Abstract
Aquaculture sediments are a purported sizable pool of antibiotic resistance genes (ARGs). However, the pathways for transmission of ARGs from sediments to animals and humans remain unclear. We conducted an ARG survey in sediments from a bullfrog production facility located in Guangdong, China, and simulated zebrafish breeding systems were constructed, with or without biochar addition in sediments, to explore the effects of biochar on ARGs and their precursors of the sediment and zebrafish gut. After 60 days, 6 subtypes of ARGs and intI1 were detected, with sediments harboring more ARGs than zebrafish gut. The addition of biochar reduced the abundance of ARGs in the sediment and zebrafish gut, as well as suppressed the horizontal transmission of ARGs from sediment to zebrafish gut. Network analysis and partial least squares path modeling revealed that ARG enrichment was mainly affected by bacterial groups dominated by Nitrospirae, Gemmatimonades, Chloroflexi, and Cyanobacteria and intI1. Our findings provide insights into the transmission of ARGs from sediment to animals and highlight the efficacy of biochar amendments to aquaculture sediments to reduce the transmission of ARGs.

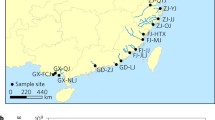
source water), S2 = bullfrog pond, and S3 = gutterway (wastewater discharge)







Similar content being viewed by others
Data availability
All data generated or analyzed during this study are included in this published article and its supplementary information files.
References
Abe K, Nomura N, Suzuki S (2020): Biofilms: hot spots of horizontal gene transfer (HGT) in aquatic environments, with a focus on a new HGT mechanism. FEMS Microbiol Ecol 96 https://doi.org/10.1093/femsec/fiaa031
Allen HK, Donato J, Wang HH, Cloud-Hansen KA, Davies J, Handelsman J (2010) Call of the wild: antibiotic resistance genes in natural environments. Nat Rev Microbiol 8:251–259. https://doi.org/10.1038/nrmicro2312
Anjum R, Krakat N (2016) Detection of multiple resistances, biofilm formation and conjugative transfer of Bacillus cereus from contaminated soils. Curr Microbiol 72:321–328. https://doi.org/10.1007/s00284-015-0952-1
Caporaso JG et al (2010) QIIME allows analysis of high-throughput community sequencing data. Nat Methods 7:335–336. https://doi.org/10.1038/nmeth.f.303
Chen C, Pankow CA, Oh M, Heath LS, Zhang L, Du P, Xia K, Pruden A (2019) Effect of antibiotic use and composting on antibiotic resistance gene abundance and resistome risks of soils receiving manure-derived amendments. Environ Int 128:233–243. https://doi.org/10.1016/j.envint.2019.04.043
Chen Q-L, Fan X-T, Zhu D, An X-L, Su J-Q, Cui L (2018) Effect of biochar amendment on the alleviation of antibiotic resistance in soil and phyllosphere of Brassica chinensis L. Soil Biol Biochem 119:74–82. https://doi.org/10.1016/j.soilbio.2018.01.015
Cheng JH, Tang XY, Su JQ, Liu C (2021) Field aging alters biochar’s effect on antibiotic resistome in manured soil. Environ Pollut 288:117719. https://doi.org/10.1016/j.envpol.2021.117719
Cohen ME, Salmasian H, Li J, Liu J, Zachariah P, Wright JD, Freedberg DE (2017) Surgical antibiotic prophylaxis and risk for postoperative antibiotic-resistant infections. J Am Coll Surg 225(631–638):e3. https://doi.org/10.1016/j.jamcollsurg.2017.08.010
