Abstract
Pecan (Carya illinoinensis) is a nut-producing tree native to North America grown for its culinary, ornamental, and lumber characteristics. A newly developed chromosome-scale reference genome for pecan was used to create genotyping-by-sequencing (GBS)-based high-density genetic linkage maps of 151 full-sibling progeny of ‘Elliott’ and ‘VC1-68’. These maps incorporate 6142 SNPs segregating in a testcross pattern into 32 linkage groups representing the 16 chromosomes of pecan across the two parents. The average distance between markers was 0.46 cM and the two maps totaled 1376.4 cM and 1463.1 cM for ‘Elliott’ and ‘VC1-68’, respectively. These markers, plus an additional 1096 intercross markers, were used to create a 1557.8-cM pecan consensus genetic linkage map. Quantitative trait locus (QTL) analyses revealed 1 major and 2 minor effect QTL for budbreak and 1 minor effect QTL for pecan scab susceptibility. The major effect locus inherited from ‘VC1-68’ explained up to 30% of the variation in budbreak and appeared across 3 years of observations. This QTL is syntenic to recently identified major effect QTL for budbreak in English walnut. The techniques reported herein, and the resulting genetic maps will facilitate future discoveries of valuable fruiting trait loci as this pecan population completes the transition to sexual maturity. In addition, the major budbreak QTL coincident in pecan and English walnut may represent a conserved mechanism effecting budbreak across the Juglandaceae providing critical knowledge for future investigations of variability in phenology.




Similar content being viewed by others
References
Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ (1990) Basic local alignment search tool. J Mol Biol 215:403–410. https://doi.org/10.1016/S0022-2836(05)80360-2
Anderson JL, Mari AR, Braasch I, Amores A, Hohenlohe P, Batzel P, Postlethwait JH (2012) Multiple sex-associated regions and a putative sex chromosome in zebrafish revealed by RAD mapping and population genomics. PLoS One 7. https://doi.org/10.1371/journal.pone.0040701
Arends D, Prins P, Broman KW, Jansen RC (2016) Tutorial-multiple-QTL Mapping (MQM) Analysis for R/qtl
Bah T (2011) Inkscape: guide to a vector drawing program. Prentice Hall, Upper Saddle River
Beedanagari SR, Dove SK, Wood BW, Conner PJ (2005) A first linkage map of pecan cultivars based on RAPD and AFLP markers. Theor Appl Genet 110:1127–1137. https://doi.org/10.1007/s00122-005-1944-5
Bentley NB, Grauke LJ, Klein PE (2019) Genotyping by sequencing (GBS) and SNP marker analysis of diverse accessions of pecan (Carya illinoinensis). Tree Genet Genomes 15:8. https://doi.org/10.1007/s11295-018-1314-5
Bernard A, Marrano A, Donkpegan A, Brown PJ, Leslie CA, Neale DB, Lheureux F, Dirlewanger E (2020) Association and linkage mapping to unravel genetic architecture of phenological traits and lateral bearing in Persian walnut (Juglans regia L.). BMC Genomics 21:1–25. https://doi.org/10.1186/s12864-020-6616-y
Bilton TP, Schofield MR, Black MA, Chagne D, Wilcox PL, Dodds KG (2018) Accounting for errors in low coverage high-throughput sequencing data when constructing genetic maps using biparental outcrossed populations. Genetics 209:65–76. https://doi.org/10.1534/genetics.117.300627
Bock CH, Cottrell TE, Hotchkiss MW, Wood BW (2013a) Vertical distribution of scab in large pecan trees. Plant Dis 97:626–634. https://doi.org/10.1094/Pdis-05-12-0453-Re
Bock CH, Wood BW, Gottwald TR (2013b) Pecan scab severity-effects of assessment methods. Plant Dis 97:675–684. https://doi.org/10.1094/Pdis-07-12-0642-Re
Bock CH, Grauke LJ, Conner P, Burrell SL, Hotchkiss MW, Boykin D, Wood BW (2016) Scab susceptibility of a provenance collection of pecan in three different seasons in the Southeastern USA. Plant Dis 100(9):1937–1945. https://doi.org/10.1094/PDIS-12-15-1398-RE
Bock CH, Hotchkiss MW, Young CA, Charlton ND, Chakradhar M, Stevenson KL, Wood BW (2017) Population genetic structure of Venturia effusa, cause of pecan scab, in the Southeastern United States. Phytopathology 107:607–619. https://doi.org/10.1094/PHYTO-10-16-0376-R
Broman KW, Wu H, Sen Ś, Churchill GA (2003) R/qtl: QTL mapping in experimental crosses. Bioinformatics 19:889–890. https://doi.org/10.1093/bioinformatics/btg112
Conner PJ (2002) A detached leaf technique for studying race-specific resistance to Cladosporium caryigenum in pecan. J Am Soc Hortic Sci 127:781–785. https://doi.org/10.21273/JASHS.127.5.781
