Abstract
Objective
Runt-related transcription factor 1 (RUNX1) has been proven to be over-expressed and vital in many malignancies. However, its role in cervical cancer is still unclear.
Methods
Some online databases (Oncomine, GEPIA, UALCAN, LinkedOmics, and others) were used to explore the expression level, prognostic significance, and gene mutation characteristics of RUNX1 in cervical cancer. The protein levels of RUNX1 in cervical cancer were measured by immunohistochemistry (IHC). The functional changes of cervical cancer cells were measured in vitro after decreasing RUNX1.
Results
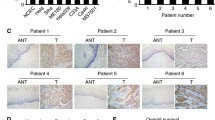
Bioinformatic results revealed that RUNX1 was upregulated in cervical cancer compared to normal tissues. Moreover, over-expression of RUNX1 was significantly correlated with cervical cancer patients’ clinical parameters (e.g., individual cancer stages, patients’ age, nodal metastasis status, and others). Meanwhile, functional enrichment analysis of RUNX1-related genes indicated that RUNX1 was mainly involved in the epithelial-mesenchymal transition (EMT) process in cervical cancer. Furthermore, RUNX1 may be upregulated by hsamiR-616-5p and hsa-miR-766 identified by miRDB, TargetScan, and miRWalk. Finally, RUNX1 was upregulated in cervical cancer compared to normal tissues by IHC in collected cervical cancer samples. The invasion and migration abilities of cervical cancer cells were significantly reduced by repressing EMT after knocking down RUNX1 in vitro.
Conclusion
RUNX1 was highly expressed in cervical cancer, and upregulated RUNX1 could significantly promote the invasive abilities of cervical cancer cells by inducing EMT. Therefore, RUNX1 may be a potential biomarker for early diagnosis and targeted therapy of cervical cancer.
Similar content being viewed by others
References
Hull R, Mbele M, Makhafola T, et al. Cervical cancer in low and middle-income countries. Oncol Lett, 2020,20(3):2058–2074
Bray F, Ferlay J, Soerjomataram I, et al. Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin, 2018,68(6):394–424
Bao HL, Liu YN, Wang LJ, et al. Analysis on mortality of cervical cancer and its temporal trend in women in China, 2006–2012. Zhonghua Liu Xing Bing Xue Za Zhi (Chinese), 2017,38(1):58–64
Lei J, Ploner A, Elfström KM, et al. HPV Vaccination and the Risk of Invasive Cervical Cancer. N Engl J Med, 2020,383(14):1340–1348
Buskwofie A, David-West G, Clare CA. A Review of Cervical Cancer: Incidence and Disparities. J Natl Med Assoc, 2020,112(2):229–232
Chopra S, Gupta M, Mathew A, et al. Locally advanced cervical cancer: A study of 5-year outcomes. Indian J Cancer, 2018,55(1):45–49
Salunkhe R, Chopra S, Kulkarni S, et al. Outcomes of locally advanced cervical cancer presenting with obstructive uropathy: An institutional audit. Indian J Cancer, 2020,57(4):416–422
Kumar L, Harish P, Malik PS, et al. Chemotherapy and targeted therapy in the management of cervical cancer. Curr Probl Cancer, 2018,42(2):120–128
Gadducci A, Cosio S. Neoadjuvant Chemotherapy in Locally Advanced Cervical Cancer: Review of the Literature and Perspectives of Clinical Research. Anticancer Res, 2020,40(9):4819–4828
Cibula D, Pötter R, Planchamp F, et al. The European Society of Gynaecological Oncology/European Society for Radiotherapy and Oncology/European Society of Pathology guidelines for the management of patients with cervical cancer. Radiother Oncol, 2018,127(3):404–416
