subunit. In this case, high deamination and amination activity was observed in plants heterologously overexpressing this gene (Skopelitis et al. 2007). Fontaine et al. (2006) applied an antisense strategy and obtained transgenic tobacco plants with impaired production of either the α- or β-subunit. A similar investigation was performed in A. thaliana mutants gdh1 (SALK_042736) and gdh2 (SALK_102711), which fails to express the β- and α-subunit, respectively.
The gene encoding the GDH β subunit was previously cloned in triticale (Grabowska et al. 2011). The protein product of this gene was hypothesized to undergo posttranslational modifications and form a hexameric enzyme with the same electrophoretic mobility as that of the heterohexameric GDH enzyme in A. thaliana. In the present study, a full-length cDNA encoding GDH α subunit was isolated from triticale and designated as TsGDH2. Since only one GDH isoform is present in triticale, this study investigates the function of the cloned gene. To analyze the function of TsGDH2, transgenic A. thaliana plants were obtained by introducing the coding region of TsGDH2 linked to the 35S CaMV promoter, in either the sense or antisense orientation, via Agrobacterium tumefaciens transformation. We generated transgenic A. thaliana lines with increased and decreased GDH activity via alternation of α-subunit levels.
Materials and methods
Plant material and growth conditions
Seeds of triticale (×Triticosecale Wittm.) cv. Witon were obtained from the Breeding Station in Laski, Poland. The grains were surface sterilized with 10% hypochlorite and then germinated in darkness at 20 °C and 100% relative humidity. Samples were collected after 72 h of germination, immediately frozen in liquid nitrogen and stored at −80 °C until use.
The seeds of the A. thaliana ecotype Columbia (Col-0) were obtained from the Nottingham Arabidopsis Stock Center. Germination of the seeds was synchronized with treatment at 4 °C for 48 h in the dark. Plants were grown to the flowering stage in a growth chamber (Versatile Environmental Test Chamber MLR-35OH, Sanyo, UK) at 22 °C with 10 h light (250 μE m−2 s−1) during the vegetative stage and 14 h light during generative growth.
Cloning procedures
The RNA was isolated from the shoots and roots of 3-day-old triticale seedlings using the guanidinium thiocyanate/acidic phenol-extraction method (Chomczynski and Saschi 2006). To eliminate any genomic DNA contamination, each RNA sample was treated with RNAse-free DNAse I (Fermentas). First-strand cDNA was synthesized using 2 μg of total RNA primed with an oligo(dT)12–18 primer and an avian myeloblastosis virus reverse transcriptase (AMV RT) following the manufacturer’s protocol (Promega).
Two oligonucleotide primers, GDH2-F1 (5′GGCTCTCCTGGCCGAATATGGGAAGTCT3′) and GDH2-R1 (5′CCTAAGGTTGCAATCTTGAGACTT3′), were used to amplify the internal region of TsGDH2 cDNA with GoTaq Flexi DNA Polymerase (Promega). Both primers corresponded to a fragment of the Brachypodium distachyon gene encoding glutamate dehydrogenase 2 (GenBank accession number: XM_003580211). PCR reactions were performed in a PTC-200 Peltier Thermal Cycler (MJ Research) under the following conditions: 2 min at 94 °C, 35 cycles of 30 s at 94 °C, 30 s at 58 °C, 1 min at 72 °C, and a final extension step for 7 min. The PCR amplification product was separated by agarose gel electrophoresis and extracted with a Silica Bead DNA gel Extraction kit (Thermo Scientific). The resulting amplified fragment was cloned into pGEM-T Easy (Promega) and sequenced.
Full-length TsGDH2 cDNA was obtained using the GeneRacer Kit (Invitrogen). The gene specific primers used for RACE were designed from the above partial TsGDH2 cDNA sequence. The primer GDH2-R2 (5′AATAACAAAGGTTGATCCAGAAATAGAC3′) was used for 5′ RACE, and GDH2-F2 (5′-GTACCTGCTCTGATGAAGCACAGAAAT3′) was used for 3′ RACE. PCR reactions were performed under the following conditions: 2 min at 94 °C, 35 cycles of 30 s at 94 °C, 30 s at 58 °C, 1 min at 68 °C, and a final 7 min at 68 °C. The amplification was performed with Platinum Taq DNA Polymerase (Invitrogen). The resulting amplified fragment was cloned using the TOPO TA Cloning Kit (Invitrogen) and sequenced. Sequencing was performed with the ABI Prism BigDye Terminator Cycle Sequencing Kit on the ABI Prism 3730 DNA analyzer (Applied Biosystems), at the DNA Sequencing and Oligonucleotide Synthesis Laboratory, The Institute of Biochemistry and Biophysics, Polish Academy of Sciences.
Bioinformatics analysis
The obtained nucleotide sequences date from this article have been deposited in GenBank. Sequences were verified by a database search in the National Center for Biotechnology Information server using the BLAST algorithm (http://www.ncbi.nlm.nih.gov). The deduction of the amino acid sequence, calculation of the theoretical molecular mass and pI was performed with ExPASy (http://www.expasy.ch/tools/). Multiple-sequence alignments of GDH amino acids sequences were generated using the CLUSTAL W program (Thompson et al. 1997). A phylogenetic analysis was performed using the neighbor-joining (NJ) method, as implemented in the PhyML program (http://www.phylogeny.fr/version2_cgi/index.cgi) (Dereeper et al. 2008).
