Abstract
Purpose
Ligulosis caused by Ligula intestinalis adversely affects the fisheries carried out in the lakes and ponds, causing economic losses in the fish industry. In this study, it was aimed to reveal the molecular characterization of L. intestinalis isolates obtained from woodfish (Acanthobrama marmid) in Keban Dam Lake in Elazig province of Turkey by using mt-CO1 gene sequences and to determine the genetic differences and haplotypes between the isolates.
Methods
In the examination made in terms of L. intestinalis, the intestine of the fish was opened with the help of fine-tipped scissors, the contents were allowed to come out, and the parasites were taken into a petri dish containing phosphate buffered saline (PBS). Then, L. intestinalis plerocercoids were taken into 15 ml falcon tubes containing 70% ethanol and stored at − 20 °C until further analysis. From each isolate, total gDNA was extracted from the plerocercoids. A partial (480 bp) mt-CO1 gene was amplified by PCR and sequenced unidirectionally. The final size of the trimmed sequences was 392 bp for 43 sequences. Sequence and haplotype analyses were performed, followed by phylogenetic analyses.
Results
All isolates were confirmed as L. intestinalis by BLAST analysis. In addition, 87 nucleotide mutation positions were determined among 43 CO1 gene sequences. As a result of the haplotype network performed for the mt-CO1 gene region of L. intestinalis isolates; arranged in a star-like configuration with the main haplotype (Hap05), separated from other haplotypes by 1–6 mutation steps, and 29 haplotypes were identified, covering 13.9% (6/43) of the total isolates. Also, 75 variable (polymorphic) sites were determined, 52 of which were parsimony informative sites.
Conclusions
The molecular characterization of L. intestinalis in woodfish (A. marmid) was identified for the first time in Turkey.
Similar content being viewed by others
Introduction
Seafood is a cheap and rich source of protein to cover animal protein needs, and it is an industry branch that represent a significant proportion of the world's nutritional needs [1]. Fish infection,, especially parasites, causes significant problems in aquatic ecosystems. Sometimes even a parasite species can threaten the existing reserves [2]. Ligula intestinalis (Linnaeus, 1758) ranks first as cestodes among the parasitic agents of fish. It has a significant prevalence in Turkey and creates a problem especially in dam lakes [3]. Ligulosis caused by Ligula intestinalis (Linnaeus, 1758) adversely affects the fisheries carried out in the lakes and ponds of in Turkey [4, 5].
Ligula intestinalis (Linnaeus, 1758), classified in the Diphyllobothriidae family, is a cestode reaching 28 cm as adults and 40 cm as plerocercoids [6, 7]. It is common in fresh waters in the Northern Hemisphere, including Turkey. Ligula intestinalis; has been reported in many fish families such as Cyprinidae, Catostomidae, Salmonidae or Galaxiidae. However, it shows a remarkable heterogeneity in host preference according to the geographical area studied [8]. Nowadays, the species in the Ligula taxon are classified as two genus (Ligula and Digramma) and four species (Ligula intestinalis (Linnaeus, 1758), Ligula colymbid (Zeder, 1803) and Digramma interrupta (Rudolphi, 1810), Digramma nemachili (Dubinina, 1957)) according to their morphological and anatomical features [9]. Ligula intestinalis is a pseudophyllidean cestode species scolex typically has dorsal and ventral bothriums. Sometimes bothriums are absent, and instead they have slits in the central part of the dorsal and ventral sides. The neck is quite short, almost absent. There are procercoid and plerocercoid stages in their development [10,11,12].
Ligula intestinalis has a three-host life cycle. There are two larval stages, procercoid and plerocercoid. The dominant phase of it life cycle is the plerocercoid phase, which lives in the body cavity of fish [10,11,12]. The first intermediate hosts are some copepod species. The larval form in these intermediate hosts are procercoids. The second intermediate hosts are freshwater fish, and their larval form is plerocercoids. Waterfowl such as seagulls, ducks, water divers and sea divers are the final hosts [13]. Freshwater fish become infected by eating parasitic copepods [3, 13, 14].