Westerveld H, Nesvacil N, Fokdal L, et al. Definitive radiotherapy with image-guided adaptive brachytherapy for primary vaginal cancer. Lancet Oncol, 2020,21(3):e157–e167
Feng CH, Mell LK, Sharabi AB, et al. Immunotherapy With Radiotherapy and Chemoradiotherapy for Cervical Cancer. Semin Radiat Oncol, 2020,30(4):273–280
Chung HC, Ros W, Delord JP, et al. Efficacy and Safety of Pembrolizumab in Previously Treated Advanced Cervical Cancer: Results From the Phase II KEYNOTE-158 Study. J Clin Oncol, 2019,37(17):1470–1478
Stevanović S, Draper LM, Langhan MM, et al. Complete regression of metastatic cervical cancer after treatment with human papillomavirus-targeted tumor-infiltrating T cells. J Clin Oncol, 2015,33(14):1543–1550
Frenel JS, Le Tourneau C, O’Neil B, et al. Safety and Efficacy of Pembrolizumab in Advanced, Programmed Death Ligand 1-Positive Cervical Cancer: Results From the Phase Ib KEYNOTE-028 Trial. J Clin Oncol, 2017,35(36):4035–4041
Simon L, Spinella JF, Yao CY, et al. High frequency of germline RUNX1 mutations in patients with RUNX1-mutated AML. Blood, 2020,135(21):1882–1886
Choi A, Illendula A, Pulikkan JA, et al. RUNX1 is required for oncogenic Myb and Myc enhancer activity in T-cell acute lymphoblastic leukemia. Blood, 2017,130(15):1722–1733
Tang CY, Wu M, Zhao D, et al. Runx1 is a central regulator of osteogenesis for bone homeostasis by orchestrating BMP and WNT signaling pathways. PLoS Genet, 2021,17(1):e1009233
Tang CY, Chen W, Luo Y, et al. Runx1 up-regulates chondrocyte to osteoblast lineage commitment and promotes bone formation by enhancing both chond-rogenesis and osteogenesis. Biochem J, 2020,477(13): 2421–2438
Li Q, Lai Q, He C, et al. RUNX1 promotes tumour metastasis by activating the Wnt/β-catenin signalling pathway and EMT in colorectal cancer. J Exp Clin Cancer Res, 2019,38(1):334
Zhou T, Luo M, Cai W, et al. Runt-Related Transcription Factor 1 (RUNX1) Promotes TGF-β-Induced Renal Tubular Epithelial-to-Mesenchymal Transition (EMT) and Renal Fibrosis through the PI3K Subunit p110δ. EBioMedicine, 2018,31:217–225
Rhodes DR, Yu J, Shanker K, et al. ONCOMINE: a cancer microarray database and integrated data-mining platform. Neoplasia, 2004,6(1):1–6
Tang Z, Li C, Kang B, et al. GEPIA: a web server for cancer and normal gene expression profiling and interactive analyses. Nucleic Acids Res, 2017,45(W1): W98–W102
Chandrashekar DS, Bashel B, Balasubramanya SAH, et al. UALCAN: A Portal for Facilitating Tumor Subgroup Gene Expression and Survival Analyses. Neoplasia, 2017,19(8):649–658
Liu CJ, Hu FF, Xia MX, et al. GSCALite: a web server for gene set cancer analysis. Bioinformatics, 2018,34(21):3771–3772
Pathan M, Keerthikumar S, Ang CS, et al. FunRich: An open access standalone functional enrichment and interaction network analysis tool. Proteomics, 2015,15(15):2597–2601
Vasaikar SV, Straub P, Wang J, et al. LinkedOmics: analyzing multi-omics data within and across 32 cancer types. Nucleic Acids Res, 2017,46(D1):D956–D963
Hänzelmann S, Castelo R, Guinney J. GSVA: gene set variation analysis for microarray and RNA-seq data. BMC Bioinformatics, 2013,14:7
Yoshihara K, Shahmoradgoli M, Martínez E, et al. Inferring tumour purity and stromal and immune cell admixture from expression data. Nat Commun, 2013,4:2612
