Abstract
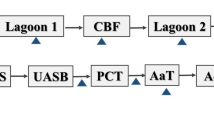
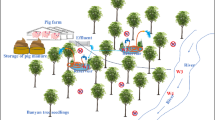
Swine manure treatment plants are important reservoirs of plasmid-harboring antibiotic resistance genes (ARGs) and physicochemical contaminants, but the changes in the abundances of plasmids and ARGs, and their interactions with the physicochemical properties of manure, are still unclear. Thus, in the present study, plasmidome and metagenome analyses were conducted for samples collected at different stages in the swine manure treatment process. The results indicated that anaerobic digestion and aerobic digestion were the most efficient stages for reducing the abundances of ARGs in swine manure. However, the plasmids associated with ARGs were not effectively removed in these stages. Through the whole treatment process, the IncL/M, IncQ1, IncHI2A, IncA/C, and IncN plasmid groups had strong correlations (r > 0.8, P < 0.01) with most ARG types, thereby indicating that these plasmids play important roles in the persistence of ARGs in this environment. Furthermore, the pH, total nitrogen, total phosphorus, and four heavy metals (Cu, Zn, As, and Fe) significantly affected the abundances of seven ARG subtypes (tetB(P), ant(6)-Ia, tet44, aph(3′′)-Ib, mefB, tet(L), and tet(39)). In particular, florfenicol had the most positive correlations with ARGs. Our results indicated that nutrients, heavy metals, and antibiotics all contributed to the presence and persistence of plasmid-harboring ARGs. This study provides insights into the fate of plasmids and ARGs, and related factors during the swine manure treatment process, thereby facilitating the development of a new treatment technique for removing ARGs and reducing the public health risk associated with livestock production.
Graphical abstract






Similar content being viewed by others
Data availability
All of the sequences in the plasmid database analyzed in the present study are available via the NCBI RefSeq repository (https://ftp.ncbi.nlm.nih.gov/refseq/release/plasmid/) (2020–11-16). The nucleotide sequences in the plasmidome and metagenome were deposited in the GSA database, and they are publicly available under accession number PRJCA004346.
Code availability
The code used in this study to generate the plasmid database is available at Github (https://gist.github.com/Alice-shui/88d39ffe3509172e32826772b5b88aa6).
References
Alcock BP, Raphenya AR, Lau TTY, Tsang KK, Bouchard M, Edalatmand A, Huynh W, Nguyen AV, Cheng AA, Liu S, Min SY, Miroshnichenko A, Tran HK, Werfalli RE, Nasir JA, Oloni M, Speicher DJ, Florescu A, Singh B, Faltyn M, Hernandez-Koutoucheva A, Sharma AN, Bordeleau E, Pawlowski AC, Zubyk HL, Dooley D, Griffiths E, Maguire F, Winsor GL, Beiko RG, Brinkman FSL, Hsiao WWL, Domselaar GV, McArthur AG (2020) CARD 2020: antibiotic resistome surveillance with the comprehensive antibiotic resistance database. Nucleic Acids Res 48(D1):D517-d525
Antipov D, Raiko M, Lapidus A, Pevzner PA (2019) Plasmid detection and assembly in genomic and metagenomic data sets. Genome Res 29(6):961–968
Baker-Austin C, Wright MS, Stepanauskas R, McArthur JV (2006) Co-selection of antibiotic and metal resistance. Trends Microbiol 14(4):176–182
Batt AL, Kostich MS, Lazorchak JM (2008) Analysis of ecologically relevant pharmaceuticals in wastewater and surface water using selective solid-phase extraction and UPLC− MS/MS. Anal Chem. 80(13):5021–5030
