Abstract
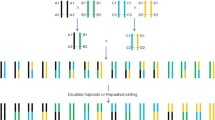
Most important agronomic and quality traits of crops are quantitative in nature. The genetic variations in such traits are usually controlled by sets of genes called quantitative trait loci (QTLs), and the interactions between QTLs and the environment. It is crucial to understand the genetic architecture of complex traits to design efficient strategies for plant breeding. In the present study, a new experimental design and the corresponding statistical method are presented for QTL mapping. The proposed mapping population is composed of double backcross populations derived from backcrossing both homozygous parents to DH (double haploid) or RI (recombinant inbreeding) lines separately. Such an immortal mapping population allows for across-environment replications, and can be used to estimate dominance effects, epistatic effects, and QTL-environment interactions, remedying the drawbacks of a single backcross population. In this method, the mixed linear model approach is used to estimate the positions of QTLs and their various effects including the QTL additive, dominance, and epistatic effects, and QTL-environment interaction effects (QE). Monte Carlo simulations were conducted to investigate the performance of the proposed method and to assess the accuracy and efficiency of its estimations. The results showed that the proposed method could estimate the positions and the genetic effects of QTLs with high efficiency.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
References
Lander E S, Green P, Abrahamson J, et al. MAPMAKER: An interactive computer package for constructing primary genetic linkage maps of experimental and natural populations. Genomics, 1987; 1: 174–181
Wang D L, Zhu J, Li Z K, et al. Mapping QTLs with epistatic effects and QTLxenvironment interactions by mixed linear model approaches. Theor Appl Genet, 1999; 99: 1255–1264
Yan J Q, Zhu J, He C X, et al. Molecular marker-assisted dissection of genotype × environment interaction for plant type traits in rice (Oryza sativa L.). Crop Sci, 1999; 39: 538–544
Liu Z H, Ji H Q, Cui Z T, et al. QTL detected for grain-filling rate in maize using a RIL population. Mol Breed, 2011; 27: 25–36
Cui F, Li J, Ding A M, et al. Conditional QTL mapping for plant height with respect to the length of the spike and internode in two mapping populations of wheat. Theor Appl Genet, 2011; 122: 1517–1536
Diaz U, Saliba-Colombani V, Loudet O, et al. Leaf yellowing and anthocyanin accumulation are two genetically independent strategies in response to nitrogen limitation in Arabidopsis thaliana. Plant Cell Physiol, 2006; 47: 74–83
Salas P, Oyarzo-Llaipen J C, Wang D, et al. Genetic mapping of seed shape in three populations of recombinant inbred lines of soybean (Glycine max L. Merr.). Theor Appl Genet, 2006; 113: 1459–1466
Spickett S G, Thoday J M. Regular Responses to Selection 3. Interaction between located polygenes. Genet Res Camb, 1966; 7: 96–121
Falconer D S. Introduction to Quantitative Genetics. 2nd ed. New York: Longman Press, 1981
Yu S B, Li J X, Tan Y F, et al. Importance of epistasis as the genetic basis of heterosis in an elite rice hybrid. Proc Natl Acad Sci USA, 1997; 94: 9226–9231
Yu S B, Li J X, Xu C G, et al. Epistasis plays an important role as genetic basis of heterosis in rice. Sci China Ser C-Life Sci, 1998; 41: 293–302
Hua J P, Xing Y Z, Wu W R, et al. Single-locus heterotic effects and dominance by dominance interactions can adequately explain the genetic basis of heterosis in an elite rice hybrid. Proc Natl Acad Sci USA, 2003; 100: 2574–2579
Mackay T F C, Stone E A, Ayroles J F. The genetics of quantitative traits: Challenges and prospects. Nat Rev Genet, 2009; 10: 565–577
Frerot H, Faucon M P, Willems G, et al. Genetic architecture of zinc hyperaccumulation in Arabidopsis halleri: The essential role of QTL × environment interactions. New Phytol, 2010; 187: 355–367