Sticht C, De La Torre C, Parveen A, et al. miRWalk: An online resource for prediction of microRNA binding sites. PLoS One, 2018,13(10):e0206239
Du A, Zhao S, Wan L, et al. MicroRNA expression profile of human periodontal ligament cells under the influence of Porphyromonas gingivalis LPS. J Cell Mol Med, 2016,20(7):1329–1338
Liu W, Wang X. Prediction of functional microRNA targets by integrative modeling of microRNA binding and target expression data. Genome Biol, 2019,20(1):18
Agarwal V, Bell GW, Nam JW, et al. Predicting effective microRNA target sites in mammalian mRNAs. Elife, 2015,4:e05005
Kagabu M, Nagasawa T, Sato C, et al. Immunotherapy for Uterine Cervical Cancer Using Checkpoint Inhibitors: Future Directions. Int J Mol Sci, 2020,21(7):2335
Ventriglia J, Paciolla I, Pisano C, et al. Immunotherapy in ovarian, endometrial and cervical cancer: State of the art and future perspectives. Cancer Treat Rev, 2017,59:109–116
Ferrall L, Lin KY, Roden RBS, et al. Cervical Cancer Immunotherapy: Facts and Hopes. Clin Cancer Res, 2021,27(18):4953–4973
Wang L, Brugge JS, Janes KA. Intersection of FOXO- and RUNX1-mediated gene expression programs in single breast epithelial cells during morphogenesis and tumor progression. Proc Natl Acad Sci USA, 2011,108(40):E803–812
Takayama K, Suzuki T, Tsutsumi S, et al. RUNX1, an androgen- and EZH2-regulated gene, has differential roles in AR-dependent and -independent prostate cancer. Oncotarget, 2015,6(4):2263–2276
Liu C, Xu D, Xue B, et al. Upregulation of RUNX1 Suppresses Proliferation and Migration through Repressing VEGFA Expression in Hepatocellular Carcinoma. Pathol Oncol Res, 2020,26(2):1301–1311
Miyagawa K, Sakakura C, Nakashima S, et al. Down-regulation of RUNX1, RUNX3 and CBFbeta in hepatocellular carcinomas in an early stage of hepatocarcinogenesis. Anticancer Res, 2006,26(5b): 3633–3643
Alonso-Alconada L, Muinelo-Romay L, Madissoo K, et al. Molecular profiling of circulating tumor cells links plasticity to the metastatic process in endometrial cancer. Mol Cancer, 2014,13:223
Lamouille S, Xu J, Derynck R. Molecular mechanisms of epithelial-mesenchymal transition. Nat Rev Mol Cell Biol, 2014,15(3):178–196
Nieto MA, Huang RY, Jackson RA, et al. EMT: 2016. Cell, 2016,166(1):21–45
Kavanagh DP, Robinson J, Kalia N. Mesenchymal stem cell priming: fine-tuning adhesion and function. Stem Cell Rev Rep, 2014,10(4):587–599
Jolly MK, Ware KE, Gilja S, et al. EMT and MET: necessary or permissive for metastasis? Mol Oncol, 2017,11(7):755–769
Pastushenko I, Blanpain C. EMT Transition States during Tumor Progression and Metastasis. Trends Cell Biol, 2019,29(3):212–226
Gonzalez-Avila G, Sommer B, Garcia-Hernandez AA, et al. Matrix Metalloproteinases’ Role in Tumor Microenvironment. Adv Exp Med Biol, 2020,1245:97–131
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
The authors declare that there is no conflict of interest regarding the publication of this paper.
Supplementary data
Rights and permissions
About this article
Cite this article
Zheng, Ll., Cai, L., Zhang, Xq. et al. Dysregulated RUNX1 Predicts Poor Prognosis by Mediating Epithelialmesenchymal Transition in Cervical Cancer. CURR MED SCI 42, 1285–1296 (2022). https://doi.org/10.1007/s11596-022-2661-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11596-022-2661-x