Plerocercoids of L. intestinalis most commonly develop in tench, red-eye, perch and pike perch. These plerocercoids are found in the abdominal cavity of fish. It causes swollen and the swimming slows down. In severe infections, the abdomen bursts and causes the death of the fish. Plerocercoids exert mechanical pressure on the reproductive organs and destroy the gonads. However, it is known that it adversely affects reproductive physiology by secreting antigonadotropic hormone. This effect causes infertility and pernicious anemia in fish [15]. In the fight against ligulosis, it is not practical and almost impossible to fight with copepod species belonging to the Cyclops and Diaptomus genus, which are the first intermediate host. In order to reduce the number of adult parasite-carrying and fish-eating waterfowl, measures should be taken periodically around the lake. Infected fish that move passively in the water should be collected and destroyed as much as possible [16,17,18].
Molecular techniques are used to identify species that cannot be distinguished morphologically according to their genetic differences [19]. Nuclear 28S rDNA and mitochondrial cytochrome c oxidase subunit 1 (mt-CO1) gene sequencing of L. intestinalis are used for molecular identification of the parasite [20]. In the case that both the existing fish species in aquatic ecosystems and the species that are grown intensively in net cages are exposed to this parasitic effect, molecular identification is important avoid economic depreciation and to take the necessary measures [18].
In this study, it was aimed to reveal the molecular characterization of L. intestinalis obtained from woodfish (Acanthobrama marmid) in Keban Dam Lake in Elazig province of Turkey by using mt-CO1 gene sequences and to determine the genetic differences and haplotypes between the isolates.
Materials and Methods
Study Area and Sampling
The fish to be examined for the study were obtained from Keban Dam Lake (Coordinates: 38°50′39″N 39°14′24″E) in Elazig province of Turkey between June and August, 2021. The fish were caught by fishing with gill nets number 22 in the lake. The fish, either alive or dead, were procured from the fishermen and brought to the Bingol University Faculty of Veterinary Fisheries Diseases Laboratory with ice in styrofoam boxes. After the external examination of the fish, the internal examination was carried out in accordance with the necropsy technique and examined for L. intestinalis plerocercoids. In the examination of L. intestinalis, the intestine of the fish was opened with the help of fine-tipped scissors, the contents were allowed to come out, and the parasites were taken into a petri dish containing physiological water. Then, L. intestinalis plerocercoids were taken into 15 ml falcon tubes containing 70% ethanol and stored at -20 °C until further analysis. The terminal segments were stained with aceto-carmine and mounted with Canada balsam. The specimens were identified as L. intestinalis by using taxonomic keys [21]. The morphological characterization of plerocercoid was completed by observing them under a light microscope.
Genomic DNA Isolation
Total genomic DNA (gDNA) was extracted using a Hibrigen Genomic DNA Isolation Kit (Hibrigen, Kocaeli-Turkey) according to the manufacturer’s instructions, with some slight modifications. The samples were washed five times using phosphate buffered saline (PBS) to remove the alcohol and then the plerocercoids was cut into small pieces with a scalpel, and transferred into Eppendorf tubes (1.5 ml). Two hundred microliters of lysis solution and 20 µl of Proteinase-K (20 mg/ml) were included in the tubes, mixed, then incubated overnight at 65 °C. To support the lysis, the samples were vortexed every 1 h for at least two to three times. The next day kit procedures were followed and total gDNA was isolated. The quantity of gDNA was (Nanodrop Technologies, Wilmington) measured and those samples containing insufficient gDNA were re-extracted to obtain a sufficient amount of gDNA. Isolated samples were stored at − 20 °C until further use.
Amplification of mt-CO1 Gene Fragment by PCR
To determine the genetic diversity of L. intestinalis isolates, the mt-CO1 gene region (480 bp) was amplified with the specific primers COIA2 (5’CATATGTTTTGATTTTTTGG3’) and COIB2 (5’AKAACATAATGAAAATGAGC3’) as previously described [22]. The PCR reaction comprised: 10 X PCR buffer (5 µl), MgCl2 (25 mM), dNTP’s (400 uM), from each primers (20 pmol), TaqDNA Polymerase (5 U/µl), PCR distilled water (28.8 µl) and gDNA (5 µl). The thermal cycling parameters of the PCR reaction were as follows: pre-denaturation for 5 min at 95 °C; 35 cycles of 45 s at 95 °C, 45 s at 57 °C, and 1 min at 72 °C, and extra extension step for 5 min at 72 °C. A Blue-Ray Biotech from (Taiwan, R.O.C) thermal cycler was used for PCR amplification and PCR products were separated using 1.4% agarose gel during 30 min. at 90 V constant current.
