Abstract
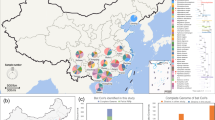
Bats are connected with the increasing numbers of emerging and re-emerging viruses that may break the species barrier and spread into the human population. Coronaviruses are one of the most common viruses discovered in bats, which were considered as the natural source of recent human-susceptible coronaviruses, i.e. SARS-COV and MERS-CoV. Our previous study reported the discovery of a bat-derived putative cross-family recombinant coronavirus with a reovirus gene p10, named as Ro-BatCoV GCCDC1. In this report, through a two-year follow-up of a special bat population in one specific cave of south China, we illustrate that Ro-BatCoV GCCDC1 persistently circulates among bats. Notably, through the longitudinal observation, we identified the dynamic evolution of Ro-BatCoV GCCDC1 in bats represented by continuously recombination events. Our study provides the first glimpse of the virus evolution in one longitudinally observed bat population cohort and underlines the surveillance and pre-warning of potential interspecies transmittable viruses in bats.
Article PDF
Similar content being viewed by others
References
Baric, R.S., Fu, K., Chen, W., and Yount, B. (1995). High recombination and mutation rates in mouse hepatitis virus suggest that coronaviruses may be potentially important emerging viruses. Adv Exp Med Biol 380, 571–576.
Calisher, C.H., Childs, J.E., Field, H.E., Holmes, K.V., and Schountz, T. (2006). Bats: important reservoir hosts of emerging viruses. Clin Microbiol Rev 19, 531–545.
Carrasco, L., Otero, M.J., and Castrillo, J.L. (1989). Modification of membrane permeability by animal viruses. Pharmacol Ther 40, 171–212.
Carrasco, L., Perez, L., Irurzun, A., Lama, J., Martinez-Abarca, F., Rodriguez, P., Guinea, R., Castrillo, J.L., Sanz, M.A., and Ayala, M.J. (1993). Regulation of Gene Expression in Animal Viruses. (New York: Plenum Publishing Corp.), pp. 283–305.
Du, J., Yang, L., Ren, X., Zhang, J., Dong, J., Sun, L., Zhu, Y., Yang, F., Zhang, S., Wu, Z., and Jin, Q. (2016). Genetic diversity of coronaviruses in Miniopterus fuliginosus bats. Sci China Life Sci 59, 604–614.
Ge, X.Y., Li, J.L., Yang, X.L., Chmura, A.A., Zhu, G., Epstein, J.H., Mazet, J.K., Hu, B., Zhang, W., Peng, C., Zhang, Y.J., Luo, C.M., Tan, B., Wang, N., Zhu, Y., Crameri, G., Zhang, S.Y., Wang, L.F., Daszak, P., and Shi, Z.L. (2013). Isolation and characterization of a bat SARS-like coronavirus that uses the ACE2 receptor. Nature 503, 535–538.
Hu, B., Zeng, L.P., Yang, X.L., Ge, X.Y., Zhang, W., Li, B., Xie, J.Z., Shen, X.R., Zhang, Y.Z., Wang, N., Luo, D.S., Zheng, X.S., Wang, M.N., Daszak, P., Wang, L.F., Cui, J., and Shi, Z.L. (2017). Discovery of a rich gene pool of bat SARS-related coronaviruses provides new insights into the origin of SARS coronavirus. PLoS Pathog 13, e1006698.
Huang, C., Liu, W.J., Xu, W., Jin, T., Zhao, Y., Song, J., Shi, Y., Ji, W., Jia, H., Zhou, Y., Wen, H., Zhao, H., Liu, H., Li, H., Wang, Q., Wu, Y., Wang, L., Liu, D., Liu, G., Yu, H., Holmes, E.C., Lu, L., and Gao, G.F. (2016). A bat-derived putative cross-family recombinant coronavirus with a reovirus gene. PLoS Pathog 12, e1005883.
Jackwood, M.W., Hall, D., and Handel, A. (2012). Molecular evolution and emergence of avian gammacoronaviruses. Infect Genet Evol 12, 1305–1311.
Lai, M.M.C. (1996). Recombination in large RNA viruses: coronaviruses. Semin Virol 7, 381–388.
Lau, S.K.P., Woo, P.C.Y., Li, K.S.M., Huang, Y., Tsoi, H.W., Wong, B.H.L., Wong, S.S.Y., Leung, S.Y., Chan, K.H., and Yuen, K.Y. (2005). Severe acute respiratory syndrome coronavirus-like virus in Chinese horseshoe bats. Proc Natl Acad Sci USA 102, 14040–14045.