Gene constructs
Agrobacterium tumefaciens strain EHA105 carrying pCAMBIA1380 was used to transform A. thaliana plants. The TsGDH2 coding sequence (1236 bp) was amplified with gene specific primers (forward: 5′-ggaaagctt/ggaaagctt-ATGAACGCGCTCGCCGCGACCAGCCGC3′; reverse: 5′taggttacc/ggcgaattc-TCATGCCTCCCAGCCCCTCAAGAT3′). HindIII and Eco91I or HindIII and EcoRI restriction sites were introduced at the ends of the primers for the sense and antisense orientations of TsGDH2, respectively. The amplification was performed with GoTaq Flexi DNA Polymerase (Promega). PCR reactions were performed under the following conditions: 3 min at 95 °C, 35 cycles of 30 s at 95 °C, 30 s at 68 °C, 60 s at 72 °C, and a final 5 min at 72 °C. After a restriction analysis, purified fragments were ligated to the pCAMBIA1380 vector. These T-DNA constructs were designated as 35S::TsGDH2s for the sense and 35S::TsGDH2as for the antisense orientation. All constructs for transformation were verified by sequencing.
Plant transformation
The floral dip method was used for plant transformation (Clough and Bent 1998). Plants designated as T0 were grown to maturity, and their seeds were harvested. Seeds from transformed plants (T1 generation) were surface sterilized by immersion in 96% (v/v) ethanol for 1 min, washed with sterile water and then diluted in bleach solution (final concentration 20% v/v sodium hypochlorite) for 5 min, washed five times with sterile water, suspended in 0.1% sterile agarose and plated on selective medium (1/2 MS) (Murashige and Skoog 1962) containing 1% sucrose, 0.8% agar, 20 mg L−1 hygromycin, and 200 mg L−1 cefotaxime. The growth conditions were as described above. After 2 weeks, resistant seedlings were transferred to fresh selection medium with 1.5% agar. After 1 week, seedlings were transferred to soil. The T2 seeds from each T1 plant were harvested. Progeny (T3) were selected from each T2 line that showed a 3:1 ratio of hygromycin resistance consistent with single-locus insertion of the transgene.
Analysis of A. thaliana transformants
Genomic DNA was isolated from T3 transformants and wild-type plants (control) using the CTAB extraction protocol adapted from Weigel and Glazebrook (2009). Antibiotic-resistant plants were screened by PCR for the hygromycin phosphotransferase gene (hpt) (primers: 5′GGCGAGTACTTCTACACA3′ and 5′GCGAAG AATCTCGTGCTT3′), the 35S promoter (primers: 5′CATGGAGTCAAAGATTCAAATAGAGGA3′ and 5′CTCTCCAAATGAAATGAACTTCCTTA3′) and the TsGDH2 cDNA sequence (primers: 5′ATGAACGCGCTCGCCGCGACCAGCCGC3′ and 5′TCATGCCTCCCAGCCCCTCAAGAT3′). PCR reactions were performed under the following conditions: 3 min at 94 °C, 35 cycles of 30 s at 94 °C, 30 s at 55 °C (for hpt gene and 35S promoter) or 65 °C (for TsGDH2 gene), 30 s (for hpt gene and 35S promoter) or 1 min (for TsGDH2 gene) at 72 °C, and a final 3 min at 72 °C. Amplification was performed with GoTaq Flexi DNA Polymerase (Promega).
Expression analysis by semi-quantitative RT-PCR
Total RNA was extracted from 100 mg of young leaves of A. thaliana transgenic lines and control plants. Semi-quantitative RT-PCR analysis was performed using the One-Step RT-PCR Kit (Novagen) according to the manufacturer’s instructions. In the expression analysis, three primer pairs were used (Table 1). The first pair was designed for the triticale gene (TsGDH2), the second for the endogenous GDH2 gene (AT5G07440), and the third pair for the reference Actin 2 gene in A. thaliana. Reactions were performed under the following conditions: 30 min at 60 °C, 2 min at 94 °C, 35 cycles (for TsGDH2, GDH2) and 30 cycles (for Actin 2) of 30 s at 94 °C, 30 s at X °C (Table 1), 30 s at 72 °C and a final extension step for 5 min at 72 °C. Each sample was analyzed with at least three independent experiments. The amplified products were analyzed by electrophoresis on 1.5% agarose gels containing ethidium bromide.
Protein extraction, enzyme assays and electrophoresis
GDH was extracted by homogenizing 100 mg of leaves in 1 mL of extraction buffer containing 100 mM Tris–HCl (pH 7.6), 5 mM 2-mercaptoethanol and 20 µM PMSF. The homogenates were centrifuged for 15 min at 13,000×g at 4 °C, and the supernatant was used as the enzyme extract. GDH activity was determined in both the aminating (NADH-GDH) and deaminating (NAD+-GDH) directions according to the method of Barash et al. (1973). The standard amination reaction mixture contained 100 mM Tris–HCl (pH 8.3), 200 mM NH4Cl, 0.28 mM NADH, 32 mM 2-oxoglutarate, 0.05 mL of enzyme extract and deionized water to a final volume of 0.6 mL. The standard deamination reaction mixture contained 100 mM Tris–HCl (pH 9.2), 200 mM l-glutamate, 0.25 mM NAD+, 0.05 mL of enzyme extract and deionized water to a final volume of 0.6 mL. All assays were performed at 30 °C. One unit of GDH activity was defined as the reduction or oxidation of 1 µmol of coenzyme (NAD+ or NADH, respectively) min−1 g−1 DW (dry weight). Enzyme activity measurements are presented as the mean ± SD for three independent experiments, with two replicates each. Statistical significance was determined using Student’s t test.