DNA Sequences
The PCR products of the mt-CO1 gene belonging to L. intestinalis isolates (n = 50) were sent to a commercial company (DNA Laboratory Systems, Istanbul, Turkey) for one-way Sanger DNA sequence analyses.
Data Analysis of the Sequences
FinchTV 1.4.0 was utilized to analyze the sequence data. BLAST search (www.ncbi.nlm.nih.gov/BLAST/) has been used to compare and trim our nucleotide sequences with published reference sequences. The MEGA X software was used to evaluate the trimmed sequences [23]. Using NCBI PubMed published reference sequences as outgroups, alignment of the sequences was performed. Ligula intestinalis (MZ359928, NC039445) was used as the reference sequence for phylogenetic tree construction while Taenia saginata (AY684274) and Echinococcus granulosus (NC044548) sequences were used as outgroup. The most suitable phylogenetic tree model for the sequences were identified as Tamura-Nei + Gamma distribution (+ G) by the MEGAX program. The phylogenetic tree was constructed using the Maximum Likelihood method in MEGA X, after finding the most accurate and suitable DNA model [23].
Nucleotide Diversity, Polymorphism and Neutrality Indices and Haplotype Networks of the Sequences
All sequence data were uploaded into DNASP6 software for more comprehensive evaluation as described by Rozas et al. [24]. The software was utilized to identify the diversity of neutrality and population indices. Population diversity indices included nucleotide diversity (πd), haplotype diversity (hd) and haplotype number (hn), while neutrality indices consisted of Tajima’s D [25], Fu’s statistics [26], Fu’s Fs [26], Fu and Li’s D test (FLD), Fu and Li's F (FLF) statistics test and parsimony informative analysis were performed using DnaSP6. Data in a number of output formats, such as NEXUS, was created using the DNASP6 (DNA Sequence Polymorphism) software program. To generate the haplotype network, the Minimum Spanning Networks (MSN) method was used in the PopART-1.7 software (Population Analysis with Reticulate Trees) [27, 28]. Haplotype analysis of the 43 sequences with a size of 392 bp belonging to the mt-CO1 gene fragment of L. intestinalis isolates was performed.
Results
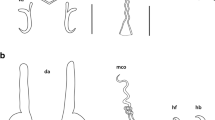
A total of 50 L. intestinalis plerocercoids were obtained from the abdominal cavity of fishes (A. marmid) (Fig. 1). To identify the exact species of the isolates, the mt-CO1 gene fragment (480 bp) was amplified using PCR. After sequence analysis, nucleotide sequences were compared with published reference sequences. The final size of the trimmed sequences was 392 bp and 7 sequences were excluded because of the low quality sequence reads although twice attempts. Totally 43 sequences of mt-CO1 samples were submitted to GenBank.
Following the alignment, point mutations in the sequences were detected. Nucleotide variation positions based on the reference sequence are demonstrated in the Table 1. A phylogenetic tree was generated with the bootstrap test (1000 replicates) by applying the Maximum Likelihood statistical method and is shown in Fig. 2.