D’Costa VM, King CE, Kalan L, Morar M, Sung WW, Schwarz C, Froese D, Zazula G, Calmels F, Debruyne R, Golding GB, Poinar HN, Wright GD (2011) Antibiotic resistance is ancient. Nature 477:457–461. https://doi.org/10.1038/nature10388
Ding C, Ma J, Jiang W, Zhao H, Shi M, Cui G, Yan T, Wang Q, Li J, Qiu Z (2021) Chironomidae larvae: A neglected enricher of antibiotic resistance genes in the food chain of freshwater environments. Environ Pollut 285:117486. https://doi.org/10.1016/j.envpol.2021.117486
Ding J, Zhu D, Hong B, Wang HT, Li G, Ma YB, Tang YT, Chen QL (2019) Long-term application of organic fertilization causes the accumulation of antibiotic resistome in earthworm gut microbiota. Environ Int 124:145–152. https://doi.org/10.1016/j.envint.2019.01.017
Duan M, Li H, Gu J, Tuo X, Sun W, Qian X, Wang X (2017) Effects of biochar on reducing the abundance of oxytetracycline, antibiotic resistance genes, and human pathogenic bacteria in soil and lettuce. Environ Pollut 224:787–795. https://doi.org/10.1016/j.envpol.2017.01.021
Duan M, Gu J, Wang X, Li Y, Zhang R, Hu T, Zhou B (2019) Factors that affect the occurrence and distribution of antibiotic resistance genes in soils from livestock and poultry farms. Ecotoxicol Environ Saf 180:114–122. https://doi.org/10.1016/j.ecoenv.2019.05.005
Ellabaan MMH, Munck C, Porse A, Imamovic L, Sommer MOA (2021) Forecasting the dissemination of antibiotic resistance genes across bacterial genomes. Nat Commun 12:2435. https://doi.org/10.1038/s41467-021-22757-1
Gao P, Mao D, Luo Y, Wang L, Xu B, Xu L (2012) Occurrence of sulfonamide and tetracycline-resistant bacteria and resistance genes in aquaculture environment. Water Res 46:2355–2364. https://doi.org/10.1016/j.watres.2012.02.004
Giatsis C, Sipkema D, Smidt H, Heilig H, Benvenuti G, Verreth J, Verdegem M (2015) The impact of rearing environment on the development of gut microbiota in tilapia larvae. Sci Rep 5:18206. https://doi.org/10.1038/srep18206
Gillings MR (2013) Evolutionary consequences of antibiotic use for the resistome, mobilome and microbial pangenome. Front Microbiol 4:4. https://doi.org/10.3389/fmicb.2013.00004
Gillings MR, Gaze WH, Pruden A, Smalla K, Tiedje JM, Zhu YG (2015) Using the class 1 integron-integrase gene as a proxy for anthropogenic pollution. ISME J 9:1269–1279. https://doi.org/10.1038/ismej.2014.226
Guglielmetti E, Korhonen JM, Heikkinen J, Morelli L, von Wright A (2009) Transfer of plasmid-mediated resistance to tetracycline in pathogenic bacteria from fish and aquaculture environments. FEMS Microbiol Lett 293:28–34. https://doi.org/10.1111/j.1574-6968.2009.01512.x
Guo HG, Chen QL, Hu HW, He JZ (2020) Fate of antibiotic resistance genes during high-solid anaerobic co-digestion of pig manure with lignite. Bioresour Technol 303:122906. https://doi.org/10.1016/j.biortech.2020.122906
He Y, Liu C, Tang X-Y, Xian Q-S, Zhang J-Q, Guan Z (2019) Biochar impacts on sorption-desorption of oxytetracycline and florfenicol in an alkaline farmland soil as affected by field ageing. Sci Total Environ 671:928–936. https://doi.org/10.1016/j.scitotenv.2019.03.414
Iwu CD, Patrick SM (2021) An insight into the implementation of the global action plan on antimicrobial resistance in the WHO African region: a roadmap for action. Int J Antimicrob Agents 58:106411. https://doi.org/10.1016/j.ijantimicag.2021.106411