Native PAGE of GDH extracts was performed using the modified Laemmli method (1970) with 7.5% resolving and 4% stacking gels. Bands with GDH activity were visualized on the gel using the tetrazolium system (Lehmann et al. 1990).
Protein concentration was determined colorimetrically according to the Bradford method (1976) using bovine serum albumin as a standard.
Results
Sequence analyses of TsGDH2
The TsGDH2 gene was isolated using RT-PCR. The full-length TsGDH2 cDNA (Gene Bank, accession number: KU311056) comprised 1538 bp, with a 1236-bp open reading frame (ORF) and a 302-bp 3′untranslated region (3′UTR). The putative protein sequence of TsGDH2 contained 411 amino acids, with a calculated molecular mass of 44.5 kDa and a predicated isoelectric point (pI) of 6.20. The deduced amino acid sequence of TsGDH2 was screened for conserved domains at the Conserved Domain Database at NCBI (http://www.ncbi.nlm.nih.gov/Structure/cdd/cdd.shtml). Two features were present in the TsGDH2 protein: an NAD(P) binding domain of glutamate dehydrogenase from Gly-176 to Arg-402 and a Glu/Leu/Phe/Val dehydrogenase (ELFV_dehydrog_N, pfam02812) dimerization domain from Pro-31 to His-161 (Marchler-Bauer et al. 2013). Three conserved amino acids residues, Lys-90, Thr-169 and Ser-344, are involved in Glu binding according to Britton et al. (1992). In TsGDH2, the region Asp-265 to Glu-276 resembles an EF-hand loop motif, which is associated with Ca2+ binding in other proteins (Denessiouk et al. 2014). Additional the TargetP program predicted that TsGDH2 contains an 18-residue mitochondrial target peptide (mTP) at the N terminus (score 0.795, reliability class 2) (Fig. 1).
The nucleotide and putative amino acid sequences of TsGDH2. The mitochondrial target peptide is in bold, the NAD(P)-binding domain of glutamate dehydrogenase is underlined, the dehydrogenase dimerization domain is double underlined, the three amino acids residues involved in Glu binding are marked by an ellipse, and the region containing the EF-hand loop motif is boxed
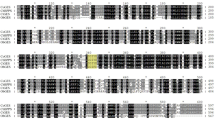
The overall sequence alignment analysis revealed that TsGDH2 is highly homologous to GDHs in other plants (Supplementary Material Fig. 1). The TsGDH2 protein showed high identity to the selected GDH of B. distachyon (XP_003580259, 94% identity, score 2058), O. sativa (BAE48298.1, 93% identity, score 2044), N. sylvestris (XP_009758042.1, 82% identity, score 1834), N. plumbaginifolia (CAA69601, 79% identity, score 1777), V. vinifera (XP_010662019.1, 82% identity, score 1812). The deduced sequence of TsGDH2 and A. thaliana GDH2 (NP_196361.1) showed 83% identity and a score of 1861, whereas TsGDH2 and A. thaliana GDH1 (NP_197318.1) showed 79% identity and a score of 1760 (Fig. 2).
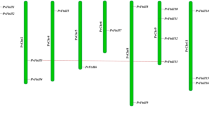
Phylogenetic tree of glutamate dehydrogenase from different plant species. Numbers above the branches represent bootstrap support for 100 replicates. The analysis was based on the deduced amino acid sequences of GDH genes. Oryza sativa—GDH2 (AB189166.1); Brachypodium distachyon—GDH2 (XM_003580211); ×Triticosecale—GDH2 (KU311056); Arabidopsis thaliana—GDH2 (NM_120826.2); Nicotiana plumbaginifolia—GDH2 (Y08293.2); Nicotiana sylvestris—GDH2 (XM_009759740.1); Vitis vinifera—GDH2 (AJ303070.1); Arabidopsis thaliana—GDH1 (NM_121822.3); Nicotiana plumbaginifolia—GDH1 (Y08292.1); Zea mays—GDH1 (NM_001111831.1); ×Triticosecale—GDH1 (HQ658905.1)
In the majority of plant species, GDH is encoded by at least two genes encoding the α and β subunits that make up the enzyme. To determine which subunit (α or β) is encoded by TsGDH2, as well as the evolutionary relationship between TsGDH2 and GDHs from other plants, a phylogenetic tree was constructed using the neighbor-joining method. The phylogenetic tree was generated using the deduced amino acid sequences of monocot and dicot GDHs, encoding both α and β subunits. The phylogenetic analysis indicated that GDH2 (α-subunit) and GDH1 (β-subunit) were each classified into one group containing all known members. Each group contained sequences from monocots and dicots, suggesting that the genes were grouped according to the nature of the subunit encoded, rather than according to the species of origin. The phylogenetic analysis clearly indicated that TsGDH2 belongs to the GDH2-NAD(H) subgroup.