Phylogenetic tree view created using reference sequences from L. intestinalis (MZ359928, NC039445) and outgroup sequence Taenia saginata (AY684274), Echinococcus granulosus (NC044548) with 392 bp mt-CO1 gene sequences (TRLig001-TRLig43) of L. intestinalis isolates. The tree was created based on the Tamura-Nei + Gamma distribution (+ G) (TN93 + G) model with the Maximum Likelihood method in MEGA-X, and the reliability of the tree was ensured by 1,000 bootstrap tests
As a result of the haplotype network performed for the mt-CO1 gene region of L. intestinalis isolates; arranged in a star-like configuration with the main haplotype (Hap05), separated from other haplotypes by 1–6 mutation steps, and 29 haplotypes were detected, covering 13.9% (6/43) of the total isolates. (Fig. 3; Table 2). As a result of haplotype analysis, 75 variable (polymorphic) sites were determined, 52 of which were parsimony informative sites. Low nucleotide diversity and high haplotype variation were reported for the mt-CO1 gene fragment (Table 3). Tajima’s D value was demonstrated population expansion and/or refinement of selection. Uncommon haplotypes from recent population expansion or hitchhiking were expected because of the significant negative Fu’s Fs value. Therefore, the fact that 79.3% (23/29) of the haplotype groups were single haplotypes supported these results. The sequences identity among the published reference sequence (NC_039445) and the current sequences (OM760000-OM760042) for L. intestinalis are presented in Table 2. After a BLAST search, several novel L. intestinalis haplotypes arose. The Hap02 and Hap29 were matched 100% with the published sequences in the Genbank. Although, Hap06 and Hap21 had 99.74% similarity rate, Hap13 had the lowest similarity with 90.67%. Finally, 27 haplotypes were determined as unique haplotypes in the current study (Table 4).
Discussion
One of the health problems encountered in fish farming is parasitic diseases. In recent years, it has been reported that parasitic diseases are quite common in freshwater fish farming in our country. The diseases not only have harmful effects such as growth retardation and reproductive problems in fish, but also lead to death when the parasite population is high [29]. The recognition of parasitic fish diseases and the investigation of their treatment are of great importance for the developing fishery sector today. Parasites are encountered intensively when the fish density is high, malnutrition and environmental conditions change. Weakness and intense parasitic invasions developing with increasing stress factors can be fatal for the fish [17, 30, 31].
In recent studies in Turkey, parasitic diseases are frequently encountered in fish species belonging to the Cyprinidae family found in natural waters [32,33,34,35,36]. Turkey has a rich location in terms of both its geographical location and water resources (lake, river, sea, etc.) and fish population. Thus, parasitic infections encountered in fish facilities and fisheries are very important for Turkey's economy. In our study, the presence of L. intestinalis infection in the A. marmid species was investigated and the internal examinations of the fish were examined by necropsy technique and 50 L. intestinalis plerocercoids were obtained from the abdominal cavities of the fishes. In this study, detection of L. intestinalis parasite in A. marmid was in agreement with the studies in terms of detection of the parasite in freshwater fish. In some studies, the changing in L. intestinalis prevalence depend on the seasons, age groups and sex characteristics of the fish were investigated [34, 37,38,39]. Moreover, the effect of the parasite on the fish condition factor and physiological-anatomical structures was also investigated [40,41,42,43,44,45,46].
Today, PCR-based DNA sequence analysis and other molecular methods are frequently preferred for the identification of parasite agents and their phylogenetic characterization [8]. Studies on L. intestinalis species are mostly anatomical-morphologically based in Turkey [47], and the molecular studies are limited [48]. However, genetic markers such as ITS and CO1 have been frequently used in recent years to accurately determine the taxonomic positions of organisms [20]. The traditional classification of ligulid cestodes Ligula (Bloch, 1782) and Digramma (Cholodkovsky, 1914) is a controversial issue. Especially, the low nucleotide diversity between the genus Ligula and Digramma indicates that Digramma is probably not an independent genus. For this reason, it showed that Ligula and Digramma should be considered as two species within the genus Ligula. As a result of the studies, it has been reported that ITS1, ITS2 and ND1 sequences are useful genetic markers to distinguish between Ligula and Digramma [49,50,51,52]. Lagrue et al. [52] reported a case of Ligula sp. in fish species Gobiomorphus cotidianus and Oncorhynchus tshawytscha from Lake Hawea, South Island of New Zealand. Parasites have been quite recorded in large sizes (60–300 mm). In addition, low prevalences in fish populations suggested that the infection was rare or local. The ITS1 and ITS2 sequences confirmed that these isolates belong to the Ligula genus [14]. In present study, sequence of the mt-CO1 gene fragment was preferred to determine the genetic diversity and phylogenetic characterization of L. intestinalis isolates collected from A. marmid fish species belonging to the Cyprinidae family. As a result all sequences were match with L. intestinalis in the current study.