Jiao W, Du R, Ye M, Sun M, Feng Y, Wan J, Zhao Y, Zhang Z, Huang D, Du D, Jiang X (2018) “Agricultural Waste to Treasure” - Biochar and eggshell to impede soil antibiotics/antibiotic resistant bacteria (genes) from accumulating in Solanum tuberosum L. Environ Pollut 242:2088–2095. https://doi.org/10.1016/j.envpol.2018.06.059
Kim DH, Brunt J, Austin B (2007) Microbial diversity of intestinal contents and mucus in rainbow trout (Oncorhynchus mykiss). J Appl Microbiol 102:1654–1664. https://doi.org/10.1111/j.1365-2672.2006.03185.x
Knapp CW, Dolfing J, Ehlert PAI, Graham DW (2010) Evidence of increasing antibiotic resistance gene abundances in archived soils since 1940. Environ Sci Technol 44:580–587. https://doi.org/10.1021/es901221x
Langille MGI, Zaneveld J, Caporaso JG, McDonald D, Knights D, Reyes JA, Clemente JC, Burkepile DE, Vega Thurber RL, Knight R, Beiko RG, Huttenhower C (2013) Predictive functional profiling of microbial communities using 16S rRNA marker gene sequences. Nat Biotechnol 31:814–821. https://doi.org/10.1038/nbt.2676
Liang Y, Pei M, Wang D, Cao S, Xiao X, Sun B (2017) Improvement of soil ecosystem multifunctionality by dissipating manure-induced antibiotics and resistance genes. Environ Sci Technol 51:4988–4998. https://doi.org/10.1021/acs.est.7b00693
Ma J, Gu J, Wang X, Peng H, Wang Q, Zhang R, Hu T, Bao J (2019) Effects of nano-zerovalent iron on antibiotic resistance genes during the anaerobic digestion of cattle manure. Bioresour Technol 289:121688. https://doi.org/10.1016/j.biortech.2019.121688
Martinez JL (2009) Environmental pollution by antibiotics and by antibiotic resistance determinants. Environ Pollut 157:2893–2902. https://doi.org/10.1016/j.envpol.2009.05.051
McDonald D, Price MN, Goodrich J, Nawrocki EP, DeSantis TZ, Probst A, Andersen GL, Knight R, Hugenholtz P (2012) An improved Greengenes taxonomy with explicit ranks for ecological and evolutionary analyses of bacteria and archaea. ISME J 6:610–618. https://doi.org/10.1038/ismej.2011.139
McKinney CW, Dungan RS, Moore A, Leytem AB (2018): Occurrence and abundance of antibiotic resistance genes in agricultural soil receiving dairy manure. FEMS Microbiol Ecol 94 https://doi.org/10.1093/femsec/fiy010
Ochman H, Lawrence JG, Groisman EA (2000) Lateral gene transfer and the nature of bacterial innovation. Nature 405:299–304. https://doi.org/10.1038/35012500
Pal C, Bengtsson-Palme J, Kristiansson E, Larsson DG (2016) The structure and diversity of human, animal and environmental resistomes. Microbiome 4:54. https://doi.org/10.1186/s40168-016-0199-5
Qian X, Gu J, Sun W, Wang XJ, Su JQ, Stedfeld R (2018) Diversity, abundance, and persistence of antibiotic resistance genes in various types of animal manure following industrial composting. J Hazard Mater 344:716–722. https://doi.org/10.1016/j.jhazmat.2017.11.020
Razavi M, Marathe NP, Gillings MR, Flach CF, Kristiansson E, Joakim Larsson DG (2017) Discovery of the fourth mobile sulfonamide resistance gene. Microbiome 5:160. https://doi.org/10.1186/s40168-017-0379-y
Sandberg KD, LaPara TM (2016): The fate of antibiotic resistance genes and class 1 integrons following the application of swine and dairy manure to soils. FEMS Microbiol Ecol 92 https://doi.org/10.1093/femsec/fiw001
Shterzer N, Mizrahi I (2015) The animal gut as a melting pot for horizontal gene transfer. Can J Microbiol 61:603–605. https://doi.org/10.1139/cjm-2015-0049