Molecular analysis of A. thaliana transformants
The binary vectors 35S::TsGDH2s and 35S::TsGDH2as containing the cauliflower mosaic virus (CaMV) 35S promoter upstream of the x Triticosecale TsGDH2 gene in the sense and antisense orientation, respectively, were used to transform A. thaliana plants. After Agrobacterium transformation and hygromycin selection, several transformants were produced. Four transgenic lines (A7, A15—TsGDH2 antisense; S3, S4—TsGDH2 sense) were grown up to the T3 generation, from which homozygous plants (for hpt marker) were analyzed. All T3 lines had a single T-DNA insertion locus that segregated in a Mendelian fashion (results not shown). No phenotypic changes were observed transgenic plants compared with controls. Stable transformation of the transgenes was confirmed by PCR. In all the transformants into which the 35S::TsGDH2s and 35S::TsGDH2a cassettes were introduced, 887-bp (for the hpt gene), 525-bp (for the 35S promoter) and 1254-bp (for TsGDH2 gene) products were detected, indicating correct integration of the T-DNA into all the transgenic lines. DNA isolated from non-transformed plants (WT) served as a negative control, whereas the plasmid DNA (pCAMBIA1380 with TsGDH2) was used as a positive control (Fig. 3).
PCR analysis of Arabidopsis transgenic lines. DNA was isolated from transgenic plants transformed with the 35S::TsGDH2s (S3 and S4) or the 35S::TsGDH2as (A7 and A15) cassette. PCR amplification was performed using primers for the hpt gene, the 35S promoter and the TsGDH2 gene. WT-negative control, DNA from wild-type plants; CP-positive control, plasmid DNA (pCAMBIA1380 with TsGDH2); DNA molecular markers (SMO311; Thermo Scientific)
Expression analysis by semi-quantitative RT-PCR and modulation of GDH activity in transgenic plants
To evaluate the effects of TsGDH2 overexpression and antisense repression, the expression levels of the introduced gene and the GDH2 (AT5G07440) gene were analyzed. The transcription level of TsGDH2 and GDH2 genes was determined via semi-quantitative RT-PCR analysis. The results were standardized relative to the Actin 2 (AT3G18780) gene from A. thaliana. Total RNA from the leaves of plants transformed with sense and antisense constructs, as well as from control plants, was used. As seen in Fig. 4, both sets of transformants containing either the sense or antisense constructs exhibited high levels of the corresponding TsGDH2 transcript, whereas wild-type plants did not express the TsGDH2 gene. With regard to endogenous GDH2 gene expression, similar transcript levels were observed in the transgenic and the control plants. Following TsGDH2 and GDH2 expression analysis in the transformants, the next step was to measure both NADH-dependent aminating and NAD-dependent deaminating GDH activity (Fig. 5). In the two overexpressing transgenic lines, the aminating activity was increased 3.2-fold (S3 line) and 2.9-fold (S4 line), and the deaminating activity was increased 4.3-fold (S3 line) and 3.7-fold (S4 line) compared with the wild-type (WT) control plants. However, in the antisense lines, the aminating and deaminating activities were reduced 1.8-fold (A7 line), 2.3-fold (A15 line), 2.2-fold (A7 line) and 2.0-fold (A15 line) compared with the wild-type (WT) control plants.
Semi-quantitative RT-PCR analysis of the TsGDH2 and GDH2 (AT5G07440) genes. Total RNA was isolated from transgenic plants transformed with the 35S::TsGDH2s (S3 and S4) and the 35S::TsGDH2as (A7 and A15) cassette and from wild-type (WT) plants. The Actin 2 gene (AT3G18780) was used to normalize the template amounts in different samples. Each figure is representative of three separate amplification experiments that yielded similar results
GDH activity of wild-type Arabidopsis plants (WT) and transgenic lines: S3 and S4 (35S::TsGDH2s), A7 and A15 (35S::TsGDH2as). Enzyme activities were measured in leaf samples. NAD(H)-GDH aminating activity (black columns), NAD-GDH deaminating activity (white columns). GDH activity was measured on three individual plants for each line. Values are mean ± SE
In-gel activity staining was used to detect the GDH isoenzyme compositions in the leaves of transgenic plants (Fig. 6). Overexpression of the α-subunit polypeptide in 35S::TsGDH2s transformants (S3 and S4) resulted in increased abundance of the anionic isoenzymes, especially the homohexameric isoenzyme 7. The other bands of GDH activity were stronger compared with the control plants. In the two transgenic antisense lines (A7, A15), only the most cathodal isoenzymes (GDH1, 2, 3 and 4) were visible; the other activity bands were much weaker compared with the wild-type plants. For comparative purposes, in addition to the transgenic lines and wild-type A. thaliana, triticale was also analyzed. Only one enzyme isoform, characterized by medium electrophoretic mobility, was observed. This isoform corresponds to isoenzyme 4 of A. thaliana—a heterohexamer composed of three α-subunits and three β-subunits.