Several studies have been carried out on the molecular identification of L. intestinalis infection, which causes serious yield and economic losses in fish facilities. Li et al. [20], certain nucleotides of 393 bp for the CO1 gene of seven L. intestinalis samples were directly sequenced from each sample. Interestingly, no nucleotide variation was detected in the sequences of the CO1 gene among the seven Ligula samples studied. The authors reported that this homology in sequences may be due to the mutation patterns of the mitochondrial genome and the sequence similarity of CO1 genes of closely related taxon. In another study, a single pattern and reliable band were recorded in all gene sequences of 6 identified L. intestinalis isolates. The size of the bands were determined as 480 bp. No nucleotide variation was detected in any of the CO1 gene products of the samples. Only one mutation was detected at nucleotide 225 of one sample [14]. In support of the data of the above researchers, in this study similar Ligula intestinalis specific primers were used. A single pattern and a reliable band were recorded in all of the gene sequences of the 43 L. intestinalis isolates. The bands were 480 bp in size. All isolates were confirmed as L. intestinalis by BLAST analysis. In addition, 87 nucleotide mutation positions were determined among 43 CO1 gene sequences. This suggests that L. intestinalis plerocercoids in A. marmid fish show higher nucleotide diversity compared to other fish species.
Bouzid et al. [22], a study was conducted to determine the phylogeny and genetic structure analysis of L. intestinalis according to geography and fish host preference. The analyzed data consisted of 109 parasites from 13 host fish species in 18 different locations on a macrogeographic scale. The two mitochondrial genes CO1 and cytb were preferred to determine the population genetic structure of L. intestinalis on a local and global scale. Besides, the nuclear sequence of the ITS2 gene was used for genetic reconstruction. Different evolutionary diversity was determined at different locations. Although the ITS2 sequences were found to show significant intragenomic variability, their associations were generally stated to be in good agreement with the topology derived from mitochondrial genes. In another study, the genetic diversity of L. intestinalis was analyzed using ISSR markers. In the study, ten host species from nine different locations and nine ISSR markers were used to analyze the genetic diversity of L. intestinalis populations. The 110 loci selected from the ISSR patterns produced revealed high variability among the analyzed samples, with a polymorphism of 100% and a global coefficient of gene differentiation estimated by Nei’s index (GST) of 0.776. Major genetic variation was associated with five different geographic locations (Algeria, Canada, Australia, China and Europe). However, no remarkable genetic variation was detected, even though samples in Europe were obtained from different geographic regions and different hosts. In conclusion, the ISSR approach has been shown to be quick and cheap and provides reliable indicators to evaluate the genetic diversity of L. intestinalis [8]. In present study, we determined 29 haplotypes among the 43 L. intestinalis isolates. We observed 87 multiple nucleotide mutation positions within 29 haplotypes. By analysis of the sequences, the most common haplotypes were Hap05 and Hap12, respectively. Identification of 29 haplotypes in 43 isolates revealed the important genetic diversity of L. intestinalis By haplotype analysis, low nucleotide variation and high haplotype diversity were recorded for the mt-CO1 partial gene sequences of L. intestinalis. Tajima’s D value indicated expansion of the population and the influence of purifying selection. The negative value of Fu’s Fs indicated that the presence of the rare haplotypes occurred because of hitchhiking or recent population enlargement. Therefore, it was not surprising that there were 29 haplotypes out of 43 sequences. Besides, 79.3% (23/29) of the haplotype consisted of a single sequence.
Conclusion
This study was conducted by taking into account that it is necessary the molecular characterization of L. intestinalis, which causes serious economic losses in fishing. Molecular characterization studies of L. intestinalis are limited in Turkey. As a result of this study, the haplotype analysis of L. intestinalis was revealed for the first time. Hence, the molecular characterization of L. intestinalis in woodfish (A. marmid) was identified for the first time. The data of this study will also contribute to the epidemiology, control and treatment options of L. intestinalis infection, which is one of the most serious economic problems of the country's fishing. With this study, will open the way for more original and comprehensive research on molecular identification of parasitic disease agents in all regions of our country in the future.