Zheng B S, Le Gouis J, Leflon M, et al. Using probe genotypes to dissect QTL × environment interactions for grain yield components in winter wheat. Theor Appl Genet, 2010; 121: 1501–1517
Gauch H G, Rodrigues P C, Munkvold J D, et al. Two New Strategies for detecting and understanding QTL × environment interactions. Crop Sci, 2011; 51: 96–113
Han Y P, Teng W L, Sun D S, et al. Impact of epistasis and QTL × environment interaction on the accumulation of seed mass of soybean (Glycine max L. Merr.). Genet Res, 2008; 90: 481–491
Paterson A H, Damon S, Hewitt J D, et al. Mendelian factors underlying quantitative traits in tomato: Comparison across species, generations, and environments. Genetics, 1991; 127: 181–197
Lu C F, Shen L S, Tan Z B, et al. Comparative mapping of QTLs for agronomic traits of rice across environments by using a doubled-haploid population. Theor Appl Genet, 1997; 94: 145–150
Zhuang J Y, Lin H X, Lu J, et al. Analysis of QTL × environment interaction for yield components and plant height in rice. Theor Appl Genet, 1997; 95: 799–808
Gao Y M, Zhu J. Mapping QTLs with digenic epistasis under multiple environments and predicting heterosis based on QTL effects. Theor Appl Genet, 2007; 115: 325–333
Yang J, Zhu J, Williams R W. Mapping the genetic architecture of complex traits in experimental populations. Bioinformatics, 2007; 23: 1527–1536
Zeng Z B. Precision mapping of quantitative trait loci. Genetics, 1994; 136: 1457–1468
Searle S R, Casella G, McCulloch C E. Variance Components. New York: John Wiley & Sons Inc, 1992
Yang J. Developing methods and software for genetic analysis of complex traits. Doctor Dissertation. Hangzhou: Zhejiang University, 2008
Piepho H P, Gauch H G Jr. Marker pair selection for mapping quantitative trait loci. Genetics, 2001; 157: 433–444
Doerge R W, Churchill G A. Permutation tests for multiple loci affecting a quantitative character. Genetics, 1996; 142: 285–294
Koseki M, Kitazawa N, Yonebayashi S, et al. Identification and fine mapping of a major quantitative trait locus originating from wild rice, controlling cold tolerance at the seedling stage. Mol Genet Genomics, 2010; 284: 45–54
Gilbert H, Riquet J, Gruand J, et al. Detecting QTL for feed intake traits and other performance traits in growing pigs in a Pietrain-Large White backcross. Animal, 2010; 4: 1308–1318
Besnier F, Wahlberg P, Ronnegard L, et al. Fine mapping and replication of QTL in outbred chicken advanced intercross lines. Genet Sel Evol, 2011; 43: 3
Buerstmayr M, Lemmens M, Steiner B, et al. Advanced backcross QTL mapping of resistance to Fusarium head blight and plant morphological traits in a Triticum macha × T. aestivum population. Theor Appl Genet, 2011; 123: 293–306
Stuber C W, Lincoln S E, Wolff D W, et al. Identification of genetic factors contributing to heterosis in a hybrid from two elite maize inbred lines using molecular markers. Genetics, 1992; 132: 823–839
Tang J H, Yan J B, Ma X Q, et al. Dissection of the genetic basis of heterosis in an elite maize hybrid by QTL mapping in an immortalized F2 population. Theor Appl Genet, 2010; 120: 333–340
Greenspan R J. Opinion—The flexible genome. Nat Rev Genet, 2001; 2: 383–387
Author information
Authors and Affiliations
Corresponding author
Additional information
This article is published with open access at Springerlink.com
Rights and permissions
This article is published under an open access license. Please check the 'Copyright Information' section either on this page or in the PDF for details of this license and what re-use is permitted. If your intended use exceeds what is permitted by the license or if you are unable to locate the licence and re-use information, please contact the Rights and Permissions team.
About this article
Cite this article
Zhu, Z., Hayart, Y., Yang, J. et al. Statistical method for mapping QTLs for complex traits based on two backcross populations. Chin. Sci. Bull. 57, 2645–2654 (2012). https://doi.org/10.1007/s11434-012-5279-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11434-012-5279-8