Abstract
Introduction
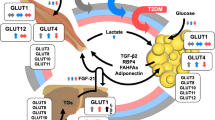
Despite the well-recognized health benefits, the mechanisms and site of action of metformin remains elusive. Metformin-induced global lipidomic changes in plasma of animal models and human subjects have been reported. However, there is a lack of systemic evaluation of metformin-induced lipidomic changes in different tissues. Metformin uptake requires active transporters such as organic cation transporters (OCTs), and hence, it is anticipated that metformin actions are tissue-dependent. In this study, we aim to characterize metformin effects in non-diabetic male mice with a special focus on lipidomics analysis. The findings from this study will help us to better understand the cell-autonomous (direct actions in target cells) or non-cell-autonomous (indirect actions in target cells) mechanisms of metformin and provide insights into the development of more potent yet safe drugs targeting a particular organ instead of systemic metabolism for metabolic regulations without major side effects.
Objectives
To characterize metformin-induced lipidomic alterations in different tissues of non-diabetic male mice and further identify lipids affected by metformin through cell-autonomous or systemic mechanisms based on the correlation between lipid alterations in tissues and the corresponding in-tissue metformin concentrations.
Methods
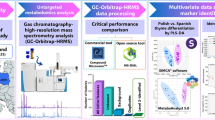
A dual extraction method involving 80% methanol followed by MTBE (methyl tert-butyl ether) extraction enables the analysis of free fatty acids, polar metabolites, and lipids. Extracts from tissues and plasma of male mice treated with or without metformin in drinking water for 12 days were analyzed using HILIC chromatography coupled to Q Exactive Plus mass spectrometer or reversed-phase liquid chromatography coupled to MS/MS scan workflow (hybrid mode) on LC-Orbitrap Exploris 480 mass spectrometer using biologically relevant lipids-containing inclusion list for data-independent acquisition (DIA), named as BRI-DIA workflow followed by data-dependent acquisition (DDA), to maximum the coverage of lipids and minimize the negative effect of stochasticity of precursor selection on experimental consistency and reproducibility.
Results
Lipidomics analysis of 6 mouse tissues and plasma allowed a systemic evaluation of lipidomic changes induced by metformin in different tissues. We observed that (1) the degrees of lipidomic changes induced by metformin treatment overly correlated with tissue concentrations of metformin; (2) the impact on lysophosphatidylcholine (lysoPC) and cardiolipins was positively correlated with tissue concentrations of metformin, while neutral lipids such as triglycerides did not correlate with the corresponding tissue metformin concentrations; (3) increase of intestinal tricarboxylic acid (TCA) cycle intermediates after metformin treatment.
Conclusion
The data collected in this study from non-diabetic mice with 12-day metformin treatment suggest that the overall metabolic effect of metformin is positively correlated with tissue concentrations and the effect on individual lipid subclass is via both cell-autonomous mechanisms (cardiolipins and lysoPC) and non-cell-autonomous mechanisms (triglycerides).








Similar content being viewed by others
Data availability
Lipidomics and metabolomics data have been uploaded to Metabolomics Workbench. The lipidomics dataset of tissue and plasma samples can be found under DataTrackID 4628, the lipid serial dilution analysis under DataTrack ID 4632, and the metabolomics dataset under DataTrack ID 4633. Other data is available upon request from the corresponding author.
Abbreviations
- LC:
-
Liquid chromatography
- HRMS:
-
High-resolution mass spectrometer
- DDA:
-
Data-dependent acquisition
- DIA:
-
Data-independent acquisition
- BRI:
-
Biologically relevant ions
References
Alnouti, Y., Petrick, J. S., & Klaassen, C. D. (2006). Tissue distribution and ontogeny of organic cation transporters in mice. Drug Metabolism and Disposition, 34(3), 477–482.
Anisimov, V. N., Popovich, I. G., Zabezhinski, M. A., Egormin, P. A., Yurova, M. N., Semenchenko, A. V., Tyndyk, M. L., Panchenko, A. V., Trashkov, A. P., Vasiliev, A. G., & Khaitsev, N. V. (2015). Sex differences in aging, life span and spontaneous tumorigenesis in 129/Sv mice neonatally exposed to metformin. Cell Cycle, 14(1), 46–55.
Bailey, C. J., Wilcock, C., & Scarpello, J. H. (2008). Metformin and the intestine. Diabetologia, 51(8), 1552–1553.