References
Yuksel F, Kuzgun NK, Ozer EI (2011) Determination of the fish consumption habits of Tunceli province. Karadeniz fen bilim derg 2:28–36
Dorucu M, Mutlu N (2008) Parasitic fish disease and their treatment. Ecol Life Sci 3:372–380
Erer H (2002) Balık hastalıkları. Selçuk Üniversitesi Basımevi, Konya.
Arslan MO, Yilmaz M, Tasci GT (2015) Infections of Ligula intestinalis on freshwater fish in Kars plateau of north-eastern Anatolia. Turkey Turkiye Parazitol Derg 39:218. https://doi.org/10.5152/tpd.2015.4168
Doosti S, Yılmaz F (2020) Occurrence of Ligula sp plerocercoids in Ladigesocypris irideus (Ladiges) from South-Western Turkey: new host and new locality records. J Inst Sci Technol 10:2416–2423. https://doi.org/10.21597/jist.688296
Ciloglu A, Biskin Z (2013) Balıklarda Görülen Parazit Hastalıkları: Respirator Sistem Deri ve Bağ Dokularda Cestod ve Nematod Hastalıkları. In: Ozcel MA (ed) Veteriner Hekimliğinde Parazit Hastalıkları. Turkiye Parazitol Dern Yay, Izmir
Hajirostamloo M (2008) The occurrence and parasite-host of Ligula intestinalis in Sattarkhan Lake (East Azerbaijan-Iran). J Anim Vet Adv 7(3):221–225
Bouzid W, Lek S, Mace M, Ben Hassine O, Etienne R, Legal L, Loot G (2008) Genetic diversity of Ligula intestinalis (Cestoda:Diphyllobothriidea) based on analysis of inter-simple sequence repeat markers. J Zool Syst Evol Res 46(4):289–296. https://doi.org/10.1111/j.1439-0469.2008.00471.x
Dubinina MN (1980) Tapeworms (Cestoda, Ligulidae) of the fauna of the USSR. Tapeworms (Cestoda, Ligulidae) of the fauna of the USSR.
Guralp N (1974) Helmintoloji. Ankara Universitesi Veteriner Fakultesi Yayınları, Ankara
Hoffman GI (1967) Parasites of North American Freshwater Fishes. University of California Pres, Berkeley and Los Angeles
Tolgay N (1973) Evcil ve yabani kanatlıların önemli parazitleri. A U Vet Fak Yayın No 249 Ankara
Olson P, Littlewood D, Griffiths D, Kennedy C, Arme C (2002) Evidence for the co-existence of separate strains or species of Ligula in Lough Neagh. Northern Ireland J Helminthol 76:171–174. https://doi.org/10.1079/JOH2002101
Islam E (2019) DNA dizisi tabanlı olarak Ligula intestinalis L.’ in moleküler tanımlaması. PhD Dissertation, Afyon Kocatepe Üniversitesi, Türkiye.
Ozcel MA, Inci A, Koroglu E, Karaer Z, Eren H (2016) Veteriner hekimliğinde parazit hastalıkları. Türkiye Parazitoloji Dernegi, İzmir Türkiye
Akkaya H, Turkmen H, Vuruşaner C (1998) Balık parazitleri. Teknik yayınları, İstanbul.
Arda M, Seçer S, Sarıeyyupoglu M (2005) Balık Hastalıkları. Medisan Yayın Serisi, Ankara.