NAD-GDH isoenzyme pattern of wild-type Arabidopsis plants (WT), transgenic lines [A7 and A15 (35S::TsGDH2as) and S3 and S4 (35S::TsGDH2s)] and triticale leaves (Ts). The soluble protein extracts of leaves were subjected to native PAGE followed by NAD-GDH in-gel deaminating activity staining. The amount of protein loaded in each lane was calculated on a similar dry-weight basis for each leaf sample. On the left-hand side of the gel, the isoenzyme pattern from triticale leaf extracts (Ts) was used as a marker to verify the corresponding isoenzymes in Arabidopsis. In-gel activity staining was performed on three individual plants for each line
Discussion
The full-length cDNA of TsGDH2, encoding an NAD(H)-GDH in triticale, was identified and characterized. This gene is the second GDH to be cloned in triticale. Previous studies carried out by Grabowska et al. (2011, 2012) analyzed the TsGDH1 gene. Sequence alignments of the previously cloned full-length TsGDH1 gene (Grabowska et al. 2011) and TsGDH2 indicated 71% nucleotide sequence similarity and 81% similarity within the 411-amino-acid sequence (Supplementary Material Figs. 2, 3). Comparable sequence similarity is observed in A. thaliana, where the sequence identity between GDH1 and GDH2 at the nucleotide and amino acid levels is 75 and 81%, respectively (Turano et al. 1997). The deduced amino acid sequence of the TsGDH2 contains a consensus calcium binding EF-loop motif between Asp-265 and Glu-276 (Grabarek 2006). This sequence is characteristic for the GDH α-subunit but is absent from the GDH β-subunit. The region responsible for binding calcium ions has also been identified in the genes encoding the GDH α-subunit in other species, for example, in A. thaliana, N. tabacum, and S. lycopersicum (Ferraro et al. 2012; Purnell et al. 2005; Turano et al. 1997). Research on subcellular localization prediction has demonstrated that GDH is mostly localized in the mitochondria of the phloem companion cells (Loulakakis and Roubelakis-Angelakis 1990; Paczek et al. 2002). Analyses of the deduced amino acid sequence of TsGDH2 revealed the presence of a signal peptide directing the protein to mitochondria. In tomato plants, in which three genes encoding the GDH α subunit were cloned, predicted mitochondrial target peptides were also identified at the N terminus (Ferraro et al. 2012).
Phylogenetic analyses distinguishes two distinct groups of GDH genes. Genes encoding the α-subunit belonged to the first group, whereas genes encoding the β-subunit were clustered in the second group. The phylogenetic analysis of the deduced amino acid sequences encoded by the GDH2 genes identified in triticale and the sequences of α- and β-GDH subunits from selected higher plants confirmed that TsGDH2 encode the α-type subunit.
To elucidate the function of the TsGDH2 gene of triticale, which shares high sequence identity with other genes coding for the GDH α subunit, an antisense repression and overexpression strategy in A. thaliana was used. Analyses of transgenic tobacco plants with modified expression of genes encoding GDH α and β subunits revealed various metabolic functions of GDH isoenzymes and indicated that, under normal growth conditions, GDH isoenzyme 1 (β-homohexamer) only deaminates glutamate, whereas GDH isoenzyme 7 (α-homohexamer) exhibits high deaminating activity but also weak aminating activity (Purnell et al. 2005; Purnell and Botella 2007; Skopelitis et al. 2007).
In the present work, A. thaliana plants were transformed using T-DNA constructs containing the full-length TsGDH2 gene coding for the triticale α-GDH subunit in the sense and antisense orientation. In all the analyzed lines, the transcript of the inserted TsGDH2 gene was present, suggests that the T-DNA was successfully transferred into the A. thaliana genome and then transcribed. In the control group, no TsGDH2 transcript was detected. Furthermore, the levels of endogenous GDH2 gene transcript were similar in the transgenic plants and in the control group. In the overexpressing transgenic lines (S3 and S4), increased GDH aminating and deaminating activity was observed in vitro compared with the control plants. Furthermore, the increase in enzyme activity measured in vitro in lines S3 and S4 was similar to that detected by native gel staining, and the expression of TsGDH2 was consistent with the synthesis of the GDH α-subunit, especially GDH isoenzyme 7. Skopelitis et al. (2007) obtained similar results, namely, increased GDH activity, upon inserting a Vvgdh-NAD derived from grapevine and encoding the α-GDH subunit into transgenic tobacco plants. The in vitro aminating activities were very high in overexpressing transgenic lines. Furthermore, overexpression of the α-subunit resulted in increased abundance of anionic isoenzymes.
In work conducted by Purnell et al. (2005), a gene encoding the β subunit in tomato plants was introduced into the antisense tobacco line. In this study, no transcript of the introduced gene was found; however, reduced GDH activity was observed. The mechanism of gene expression inhibition typically relies on the formation of duplexes between the antisense RNA fragments and the endogenous mRNA. Due to the low level of complementarity between RNA in the formed duplexes, the endogenous RNA is not silenced; therefore, the transcript was still detectable but interfered with the translation of endogenous transcripts responsible for the synthesis of the β subunit. Moreover, according to gene-silencing studies concerning carried out by Cannon et al. (1990) and Oeller et al. (1991), during inhibition by antisense RNA expression, reduced expression of endogenous gene is often observed. Therefore, a different silencing model was proposed in which antisense RNA forms duplexes with the corresponding endogenous sense RNA. Thus, the duplexes are then degraded by RNase activity, resulting in the loss of the target mRNA (Mol et al. 1990). In the obtained A3 and A4 lines, expression of the antisense gene and the endogenous gene coding for the α subunit was observed. Thus, the heteroduplexes that might have formed were likely stable. Related results were found by Temple et al. (1993), who transformed tobacco plants in a heterologous system using an antisense genetic construct containing a glutamate synthetase gene derived from the alfalfa plant. In the transgenic tobacco plants, neither the endogenous gene transcript nor the introduced alfalfa gene was observed. The level of GS polypeptides was found to be reduced. In both above studies, the formed heteroduplexes were not used in the translation process, which in our experiment resulted in decreased aminating and deaminating GDH activity in vitro compared with the control plants. These results were also confirmed by native PAGE NAD-GDH gel activity staining, in which only the most cathodal isoenzymes (GDH 1, 2, 3 and 4) were observed, and the activity of the remaining isoenzymes was lower compared with the control. These results demonstrate that the full-length triticale TsGDH2 gene, when transcribed in the antisense orientation, is capable of downregulating GDH2 and GDH1 in A. thaliana leaves. Therefore, the antisense RNA approach is effective for silencing gene expression even in a heterologous system. TsGDH2 shares 83 and 79% nucleotide sequence homology with GDH2 and GDH1 from A. thaliana. Because the sequence similarity is so large, both genes could have been silenced to some extent, and this result was confirmed by gel electrophoresis.