Bayoumy, A. B., Mulder, C. J. J., Mol, J. J., & Tushuizen, M. E. (2021). Gut fermentation syndrome: A systematic review of case reports. United European Gastroenterol J, 9(3), 332–342.
Bowman, A. P., Abzalimov, R. R., & Shvartsburg, A. A. (2017). Broad separation of isomeric lipids by high-resolution differential ion mobility spectrometry with tandem mass spectrometry. Journal of the American Society for Mass Spectrometry, 28(8), 1552–1561.
Camporez, J. P. G., Kanda, S., Petersen, M. C., Jornayvaz, F. R., Samuel, V. T., Bhanot, S., Petersen, K. F., Jurczak, M. J., & Shulman, G. I. (2015). ApoA5 knockdown improves whole-body insulin sensitivity in high-fat-fed mice by reducing ectopic lipid content. Journal of Lipid Research, 56(3), 526–536.
Duan, L., Scheidemantle, G., Lodge, M., Cummings, M. J., Pham, E., Wang, X., Kennedy, A., & Liu, X. (2022). Prioritize biologically relevant ions for data-independent acquisition (BRI-DIA) in LC-MS/MS-based lipidomics analysis. Metabolomics, 18(8), 55.
Fahy, E., Subramaniam, S., Murphy, R. C., Nishijima, M., Raetz, C. R., Shimizu, T., Spener, F., van Meer, G., Wakelam, M. J., & Dennis, E. A. (2009). Update of the LIPID MAPS comprehensive classification system for lipids. Journal of Lipid Research, 50(Suppl), S9-14.
Fontaine, E. (2018). Metformin-induced mitochondrial complex I inhibition: Facts, uncertainties, and consequences. Frontiers in Endocrinology (lausanne), 9, 753.
Geerling, J. J., Boon, M. R., van der Zon, G. C., van den Berg, S. A., van den Hoek, A. M., Lombes, M., Princen, H. M., Havekes, L. M., Rensen, P. C., & Guigas, B. (2014). Metformin lowers plasma triglycerides by promoting VLDL-triglyceride clearance by brown adipose tissue in mice. Diabetes, 63(3), 880–891.
He, L. (2020). Metformin and systemic metabolism. Trends in Pharmacological Sciences, 41(11), 868–881.
Jensen, J. B., Sundelin, E. I., Jakobsen, S., Gormsen, L. C., Munk, O. L., Frokiaer, J., & Jessen, N. (2016). [11C]-labeled metformin distribution in the liver and small intestine using dynamic positron emission tomography in mice demonstrates tissue-specific transporter dependency. Diabetes, 65(6), 1724–1730.
Kind, T., Liu, K. H., Lee, D. Y., DeFelice, B., Meissen, J. K., & Fiehn, O. (2013). LipidBlast in silico tandem mass spectrometry database for lipid identification. Nature Methods, 10(8), 755–758.
Kyle, J. E., Zhang, X., Weitz, K. K., Monroe, M. E., Ibrahim, Y. M., Moore, R. J., Cha, J., Sun, X., Lovelace, E. S., Wagoner, J., Polyak, S. J., Metz, T. O., Dey, S. K., Smith, R. D., Burnum-Johnson, K. E., & Baker, E. S. (2016). Uncovering biologically significant lipid isomers with liquid chromatography, ion mobility spectrometry and mass spectrometry. The Analyst, 141(5), 1649–1659.
LaMoia, T. E., Butrico, G. M., Kalpage, H. A., Goedeke, L., Hubbard, B. T., Vatner, D. F., Gaspar, R. C., Zhang, X. M., Cline, G. W., Nakahara, K., Woo, S., Shimada, A., Huttemann, M., & Shulman, G. I. (2022). Metformin, phenformin, and galegine inhibit complex IV activity and reduce glycerol-derived gluconeogenesis. Proceedings of the National Academy of Sciences of the United States of America, 119(10), e2122287119.
LaMoia, T. E., & Shulman, G. I. (2021). Cellular and molecular mechanisms of metformin action. Endocrine Reviews, 42(1), 77–96.
Leaptrot, K. L., May, J. C., Dodds, J. N., & McLean, J. A. (2019). Ion mobility conformational lipid atlas for high confidence lipidomics. Nature Communications, 10(1), 985.