Ozcan M, Kartal B (2020) Molecular identification of the parasite Ligula intestinalis identified in Alburnus adanensis. Asian J Sci Techn 11:10700–10707
Aksakal E, Erdoğan O (2007) PCR-RFLP Uygulamalarının Su Ürünlerinde Kullanım Alanları. Türk Sucul Yaşam Dergisi 5:747–753
Li J, Liao X, Yang H (2000) Molecular characterization of a parasitic tapeworm (Ligula) based on DNA sequences from formalin-fixed specimens. Biochem Genet 38:309–322. https://doi.org/10.1023/A:1002009117626
Chubb J, Pool D, Veltkamp C (1987) A key to the species of cestodes (tapeworms) parasitic in British and Irish freshwater fishes. J Fish Biol 31:517–543. https://doi.org/10.1111/j.1095-8649.1987.tb05256.x
Kumar S, Stecher G, Li M, Knyaz C, Tamura K (2018) MEGA X: molecular evolutionary genetics analysis across computing platforms. Mol Biol Evol 35:1547. https://doi.org/10.1093/molbev/msy096
Rozas J, Ferrer-Mata A, Sánchez-DelBarrio JC, Guirao-Rico S, Librado P, Ramos-Onsins SE, Sánchez-Gracia A (2017) DnaSP 6: DNA sequence polymorphism analysis of large data sets. Mol Biol Evol 34:3299–3302. https://doi.org/10.1093/molbev/msx248
Tajima F (1989) Statistical method for testing the neutral mutation hypothesis by DNA polymorphism. Genetics 123:585–595. https://doi.org/10.1093/genetics/123.3.585
Fu Y-X (1997) Statistical tests of neutrality of mutations against population growth, hitchhiking and background selection. Genetics 147:915–925. https://doi.org/10.1093/genetics/147.2.915
Bandelt H-J, Forster P, Röhl A (1999) Median-joining networks for inferring intraspecific phylogenies. Mol Biol Evol 16:37–48. https://doi.org/10.1093/oxfordjournals.molbev.a026036
Leigh JW, Bryant D (2015) POPART: full-feature software for haplotype network construction. Methods Ecol Evol 6:1110–1116. https://doi.org/10.1111/2041-210X.12410
Ozcan M, Yılmaz Y, Donat E, Kılavuz D, Tuncel M (2019) A research on endoparasitic fauna in fish species caught in Menzelet Dam Lake Kahramanmaraş (Turkey). Middle East J Sci 5:33–40. https://doi.org/10.23884/mejs.2019.5.1.04
Cetin ET, Ang O, Toreci K (1983) Tıbbi Parazitoloji. İstanbul Tıp Fakültesi Yayınları, İstanbul.
Ekingen G (1983) Tatlı Su Balık Parazitleri. Fırat Üniversitesi. Su Ürünleri Yüksek Okulu FÜ Basımevi, Elâzığ.
Barata S, Dorucu M (2014) Karakaya baraj gölü Kömürhan Bölgesinden yakalanan bazı balıklarda endohelmintlerin araştırılması. Master tezi, F.U. Fen Bilimleri Enstitüsü.
Gul A, Ispir U, Turk C, Kırıcı M, Taysı M, Yonar M (2014) The investigation of Diplostomum sp. metacercariae in some Cyprinids from Murat River (Genç Area), Bingöl, Turkey. Turkish J Agricultura Nat Sci 1:547–551
Ozbek M, Ozturk MO (2010) Investigations on Ligula intestinalis plerocercoid L., 1758 infection of some fishes from dam lake kunduzlar (Kırka, Eskişehir). Turkiye Parazitol Derg 34:112–117
Ozturk M (2012) A research on argulus foliaceus infection of three cyprinid fish species in Lake Dam Emre, Afyonkarahisar. J Fac Vet Med Istanbul Univ 38:183–190
Vilizzi L, Tarkan AS, Ekmekci FG (2015) Parasites of the common carp Cyprinus carpio L., 1758 (Teleostei: Cyprinidae) from water bodies of Turkey: updated checklist and review for the 1964–2014 period. Turk J Zool 39:545–554. https://doi.org/10.3906/zoo-1401-42
Kır I, Ayvaz Y, Barlas M, Ozan ST (2004) Seasonal distribution and effect of parasites on carp (Cyprinus carpio L., 1758) inhabiting the Karacaoren I dam lake. Turkiye Parazitol Derg 28:45–49
Kurupınar E, Ozturk MO (2009) A study on the helminth fauna linked to seasonal changes and size of the fish host, Leuciscus cephalus L., from Lake Dam Örenler. Afyonkarahisar Turkiye Parazitol Derg 33:248–253
Demirtas M, Altındag A (2011) The seasonal infection distribution of Ligula intestinalis plerocercoid L., 1758 on some fishes (Cyprinidae) LIVING in Terkos Lake. YYU Vet Fak Derg 22:147–151