Bennett P (2008) Plasmid encoded antibiotic resistance: acquisition and transfer of antibiotic resistance genes in bacteria. Br J Pharmacol 153(S1):S347–S357
Bolger AM, Lohse M, Usadel B (2014) Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 30(15):2114–2120
Carattoli A, Zankari E, Garcìa-Fernandez A, Larsen MV, Lund O, Villa L, Aarestrup FM, Hasman H (2014) PlasmidFinder and pMLST: in silico detection and typing of plasmids. Antimicrob Agents Chemother
Carlet J (2015) The world alliance against antibiotic resistance: consensus for a declaration. Clin Infect Dis 60(12):1837–1841
Che Y, Xia Y, Liu L, Li A-D, Yang Y, Zhang T (2019) Mobile antibiotic resistome in wastewater treatment plants revealed by Nanopore metagenomic sequencing. Microbiome 7(1):44
Che Y, Yang Y, Xu X, Břinda K, Polz MF, Hanage WP, Zhang T (2021) Conjugative plasmids interact with insertion sequences to shape the horizontal transfer of antimicrobial resistance genes. Proc Nat Acad Sci 118(6):e2008731118
Cheng W, Chen H, Su C, Yan S (2013) Abundance and persistence of antibiotic resistance genes in livestock farms: a comprehensive investigation in eastern China. Environ Int 61:1–7
Dib JR, Wagenknecht M, Farías ME, Meinhardt F (2015) Strategies and approaches in plasmidome studies—uncovering plasmid diversity disregarding of linear elements? Front Microbiol 6(463)
Dickinson AW, Power A, Hansen MG, Brandt KK, Piliposian G, Appleby P, O’Neill PA, Jones RT, Sierocinski P, Koskella B, Vos M (2019) Heavy metal pollution and co-selection for antibiotic resistance: a microbial palaeontology approach. Environ Int 132:105117
Dyson ZA, Klemm EJ, Palmer S, Dougan G (2019) Antibiotic resistance and typhoid. Clin Infect Dis 68(Suppl 2):S165-s170
Guo MT, Yuan QB, Yang J (2015) Distinguishing effects of ultraviolet exposure and chlorination on the horizontal transfer of antibiotic resistance genes in municipal wastewater. Environ Sci Technol 49(9):5771–5778
Guo H, Nasir M, Lv J, Dai Y, Gao J (2017) Understanding the variation of microbial community in heavy metals contaminated soil using high throughput sequencing. Ecotoxicol Environ Saf 144:300–306
He L-Y, Ying G-G, Liu Y-S, Su H-C, Chen J, Liu S-S, Zhao J-L (2016) Discharge of swine wastes risks water quality and food safety: Antibiotics and antibiotic resistance genes from swine sources to the receiving environments. Environ Int 92–93:210–219
Heuer H, Schmitt H, Smalla K (2011) Antibiotic resistance gene spread due to manure application on agricultural fields. Curr Opin Microbiol 14(3):236–243
Kime L, Randall CP, Banda FI, Coll F, Wright J, Richardson J, Empel J, Parkhill J, O'Neill AJ (2019) Transient silencing of antibiotic resistance by mutation represents a significant potential source of unanticipated therapeutic failure. mBio 10(5)
Liang-wei DJCB. 2001. Models of pig waste treatment. 1
Loftie-Eaton W, Yano H, Burleigh S, Simmons RS, Hughes JM, Rogers LM, Hunter SS, Settles ML, Forney LJ, Ponciano JM, Top EM (2016) Evolutionary paths that expand plasmid host-range: implications for spread of antibiotic resistance. Mol Biol Evol 33(4):885–897
Patro R, Duggal G, Love MI, Irizarry RA, Kingsford C (2017) Salmon provides fast and bias-aware quantification of transcript expression. Nat Methods 14(4):417–419
Rozwandowicz M, Brouwer M, Fischer J, Wagenaar J, Gonzalez-Zorn B, Guerra B, Mevius D, Hordijk J (2018) Plasmids carrying antimicrobial resistance genes in Enterobacteriaceae. J Antimicrob Chemother 73(5):1121–1137