Song W, Wang X, Gu J, Zhang S, Yin Y, Li Y, Qian X, Sun W (2017) Effects of different swine manure to wheat straw ratios on antibiotic resistance genes and the microbial community structure during anaerobic digestion. Bioresour Technol 231:1–8. https://doi.org/10.1016/j.biortech.2017.01.054
Su J-Q, Wei B, Ou-Yang W-Y, Huang F-Y, Zhao Y, Xu H-J, Zhu Y-G (2015) Antibiotic resistome and its association with bacterial communities during sewage sludge composting. Environ Sci Technol 49:7356–7363. https://doi.org/10.1021/acs.est.5b01012
Sun M, Ye M, Zhang Z, Zhang S, Zhao Y, Deng S, Kong L, Ying R, Xia B, Jiao W, Cheng J, Feng Y, Liu M, Hu F (2019) Biochar combined with polyvalent phage therapy to mitigate antibiotic resistance pathogenic bacteria vertical transfer risk in an undisturbed soil column system. J Hazard Mater 365:1–8. https://doi.org/10.1016/j.jhazmat.2018.10.093
Sun W, Gu J, Wang X, Qian X, Tuo X (2018) Impacts of biochar on the environmental risk of antibiotic resistance genes and mobile genetic elements during anaerobic digestion of cattle farm wastewater. Bioresour Technol 256:342–349. https://doi.org/10.1016/j.biortech.2018.02.052
Tiimub BM, Zhou ZC, Zhu L, Liu Y, Shuai XY, Xu L, Niyungeko C, Meng LX, Sun YJ, Chen H (2021) Characteristics of bacterial community and ARGs profile in engineered goldfish tanks with stresses of sulfanilamide and copper. Environ Sci Pollut Res Int 28:38706–38717. https://doi.org/10.1007/s11356-021-13239-4
Wang F, Fu Y-H, Sheng H-J, Topp E, Jiang X, Zhu Y-G, Tiedje JM (2021) Antibiotic resistance in the soil ecosystem: a One Health perspective. Current Opinion in Environmental Science & Health 20:100230. https://doi.org/10.1016/j.coesh.2021.100230
Wang J, Ben W, Yang M, Zhang Y, Qiang Z (2016) Dissemination of veterinary antibiotics and corresponding resistance genes from a concentrated swine feedlot along the waste treatment paths. Environ Int 92–93:317–323. https://doi.org/10.1016/j.envint.2016.04.020
Yu Q, Feng T, Yang J, Su W, Zhou R, Wang Y, Zhang H, Li H (2022) Seasonal distribution of antibiotic resistance genes in the Yellow River water and tap water, and their potential transmission from water to human. Environ Pollut 292:118304. https://doi.org/10.1016/j.envpol.2021.118304
Yuan K, Wang X, Chen X, Zhao Z, Fang L, Chen B, Jiang J, Luan T, Chen B (2019) Occurrence of antibiotic resistance genes in extracellular and intracellular DNA from sediments collected from two types of aquaculture farms. Chemosphere 234:520–527. https://doi.org/10.1016/j.chemosphere.2019.06.085
Zhang S, Pang S, Wang P, Wang C, Han N, Liu B, Han B, Li Y, Anim-Larbi K (2016) Antibiotic concentration and antibiotic-resistant bacteria in two shallow urban lakes after stormwater event. Environ Sci Pollut Res Int 23:9984–9992. https://doi.org/10.1007/s11356-016-6237-9
Zhang S, Yang G, Hou S, Zhang T, Li Z, Liang F (2018) Distribution of ARGs and MGEs among glacial soil, permafrost, and sediment using metagenomic analysis. Environ Pollut 234:339–346. https://doi.org/10.1016/j.envpol.2017.11.031
Zhang X, Li X, Wang W, Qi J, Wang D, Xu L, Liu Y, Zhang Y, Guo K (2020a) Diverse gene cassette arrays prevail in commensal Escherichia coli from intensive farming swine in four provinces of China. Front Microbiol 11:565349. https://doi.org/10.3389/fmicb.2020.565349