In summary, a second triticale gene, encoding GDH α subunit, was cloned. To assign a function to this gene, A. thaliana plants were transformed with two genetic constructs in the sense and antisense orientation. The resulting transgenic A. thaliana plants were characterized by an altered level of GDH-NAD(H) activity due to the alteration of α-subunit levels, which indicates that the TsGDH2 protein is functionally analogous to GDH2 in other plants. Our future studies will focus on two areas. First, the transgenic lines will be analyzed further using TsGDH2-specific antibodies. The second aim will be to obtain transgenic A. thaliana plants using the β subunit-coding gene (TsGDH1) in triticale to further clarify the in vivo reaction direction(s), physiological role(s) and regulation of the GDH isoenzymes in triticale. This work is of particular interest, as triticale only contains one enzyme isoform that might be composed of three α and three β subunits.
Author contribution statement
AG conceived and designed the experiments; AG performed the experiments with help JK and EZZ; AG and JK analyzed the data; EK technical support; JK revised the manuscript, AG wrote the manuscript. All authors read and approved the manuscript.
References
Aubert S, Bligny R, Douce R, Gout E, Ratcliffe RG, Roberts JKM (2001) Contribution of glutamate dehydrogenase to mitochondrial glutamate metabolism studied by 13C and 31P nuclear magnetic resonance. J Exp Bot 52:37–45. doi:10.1093/jexbot/52.354.37
Barash I, Sadon T, Mor H (1973) Induction of a specific isoenzyme of glutamate dehydrogenase by ammonia in oat leaves. Nat New Biol 244:150–152. doi:10.1038/newbio244150a0
Bradford MM (1976) A rapid and sensitive method for the quantification of microgram quantities of protein utilizing the principle of protein-dye binding. Anal Biochem 72:248–254. doi:10.1016/0003-2697(76)90527-3
Britton KL, Baker PJ, Rice DW, Stillman TJ (1992) Structural relationship between the hexameric and tetrameric family of glutamate dehydrogenases. Eur J Biochem 209:851–859. doi:10.1111/j.1432-1033.1992.tb17357.x
Bullen WA (1956) The isolation and characterization of glutamic dehydrogenase from corn leaves. Arch Biochem Biophys 62:173–183. doi:10.1016/0003-9861(56)90100-X
Cannon M, Platz J, O’Leary M, Sookdeo C, Cannon F (1990) Organ-specific modulation of gene expression in transgenic plants using antisense RNA. Plant Mol Biol 15:39–47. doi:10.1007/BF00017722
Chomczynski P, Saschi N (2006) The single-step method of RNA isolation by acid guanidinium thiocyanate–phenol–chloroform extraction: twenty-something years on. Nat Protoc 1:581–585. doi:10.1038/nprot.2006.83
Clough SJ, Bent AF (1998) Floral dip: a simplified method for Agrobacterium-mediated transformation of Arabidopsis thaliana. Plant J 16:735–743. doi:10.1046/j.1365-313x.1998.00343.x
Denessiouk K, Permyakov S, Denesyuk A, Permyakov E, Johnson MS (2014) Two structural motifs within canonical EF-hand calcium-binding domains identify five different classes of calcium buffers and sensors. PLoS One 9:e109287. doi:10.1371/journal.pone.010928
Dereeper A, Guignon V, Blanc G, Audic S, Buffet S, Chevenet F, Dufayard JF, Guindon S, Lefort V, Lescot M, Claverie JM, Gascuel O (2008) Phylogeny.fr: robust phylogenetic analysis for the non-specialist. Nucleic Acids Res 36:W465–W469. doi:10.1093/nar/gkn180
Ferraro G, Bortolotti S, Mortera P, Schlereth A, Stitt M, Carrari F, Kamenetzky L, Valle EM (2012) Novel glutamate dehydrogenase genes show increased transcript and protein abundances in mature tomato fruits. J Plant Physiol 169:899–907. doi:10.1016/j.jplph.2012.02.002
Fontaine JX, Saladino F, Agrimonti C, Bedu M, Tercé-Laforgue T, Tétu T, Hirel B, Restivo FM, Dubois F (2006) Control of the synthesis and subcellular targeting of the two GDH genes products in leaves and stems of Nicotiana plumbaginifolia and Arabidopsis thaliana. Plant Cell Physiol 47:410–418. doi:10.1093/pcp/pcj008
Fontaine J-X, Tercé-Laforgue T, Armengaud P, Clément G, Renou J-P, Pelletier S, Catterou M, Azzopardi M, Gibon Y, Lea PJ, Hirel B, Duboise F (2012) Characterization of a NADH-dependent glutamate dehydrogenase mutant of Arabidopsis demonstrates the key role of this enzyme in root carbon and nitrogen metabolism. Plant Cell 24:4044–4065. doi:10.1105/tpc.112.103689
Fontaine J-X, Tercé-Laforgue T, Bouton S, Pageau K, Lea PJ, Dubois F, Hirel B (2013) Further insights into the isoenzyme composition and activity of glutamate dehydrogenase in Arabidopsis thaliana. Plant Signal Behav 8:e23329. doi:10.4161/psb.23329