Harris MT, Wheeler A (1974) Ligula infestation of bleak Alburnus alburnus (L.) in the tidal Thames. J Fish Biol 6:181–188. https://doi.org/10.1111/j.1095-8649.1974.tb04535.x
Uzbilek MK, Yıldız HY (2002) A report on spontaneous diseases in the culture of grass carp (Ctenopharyngodon idella Val. 1844). Turk Turk J Vet Anim Sci 26:407–410
Ergonul M, Altındag A (2005) Ligula intestinalis pleurocercoidlerinin kadife balığının büyüme özelliklerine etkisi. Turk J Vet Anim Sci 29:1337–1341
Museth J (2001) Effects of Ligula intestinalis on habitat use, predation risk and catchability in European minnows. J Fish Biol 59:1070–1080. https://doi.org/10.1111/j.1095-8649.2001.tb00172.x
Innal D, Keskin N (2006) The infection of European Chub Leuciscus cephalus L 1758 with Ligula intestinalis plecercoids in Çamkoru Lake Turkey. J Anim Vet Adv 5:108–110
Akmirza A (2007) The effect of Ligula intestinalis L. plerocercoid on the growth of bitterling (Rhodeus amarus Bloch, 1782). J Black Sea/Mediterranean Environ 13:155–160
Tekin-Ozan S, Barlas M (2008) Concentrations of selected heavy metals in Ligula intestinalis L., 1758 plerocercoids (Cestoda) compared to it host’s (Tinca tinca L., 1758) organs from Beyşehir Lake (Turkey). Helminthologia 45:76–80. https://doi.org/10.2478/s11687-008-0014-3
Innal D, Keskin N, Erk’akan F (2007) Distribution of Ligula intestinalis (L.) in Turkey. Turk J Fish Aquat Sci 7:19–22
Logan FJ, Horák A, Štefka J, Aydogdu A, Scholz T (2004) The phylogeny of diphyllobothriid tapeworms (Cestoda: Pseudophyllidea) based on ITS-2 rDNA sequences. Parasitol Res 94:10–15. https://doi.org/10.1007/s00436-004-1164-y
Ahmadiara E, Hosseini SH, Jalousian F, Mousavi HAE (2018) The taxonomic position and phylogenetic relationship between Digramma interrupta and Ligula intestinalis based on morphological and molecular diagnosis. Iran J Parasitol 13:648
Li WX, Fu PP, Zhang D, Boyce K, Xi BW, Zou H, Li M, Wu SG, Wang GT (2018) Comparative mitogenomics supports synonymy of the genera Ligula and Digramma (Cestoda: Diphyllobothriidae). Parasit Vectors 11:324. https://doi.org/10.1186/s13071-018-2910-9
Li J, Liao X (2003) The taxonomic status of Digramma (Pseudophyllidea: Ligulidae) inferred from DNA sequences. J Parasitol 89:792–799. https://doi.org/10.1645/GE-3078
Luo H, Nie P, Yao W, Wang G, Gao Q (2003) Is the genus Digramma synonymous to the genus Ligula (Cestoda: Pseudophyllidea)? Parasitol Res 89:419–421. https://doi.org/10.1007/s00436-002-0802-5
Lagrue C, Presswell B, Dunckley N, Poulin R (2018) The invasive cestode parasite Ligula from salmonids and bullies on the South Island, New Zealand. Parasitol Res 117:151–156. https://doi.org/10.1007/s00436-017-5684-7
Funding
Open access funding provided by the Scientific and Technological Research Council of Türkiye (TÜBİTAK). This study has been supported by a grant from the Bingol University Scientific Research Projects Unit (Project No: BAP-VF.2021.00.002).
Author information
Authors and Affiliations
Contributions
HKK, FC, SGK and SS contributed to the conception and design of the study. HKK, FC, CT, SGK and SS collected the samples, performed the laboratory activities and organized the database. HKK and SS wrote the manuscript. HKK, SS and AG critically oversaw the substantial revisions of the manuscript.
Corresponding author
Ethics declarations
Conflict of Interest
All authors have read and agreed to the published version of the manuscript. The authors declare that they have no conflicts of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Kesik, H.K., Celik, F., Turk, C. et al. Sequence and Haplotype Analyses of Ligula intestinalis in Acanthobrama marmid (Cyprinidae) in Turkey. Acta Parasit. 69, 453–464 (2024). https://doi.org/10.1007/s11686-023-00762-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11686-023-00762-2