Zhang Y, Yang Z, Xiang Y, Xu R, Zheng Y, Lu Y, Jia M, Sun S, Cao J, Xiong W (2020b) Evolutions of antibiotic resistance genes (ARGs), class 1 integron-integrase (intI1) and potential hosts of ARGs during sludge anaerobic digestion with the iron nanoparticles addition. The Science of the Total Environment 724:138248. https://doi.org/10.1016/j.scitotenv.2020.138248
Zhang YJ, Hu HW, Chen QL, Singh BK, Yan H, Chen D, He JZ (2019) Transfer of antibiotic resistance from manure-amended soils to vegetable microbiomes. Environ Int 130:104912. https://doi.org/10.1016/j.envint.2019.104912
Zheng H, Feng N, Yang T, Shi M, Wang X, Zhang Q, Zhao J, Li F, Sun K, Xing B (2021) Individual and combined applications of biochar and pyroligneous acid mitigate dissemination of antibiotic resistance genes in agricultural soil. The Science of the Total Environment 796:148962. https://doi.org/10.1016/j.scitotenv.2021.148962
Zhi L, Zhipeng R, Minglong L, Rongjun B, Xiaoyu L, Haifei L, Kun C, Xuhui Z, Jufeng Z, Lianqing L, Marios D, Stephen J, Natarjan I, Genxing P (2020) Pyrolyzed biowastes deactivated potentially toxic metals and eliminated antibiotic resistant genes for healthy vegetable production. J Clean Prod 276:124208. https://doi.org/10.1016/j.jclepro.2020.124208
Zhu D, Xiang Q, Yang XR, Ke X, O’Connor P, Zhu YG (2019) Trophic transfer of antibiotic resistance genes in a soil detritus food chain. Environ Sci Technol 53:7770–7781. https://doi.org/10.1021/acs.est.9b00214
Acknowledgements
We thank zebrafish breeding center of Medical College of South China University of Technology for providing helpful suggestions.
Funding
This research was supported financially by the Ministry of Science and Technology of China for State Key Research and Development Project (2016YFC0400702), and the Overseas Outstanding Teacher Project from the Department of Science and Technology of Guangdong Province (No. 2020A1414010108).
Author information
Authors and Affiliations
Contributions
Fengjie Hu: conceptualization, methodology, software, data curation, writing—original draft preparation. Randy Dahlgren: visualization, investigation. Taiping Zhang: writing—reviewing and editing. Jinni Liang: validation. Jiahui Xiao: validation. Zidan Liu: investigation, visualization.
Corresponding author
Ethics declarations
Ethics approval and consent to participate
All experiments were performed according to the guidelines laid down by the Institutional Animal Care and Use Committee of South China University of Technology.
Consent for publication
Not applicable.
Conflict of interest
The authors declare no competing interests.
Additional information
Responsible Editor: Zhihong Xu
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Highlights
• Amendment of biochar decreases ARGs abundance in bullfrog breeding sediment and guts of zebrafishes.
• Biochar’s impacts on inhibiting ability for all ARGs weaken after excessive addition.
• IntI1 significantly affected the horizontal gene transfer of ARGs in sediment.
• Possible transfer mechanism and pathway of ARGs between sediment and animals were proposed.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Hu, F., Zhang, T., Liang, J. et al. Impact of biochar on persistence and diffusion of antibiotic resistance genes in sediment from an aquaculture pond. Environ Sci Pollut Res 29, 57918–57930 (2022). https://doi.org/10.1007/s11356-022-19700-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11356-022-19700-2