Grabarek Z (2006) Structural basis for diversity of the EF-hand calcium-binding proteins. J Mol Biol 359:509–525. doi:10.1016/j.jmb.2006.03.066
Grabowska A, Nowicki M, Kwinta J (2011) Glutamate dehydrogenase of the germinating triticale seeds: gene expression, activity distribution and kinetic characteristics. Acta Physiol Plant 33:1981–1990. doi:10.1007/s11738-011-0801-1
Grabowska A, Kwinta J, Bielawski W (2012) Glutamine synthetase and glutamate dehydrogenase in triticale seeds: molecular cloning and genes expression. Acta Physiol Plant 34:2393–2406. doi:10.1007/s11738-012-1085-9
Kwinta J, Kolik D (2006) Glutamine synthase and glutamate dehydrogenase in cadmium-stressed triticale seedlings. Acta Physiol Plant 28:339–347. doi:10.1007/s11738-006-0030-1
Kwinta J, Bartoszewicz K, Bielawski W (2001) Purification and characteristics of glutamate dehydrogenase (GDH) from triticale roots. Acta Physiol Plant 23:399–405. doi:10.1007/s11738-001-0049-2
Labboun S, Tercé-Laforgue T, Roscher A, Bedu M, Restivo FM, Velanis CN, Skopelitis DS, Moschou PN, Roubelakis-Angelakis KA, Suzuki A, Hirel B (2009) Resolving the role of plant glutamate dehydrogenase. I. In vivo real time nuclear magnetic resonance spectroscopy experiments. Plant Cell Physiol 50:1761–1773. doi:10.1093/pcp/pcp118
Laemmli UK (1970) Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature 227:680–685. doi:10.1038/227680a0
Lea PJ, Mifflin BJ (1974) An alternative route for nitrogen assimilation in plants. Nature 251:680–685. doi:10.1038/251614a0
Lehmann T, Ratajczak L (2008) The pivotal role of glutamate dehydrogenase (GDH) in the mobilization of N and C from storage material to asparagine in germinating seeds of yellow lupine. J Plant Physiol 165:149–158. doi:10.1016/j.jplph.2006.12.010
Lehmann T, Polcyn W, Ratajczak L (1990) Glutamate dehydrogenase isoenzymes in yellow lupine root nodules. III. Affinity for ammonia. Acta Physiol Plant 12:259–263
Lehmann T, Dabert M, Nowak W (2011) Organ-specific expression of glutamate dehydrogenase (GDH) subunits in yellow lupine. J Plant Physiol 168:1060–1066. doi:10.1016/j.jplph.2010.12.016
Loulakakis CA, Roubelakis-Angelakis KA (1990) Intracellular localization and properties of NADH-glutamate dehydrogenase from Vitis vinifera L.: purification and characterization of the major leaf isoenzyme. J Exp Bot 41:1223–1230. doi:10.1093/jxb/41.10.1223
Loulakakis KA, Roubelakis-Angelakis KA (1991) Plant NAD(H)-glutamate dehydrogenase consists of two subunit polypeptides and their participation in the seven isoenzymes occurs in an ordered ratio. Plant Physiol 97:104–111
Marchler-Bauer A, Zheng C, Chitsaz F, Derbyshire MK, Geer LY, Geer RC, Gonzales NR, Gwadz M, Hurwitz DI, Lanczycki CJ, Lu F, Lu S, Marchler GH, Song JS, Thanki N, Yamashita RA, Zhang D, Bryant SH (2013) CDD: conserved domains and protein three-dimensional structure. Nucleic Acids Res 41:D348–D352. doi:10.1093/nar/gks12
Masclaux-Daubresse C, Valadier M-H, Carrayol E, Reisdorf-Cren M, Hirel B (2002) Diurnal changes in the expression of glutamate dehydrogenase and nitrate reductase are involved in the C/N balance of tobacco source leaves. Plant Cell Environ 25:1451–1462. doi:10.1046/j.1365-3040.2002.00925.x
Melo-Oliveria R, Cinha-Oliveria I, Coruzzi GM (1996) Arabidopsis mutant analysis and gene regulation define a non-redundant role for glutamate dehydrogenase in nitrogen assimilation. Proc Natl Acad Sci USA 96:4718–4723. doi:10.1073/pnas.93.10.4718
Mol JNM, van der Krol AR, van Tunen R, van Blokland R, de Lange P, Stuitje AR (1990) Regulation of plant gene expression by antisense RNA. FEBS Lett 268:427–430. doi:10.1016/0014-5793(90)81298-3
Murashige T, Skoog F (1962) A revised medium for rapid growth and bio assays with tobacco tissue cultures. Physiol Plant 15:473–497. doi:10.1111/j.1399-3054.1962.tb08052.x
Nanda BB, Mall PC, Lodh SB (1991) Glutamate dehydrogenase activity and isoenzymes, protein, and soluble amino acids in developing grains of high and low protein rice. Cereal Chem 68:351–353
Oeller PW, Min-Wong L, Taylor LP, Pike DA, Theologis A (1991) Reversible inhibition of tomato fruit senescence by antisense RNA. Science 254:437–439. doi:10.1126/science.1925603
Paczek V, Dubois F, Sangwan R, Morot-Gaudry JF, Roubelakis-Angelakis KA, Hirel B (2002) Cellular and subcellular localisation of glutamine synthetase and glutamate dehydrogenase in grapes gives new insights on the regulation of carbon and nitrogen metabolism. Planta 216:245–254. doi:10.1007/s00425-002-0854-x