San-cheng Y, Xue-bin L, Zhi-ping HJSA and Sciences V (2010) Observation of different patterns for pig waste treatment and utilization in Sichuan. 6
Segata N, Waldron L, Ballarini A, Narasimhan V, Jousson O, Huttenhower C (2012) Metagenomic microbial community profiling using unique clade-specific marker genes. Nat Methods 9(8):811–814
Selvam A, Xu D, Zhao Z, Wong JW (2012) Fate of tetracycline, sulfonamide and fluoroquinolone resistance genes and the changes in bacterial diversity during composting of swine manure. Bioresour Technol 126:383–390
Szczepanowski R, Bekel T, Goesmann A, Krause L, Krömeke H, Kaiser O, Eichler W, Pühler A, Schlüter A (2008) Insight into the plasmid metagenome of wastewater treatment plant bacteria showing reduced susceptibility to antimicrobial drugs analysed by the 454-pyrosequencing technology. J Biotechnol 136(1–2):54–64
Tao CW, Hsu BM, Ji WT, Hsu TK, Kao PM, Hsu CP, Shen SM, Shen TY, Wan TJ, Huang YL (2014) Evaluation of five antibiotic resistance genes in wastewater treatment systems of swine farms by real-time PCR. Sci Total Environ 496:116–121
Tong J, Liu J, Zheng X, Zhang J, Ni X, Chen M, Wei Y (2016) Fate of antibiotic resistance bacteria and genes during enhanced anaerobic digestion of sewage sludge by microwave pretreatment. Biores Technol 217:37–43
Van den Meersche T, Rasschaert G, Haesebrouck F, Van Coillie E, Herman L, Van Weyenberg S, Daeseleire E, Heyndrickx M (2019) Presence and fate of antibiotic residues, antibiotic resistance genes and zoonotic bacteria during biological swine manure treatment. Ecotoxicol Environ Saf 175:29–38
Wagner GP, Kin K, Lynch VJ (2012) Measurement of mRNA abundance using RNA-seq data: RPKM measure is inconsistent among samples. Theory Bio 131(4):281–285
Wang FH, Ma W, Dou Z, Ma L, Liu XL, Xu JX, Zhang FS (2006) The estimation of the production amount of animal manure and its environmental effect in China [in Chinese]. Zhongguo Huanjing Kexue/China Environ Sci 26:614–617
Wang J, Gu J, Wang X, Song Z, Dai X, Guo H, Yu J, Zhao W, Lei L (2021) Enhanced removal of antibiotic resistance genes and mobile genetic elements during swine manure composting inoculated with mature compost. J Hazard Mater 411:125135
Wen X, Mi J, Wang Y, Ma B, Zou Y, Liao X, Liang JB, Wu Y (2019) Occurrence and contamination profiles of antibiotic resistance genes from swine manure to receiving environments in Guangdong Province southern China. Ecotoxicol Environ Saf 173:96–102
Xiong W, Wang Y, Sun Y, Ma L, Zeng Q, Jiang X, Li A, Zeng Z, Zhang T (2018) Antibiotic-mediated changes in the fecal microbiome of broiler chickens define the incidence of antibiotic resistance genes. Microbiome 6(1):34
Yang C, Dong Y, Friman VP, Jousset A, Wei Z, Xu Y, Shen Q (2019) Carbon resource richness shapes bacterial competitive interactions by alleviating growth-antibiosis trade-off. Funct Ecol 33(5):868–875
Yuan CG, Shi JB, He B, Liu JF, Liang LN, Jiang GB (2004) Speciation of heavy metals in marine sediments from the East China Sea by ICP-MS with sequential extraction. Environ Int 30(6):769–783
Yuan QB, Zhai YF, Mao BY, Schwarz C, Hu N (2019) Fates of antibiotic resistance genes in a distributed swine wastewater treatment plant. Water Environ Res: Res Publ Water Environ Fed 91(12):1565–1575