Lin, M. J., Dai, W., Scott, M. J., Li, R., Zhang, Y. Q., Yang, Y., Chen, L. Z., & Huang, X. S. (2017). Metformin improves nonalcoholic fatty liver disease in obese mice via down-regulation of apolipoprotein A5 as part of the AMPK/LXRalpha signaling pathway. Oncotarget, 8(65), 108802–108809.
Mazaleuskaya, L. L., Salamatipour, A., Sarantopoulou, D., Weng, L., FitzGerald, G. A., Blair, I. A., & Mesaros, C. (2018). Analysis of HETEs in human whole blood by chiral UHPLC-ECAPCI/HRMS. Journal of Lipid Research, 59(3), 564–575.
Mbaye, B., Borentain, P., Magdy Wasfy, R., Alou, M. T., Armstrong, N., Mottola, G., Meddeb, L., Ranque, S., Gerolami, R., Million, M., & Raoult, D. (2022). Endogenous ethanol and triglyceride production by gut Pichia kudriavzevii, Candida albicans and Candida glabrata yeasts in non-alcoholic steatohepatitis. Cells, 11(21), 3390.
Omi, J., Kano, K., & Aoki, J. (2021). Current knowledge on the biology of lysophosphatidylserine as an emerging bioactive lipid. Cell Biochemistry and Biophysics, 79(3), 497–508.
Paradies, G., Paradies, V., Ruggiero, F. M., & Petrosillo, G. (2019). Role of cardiolipin in mitochondrial function and dynamics in health and disease: Molecular and pharmacological aspects. Cells, 8(7), 728.
Rena, G., Hardie, D. G., & Pearson, E. R. (2017). The mechanisms of action of metformin. Diabetologia, 60(9), 1577–1585.
Rubino, M. E. G., Carrillo, E., Alcala, G. R., Dominguez-Martin, A., & Boulaiz, H. (2019). Phenformin as an anticancer agent: Challenges and prospects. International Journal of Molecular Sciences, 20(13), 3316.
Sato, D., Morino, K., Nakagawa, F., Murata, K., Sekine, O., Beppu, F., Gotoh, N., Ugi, S., & Maegawa, H. (2017). Acute effect of metformin on postprandial hypertriglyceridemia through delayed gastric emptying. International Journal of Molecular Sciences, 18(6), 1282.
Shikata, E., Yamamoto, R., Takane, H., Shigemasa, C., Ikeda, T., Otsubo, K., & Ieiri, I. (2007). Human organic cation transporter (OCT1 and OCT2) gene polymorphisms and therapeutic effects of metformin. Journal of Human Genetics, 52(2), 117–122.
Silamikele, L., Silamikelis, I., Ustinova, M., Kalnina, Z., Elbere, I., Petrovska, R., Kalnina, I., & Klovins, J. (2021). Metformin strongly affects gut microbiome composition in high-fat diet-induced type 2 diabetes mouse model of both sexes. Frontiers in Endocrinology (lausanne), 12, 626359.
Tagesson, C., Franzen, L., Dahl, G., & Westrom, B. (1985). Lysophosphatidylcholine increases rat ileal permeability to macromolecules. Gut, 26(4), 369–377.
Tang, X., Wang, W., Hong, G., Duan, C., Zhu, S., Tian, Y., Han, C., Qian, W., Lin, R., & Hou, X. (2021). Gut microbiota-mediated lysophosphatidylcholine generation promotes colitis in intestinal epithelium-specific Fut2 deficiency. Journal of Biomedical Science, 28(1), 20.
Tsugawa, H., Ikeda, K., Takahashi, M., Satoh, A., Mori, Y., Uchino, H., Okahashi, N., Yamada, Y., Tada, I., Bonini, P., Higashi, Y., Okazaki, Y., Zhou, Z., Zhu, Z. J., Koelmel, J., Cajka, T., Fiehn, O., Saito, K., Arita, M., & Arita, M. (2020). A lipidome atlas in MS-DIAL 4. Nature Biotechnology, 38(10), 1159–1163.
Vial, G., Detaille, D., & Guigas, B. (2019). Role of mitochondria in the mechanism(s) of action of metformin. Frontiers in Endocrinology (lausanne), 10, 294.