Purnell MP, Botella JR (2007) Tobacco isoenzyme 1 of NAD(H)-dependent glutamate dehydrogenase catabolizes glutamate in vivo. Plant Physiol 143:530–539. doi:10.1104/pp.106.091330
Purnell MP, Skopelitis DS, Roubelakis-Angelakis KA, Botella JR (2005) Modulation of higher-plant NAD(H)-dependent glutamate dehydrogenase activity in transgenic tobacco via alteration of beta subunit levels. Planta 222:167–180. doi:10.1007/s00425-005-1510-z
Qiu X, Xie W, Lian X, Zhang Q (2009) Molecular analyses of the rice glutamate dehydrogenase gene family and their response to nitrogen and phosphorous deprivation. Plant Cell Rep 28:1115–1126. doi:10.1007/s00299-009-0709-z
Ratajczak L, Koroniak D, Mazurowa H, Ratajczak W, Prus-Glowacki W (1986) Glutamate dehydrogenase isoforms in lupine roots and root nodules. Immunological studies. Physiol Plant 67:685–689. doi:10.1111/j.1399-3054.1986.tb05078.x
Restivo FM (2004) Molecular cloning of glutamate dehydrogenase genes of Nicotiana plumbaginifolia: structure and regulation of their expression by physiological and stress conditions. Plant Sci 166:971–982. doi:10.1016/j.plantsci.2003.12.011
Singh RP, Srivastava HS (1982) Glutamate dehydrogenase activity and assimilation of inorganic nitrogen in maize seedlings. Biochem Physiol Pflanzen 177:633–642. doi:10.1016/S0015-3796(82)80066-8
Skopelitis DS, Paranychianakis NV, Paschalidis KA, Pliakonis ED, Delis ID, Yakoumakis DI, Kouvarakis A, Papadakis AK, Stephanou EG, Roubelakis-Anfelakis KA (2006) Abiotic stress generates ROS that signal expression of anionic glutamate dehydrogenases to form glutamate for proline synthesis in tobacco and grapevine. Plant Cell 18:2767–2781. doi:10.1105/tpc.105.038323
Skopelitis DS, Paranychianakis NV, Kouvarakis A, Spyros A, Stephanou EG, Roubelakis-Angelakis KA (2007) The isoenzyme 7 of tobacco NAD(H)-dependent glutamate dehydrogenase exhibits high deaminating and low aminating activities in vivo. Plant Physiol 145:1726–1734. doi:10.1104/pp.107.107813
Temple SJ, Knight TJ, Unkefer PJ, Sengupta-Gopalan C (1993) Modulation of glutamine synthetase gene expression in tobacco by the introduction of an alfalfa glutamine synthetase gene in sense and antisense orientation: molecular and biochemical analysis. Mol Gen Genet 236:315–325. doi:10.1007/BF00277128
Tercé-Laforgue T, Bedu M, Dargel-Grafin C, Dubois F, Gibon Y, Restivo FM, Hirel B (2013) Resolving the role of plant glutamate dehydrogenase: II. Physiological characterization of plants overexpressing the two enzyme subunits individually or simultaneously. Plant Cell Physiol 54:1635–1647. doi:10.1093/pcp/pct108
Tercé-Laforgue T, Clément G, Marchi L, Restivo FM, Lea PJ, Hirel B (2015) Resolving the role of plant NAD-glutamate dehydrogenase: III. Overexpressing individually or simultaneously the two enzyme subunits under salt stress induces changes in the leaf metabolic profile and increases plant biomass production. Plant Cell Physiol 56:1918–1929. doi:10.1093/pcp/pcv114
Thompson JD, Gibson TJ, Plewniak F, Jeanmougin F, Higgins DG (1997) The CLUSTAL_X windows interface: flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Res 25:4876–4882. doi:10.1093/nar/25.24.4876
Turano FJ, Thakkar SS, Fang T, Weisemann JM (1997) Characterization and expression of NAD(H)-dependent glutamate dehydrogenase genes in Arabidopsis. Plant Physiol 113:1329–1341. doi:10.1104/pp.113.4.1329
Weigel D, Glazebrook J (2009) Quick miniprep for plant DNA isolation. Cold Spring Harb Protoc. doi:10.1101/pdb.prot5179
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Communicated by J. Gao.
Electronic supplementary material
Below is the link to the electronic supplementary material.
11738_2016_2322_MOESM1_ESM.pdf
Supplementary Figure. 1. The alignment of the deduced amino acid sequences of TsGDH2 and other plants GDH2s. The sequences of the following species were aligned: x Triticosecale (ALT22419.1), Brachypodium distachyon (XP_003580259.1), Oryza sativa (BAE48298.1), Nicotiana plumbaginifolia (CAA69601), Nicotiana sylvestris (XP_009758042.1), Vitis vinifera (XP_010662019.1), Arabidopsis thaliana (NP_196361.1). Supplementary Figure 2. The alignment of the nucleotide sequences of TsGDH1 and TsGDH2. Supplementary Figure 3. The alignment of the deduced amino acid sequences of TsGDH1 and TsGDH2 (PDF 77 kb)
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Grabowska, A., Zdunek-Zastocka, E., Kutryn, E. et al. Molecular cloning and functional analysis of the second gene encoding glutamate dehydrogenase in triticale. Acta Physiol Plant 39, 24 (2017). https://doi.org/10.1007/s11738-016-2322-4
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11738-016-2322-4