Zankari E, Hasman H, Cosentino S, Vestergaard M, Rasmussen S, Lund O, Aarestrup FM, Larsen MV (2012) Identification of acquired antimicrobial resistance genes. J Antimicrob Chemother 67(11):2640–2644
Zhang J, Chen M, Sui Q, Wang R, Tong J, Wei Y (2016) Fate of antibiotic resistance genes and its drivers during anaerobic co-digestion of food waste and sewage sludge based on microwave pretreatment. Biores Technol 217:28–36
Zhang Y-J, Hu H-W, Gou M, Wang J-T, Chen D, He J-Z (2017) Temporal succession of soil antibiotic resistance genes following application of swine, cattle and poultry manures spiked with or without antibiotics. Environ Pollut 231:1621–1632
Zhang Y, Yang Z, Xiang Y, Xu R, Zheng Y, Lu Y, Jia M, Sun S, Cao J, Xiong W (2020) Evolutions of antibiotic resistance genes (ARGs), class 1 integron-integrase (intI1) and potential hosts of ARGs during sludge anaerobic digestion with the iron nanoparticles addition. Sci Total Environ 724:138248
Zhao X, Wang J, Zhu L, Ge W, Wang J (2017) Environmental analysis of typical antibiotic-resistant bacteria and ARGs in farmland soil chronically fertilized with chicken manure. Sci Total Environ 593–594:10–17
Zhou B, Wang C, Zhao Q, Wang Y, Huo M, Wang J, Wang S (2016) Prevalence and dissemination of antibiotic resistance genes and coselection of heavy metals in Chinese dairy farms. J Hazard Mater 320:10–17
Zhou Z-C, Zheng J, Wei Y-Y, Chen T, Dahlgren RA, Shang X, Chen H (2017) Antibiotic resistance genes in an urban river as impacted by bacterial community and physicochemical parameters. Environ Sci Pollut Res 24(30):23753–23762
Zhu Y-G, Johnson TA, Su J-Q, Qiao M, Guo G-X, Stedtfeld RD, Hashsham SA, Tiedje JM (2013) Diverse and abundant antibiotic resistance genes in Chinese swine farms. Proc Natl Acad Sci 110(9):3435–3440
Zhu YG, Johnson TA, Su JQ, Qiao M, Guo GX, Stedtfeld RD, Hashsham SA, Tiedje JM (2013) Diverse and abundant antibiotic resistance genes in Chinese swine farms. Proc Natl Acad Sci U S A 110(9):3435–3440
Funding
This study was supported by the General Program of National Natural Science Foundation of China (grant number [U21A20257]), Key R&D Program of Sichuan province (grant numbers [22ZDZX0011], [2020ZYD003], [2020YFN0147], and [2021YFH0192]), and Project of Science and Technology Bureau of Banan District Chongqing (grant number [BN2020-115]).
Author information
Authors and Affiliations
Contributions
Conceptualization: Junrui Shui, Hongmei Tuo, Anyun Zhang; Methodology: Junrui Shui, Hongmei Tuo, Anyun Zhang; formal analysis and investigation: Hongmei Tuo, Junrui Shui; writing—original draft: Junrui Shui, Hongmei Tuo, Jinxin Liu, Anyun Zhang; writing—review & editing: Junrui Shui, Anyun Zhang, Jinxin Liu, Xialan Zhang, Jingyi Feng, Yuxuan Feng, Wen Su, Cong Lin, Haoyu Zhang, Zunfang Tu; funding acquisition: Anyun Zhang, Hongning Wang.
Corresponding author
Ethics declarations
Ethics approval and consent to participate
Not applicable.
Consent for publication
Not applicable.
Competing interests
The authors declare no competing interests.
Additional information
Responsible Editor: Diane Purchase
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Shui, J., Tuo, H., Liu, J. et al. Insights into the fates of plasmids and antimicrobial resistance genes during swine manure treatment and related factors based on plasmidome and metagenome analyses. Environ Sci Pollut Res 29, 69037–69047 (2022). https://doi.org/10.1007/s11356-022-20574-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11356-022-20574-7