Lelli, D., Papetti, A., Sabelli, C., Rosti, E., Moreno, A., and Boniotti, M.B. (2013). Detection of coronaviruses in bats of various species in italy. Viruses 5, 2679–2689.
Li, W., Shi, Z., Yu, M., Ren, W., Smith, C., Epstein, J.H., Wang, H., Crameri, G., Hu, Z., Zhang, H., Zhang, J., McEachern, J., Field, H., Daszak, P., Eaton, B.T., Zhang, S., and Wang, L.F. (2005). Bats are natural reservoirs of SARS-like coronaviruses. Science 310, 676–679.
Li, Z., Liu, D., Ran, X., Liu, C., Guo, D., Hu, X., Tian, J., Zhang, X., Shao, Y., Liu, S., and Qu, L. (2016). Characterization and pathogenicity of a novel mammalian orthoreovirus from wild short-nosed fruit bats. Infect Genet Evol 43, 347–353.
López-Roig, M., Bourhy, H., Lavenir, R., and Serra-Cobo, J. (2014). Seroprevalence dynamics of european bat lyssavirus type 1 in a multispecies bat colony. Viruses 6, 3386–3399.
Minskaia, E., Hertzig, T., Gorbalenya, A.E., Campanacci, V., Cambillau, C., Canard, B., and Ziebuhr, J. (2006). Discovery of an RNA virus 3′→5′ exoribonuclease that is critically involved in coronavirus RNA synthesis. Proc Natl Acad Sci USA 103, 5108–5113.
Sokolova, T.M., Uryvaev, L.V., Selivanova, T.K., Bobrova, O.V., Lebedev, A., and Bystrov, N.S. (1996). Recombination of alpha-viruses during mixed infection in cultured cells. Formation of hybrid forms of RNA of Venezuelan equine encephalomyelitis virus and Karelian fever virus in the region of genes for nucleocapsid and envelope proteins. Vopr Virusol 41, 245–252.
Wang, Q., Qi, J., Yuan, Y., Xuan, Y., Han, P., Wan, Y., Ji, W., Li, Y., Wu, Y., Wang, J., Iwamoto, A., Woo, P.C., Yuen, K.Y., Yan, J., Lu, G., and Gao, G.F. (2014). Bat origins of MERS-CoV supported by bat coronavirus HKU4 usage of human receptor CD26. Cell Host Microbe 16, 328–337.
Woo, P.C.Y., Lau, S.K.P., Lam, C.S.F., Lau, C.C.Y., Tsang, A.K.L., Lau, J. H.N., Bai, R., Teng, J.L.L., Tsang, C.C.C., Wang, M., Zheng, B.J., Chan, K.H., and Yuen, K.Y. (2012). Discovery of seven novel mammalian and avian coronaviruses in the genus deltacoronavirus supports bat coronaviruses as the gene source of alphacoronavirus and betacoronavirus and avian coronaviruses as the gene source of gammacoronavirus and deltacoronavirus. J Virol 86, 3995–4008.
Zumla, A., Chan, J.F.W., Azhar, E.I., Hui, D.S.C., and Yuen, K.Y. (2016). Coronaviruses—drug discovery and therapeutic options. Nat Rev Drug Discov 15, 327–347.
Acknowledgements
We appreciate the great efforts of Drs. Yongming Zhou, Honghua Wen, Huaxing Liu, who participated in the collection of the bat samples. This work was supported by the National Key Research and Development Program of China (2017YFC1200202), the National Natural Science Foundation of China (81290342, 81461168030), the Major Special Projects for Infectious Disease Research of China (2016ZX10004222-003), and China National Grand S&T Special Project (2014ZX10004-001-006). Joseph O. Obameso was supported by CAS-TWAS President’s Fellowship of the University of Chinese Academy of Sciences (UCAS) and The World Academy of Sciences (TWAS). George F. Gao is a leading principle investigator of the NSFC Innovative Research Group (81621091). The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.
Author information
Authors and Affiliations
Corresponding authors
Rights and permissions
About this article
Cite this article
Obameso, J.O., Li, H., Jia, H. et al. The persistent prevalence and evolution of cross-family recombinant coronavirus GCCDC1 among a bat population: a two-year follow-up. Sci. China Life Sci. 60, 1357–1363 (2017). https://doi.org/10.1007/s11427-017-9263-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11427-017-9263-6