Viel, G., Boscolo-Berto, R., Cecchetto, G., Fais, P., Nalesso, A., & Ferrara, S. D. (2012). Phosphatidylethanol in blood as a marker of chronic alcohol use: A systematic review and meta-analysis. International Journal of Molecular Sciences, 13(11), 14788–14812.
Wheaton, W. W., Weinberg, S. E., Hamanaka, R. B., Soberanes, S., Sullivan, L. B., Anso, E., Glasauer, A., Dufour, E., Mutlu, G. M., Budigner, G. S., & Chandel, N. S. (2014). Metformin inhibits mitochondrial complex I of cancer cells to reduce tumorigenesis. eLife, 3, e02242.
Yuan, T., Li, J., Zhao, W. G., Sun, W., Liu, S. N., Liu, Q., Fu, Y., & Shen, Z. F. (2019). Effects of metformin on metabolism of white and brown adipose tissue in obese C57BL/6J mice. Diabetology and Metabolic Syndrome, 11, 96.
Zake, D. M., Kurlovics, J., Zaharenko, L., Komasilovs, V., Klovins, J., & Stalidzans, E. (2021). Physiologically based metformin pharmacokinetics model of mice and scale-up to humans for the estimation of concentrations in various tissues. PLoS ONE, 16(4), e0249594.
Zhang, D., Hop, C., Patilea-Vrana, G., Gampa, G., Seneviratne, H. K., Unadkat, J. D., Kenny, J. R., Nagapudi, K., Di, L., Zhou, L., Zak, M., Wright, M. R., Bumpus, N. N., Zang, R., Liu, X., Lai, Y., & Khojasteh, S. C. (2019). Drug concentration asymmetry in tissues and plasma for small molecule-related therapeutic modalities. Drug Metabolism and Disposition, 47(10), 1122–1135.
Zhang, E., Jin, L., Wang, Y., Tu, J., Zheng, R., Ding, L., Fang, Z., Fan, M., Al-Abdullah, I., Natarajan, R., Ma, K., Wang, Z., Riggs, A. D., Shuck, S. C., Yang, L., & Huang, W. (2022). Intestinal AMPK modulation of microbiota mediates crosstalk with brown fat to control thermogenesis. Nature Communications, 13(1), 1135.
Zhang, W., Zhong, W., Sun, X., Sun, Q., Tan, X., Li, Q., Sun, X., & Zhou, Z. (2015). Visceral white adipose tissue is susceptible to alcohol-induced lipodystrophy in rats: Role of acetaldehyde. Alcoholism, Clinical and Experimental Research, 39(3), 416–423.
Zhu, X., Shen, W., Liu, Z., Sheng, S., Xiong, W., He, R., Zhang, X., Ma, L., & Ju, Z. (2020). Effect of metformin on cardiac metabolism and longevity in aged female mice. Frontiers in Cell and Developmental Biology, 8, 626011.
Acknowledgements
We thank support from Molecular Education, Technology and Research Innovation Center (METRIC) at North Carolina State University.
Funding
This work was supported by Chemistry of Life pilot funds and startup funds (L.D., G.S., M.L., X.L., A.K.) at North Carolina State University (NCSU), National Institutes of Health 1R21DK128678-01A1 (A.K.), National Institutes of Health 1R35GM150985-01 (X.L.), Center for Human Health and the Environment Pilot Project Program Award (ES025128 to X.W.) at NCSU and Animal Reproduction Program (No. 2022-67015-36491 to X.W.) from the USDA National Institute of Food and Agriculture.
Author information
Authors and Affiliations
Contributions
A.K., X.W., M.L, and M.C. performed all animal experiments. G.S., L.D., and X.L. prepared samples and performed LC–MS analysis. X.L., L.D., G.S., A.K., and X.W. participated in experimental design. X.L, L.D., and G.S. interpreted results and wrote the manuscript. E.P. helped with text editing. All authors provided input on the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
Authors declare that they have no conflicts of interest with the contents of this article.
Ethical approval
All animal procedures were approved by the Institutional Animal Care and Use Committee (IACUC) at North Carolina State University.
Consent for publication
All authors have read and approved the manuscript.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Scheidemantle, G., Duan, L., Lodge, M. et al. Data-dependent and -independent acquisition lipidomics analysis reveals the tissue-dependent effect of metformin on lipid metabolism. Metabolomics 20, 53 (2024). https://doi.org/10.1007/s11306-024-02113-2
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11306-024-02113-2