Conner PJ (2013) Evaluation of response to pecan scab pathogen race using a detached leaflet protocol. HortScience 48:63–67. https://doi.org/10.21273/HORTSCI.48.1.63
Conner PJ, Stevenson KL (2004) Pathogenic variation of Cladosporium caryigenum isolates and corresponding differential resistance in pecan. HortScience 39:553–557. https://doi.org/10.21273/HORTSCI.39.3.553
Conner PJ, Wood BW (2001) Identification of pecan cultivars and their genetic relatedness as determined by randomly amplified polymorphic DNA analysis. J Am Soc Hortic Sci 126:474–480. https://doi.org/10.21273/JASHS.126.4.474
De Keyser E, Shu QY, Van Bockstaele E, De Riek J (2010) Multipoint-likelihood maximization mapping on 4 segregating populations to achieve an integrated framework map for QTL analysis in pot azalea (Rhododendron simsii hybrids). BMC Mol Biol 11:1. https://doi.org/10.1186/1471-2199-11-1
De Vicente M, Tanksley S (1991) Genome-wide reduction in recombination of backcross progeny derived from male versus female gametes in an interspecific cross of tomato. Theor Appl Genet 83:173–178. https://doi.org/10.1007/BF00226248
Demaree JB (1924) Pecan scab with special reference to sources of the early spring infections. J Agric Res 28:0321–0330
Garnier S, Ross N, Rudis B (2018) viridis: default color maps from “matplotlib”. R package version 05 1
Google Earth (2019) Accessed 1/27/2020
Gottwald TR (1982) Spore discharge by the pecan scab pathogen, Cladosporium caryigenum. Phytopathology 72:1193–1197. https://doi.org/10.1094/Phyto-72-1193
Grattapaglia D, Sederoff R (1994) Genetic linkage maps of Eucalyptus grandis and Eucalyptus urophylla using a pseudo-testcross: mapping strategy and RAPD markers. Genetics 137:1121–1137
Grauke LJ (2010) Pecan seed stock selection - regional implications. Proc Southeast Pecan Grow Assoc 103:42–51
Grauke LJ, Pratt J (1992) Pecan bud growth and freeze damage are influenced by rootstock. J Am Soc Hortic Sci 117:404–406. https://doi.org/10.21273/JASHS.117.3.404
Grauke LJ, Thompson TE (1996) Variability in pecan flowering. Fruit Var J 50:140–150
Grauke LJ, Iqbal MJ, Reddy AS, Thompson TE (2003) Developing microsatellite DNA markers in pecan. J Am Soc Hortic Sci 128:374–380. https://doi.org/10.21273/JASHS.128.3.0374
Grauke LJ, Mendoza-Herrera MA, Binzel ML (2010) Plastid microsatellite markers in Carya. Acta Hortic 859:237–246. https://doi.org/10.17660/ActaHortic.2010.859.27
Grauke LJ, Mendoza-Herrera MA, Miller AJ, Wood BW (2011) Geographic patterns of genetic variation in native pecans. Tree Genet Genomes 7:917–932. https://doi.org/10.1007/s11295-011-0384-4
Grauke LJ, Klein R, Grusak MA, Klein P (2015) The forest and the trees: applications for molecular markers in the repository and pecan breeding program. Acta Hortic 1070:109–126. https://doi.org/10.17660/ActaHortic.2015.1070.12
Grauke LJ, Wood BW, Harris MK (2016) Crop vulnerability: Carya. HortScience 51:653–663. https://doi.org/10.21273/HORTSCI.51.6.653
Haley CS, Knott SA (1992) A simple regression method for mapping quantitative trait loci in line crosses using flanking markers. Heredity 69:315–324
Huang BE, George AW, Forrest KL, Kilian A, Hayden MJ, Morell MK, Cavanagh CR (2012) A multiparent advanced generation inter-cross population for genetic analysis in wheat. Plant Biotechnol J 10:826–839. https://doi.org/10.1111/j.1467-7652.2012.00702.x
Huang Y, Xiao L, Zhang Z, Zhang R, Wang Z, Huang C, Huang R, Luan Y, Fan T, Wang J (2019) The genomes of pecan and Chinese hickory provide insights into Carya evolution and nut nutrition. GigaScience 8:giz036. https://doi.org/10.1093/gigascience/giz036
Hunter R, Roberts D (1978) A disease grading system for pecan scab [Fusicladium effusum]. Pecan Q
Hyma KE, Barba P, Wang M, Londo JP, Acharya CB, Mitchell SE, Sun Q, Reisch B, Cadle-Davidson L (2015) Heterozygous mapping strategy (HetMappS) for high resolution genotyping-by-sequencing markers: a case study in grapevine. PLoS One 10. https://doi.org/10.1371/journal.pone.0134880
INC (2015) Pecan. International Nut and Dried Fruit Council, Reus, Spain
INC (2016) Pecan. International Nut and Dried Fruit Council, Reus, Spain
INC (2017) Pecan. International Nut and Dried Fruit Council, Reus, Spain
INC (2018) Pecan. International Nut and Dried Fruit Council, Reus, Spain
Jenkins J, Wilson B, Grimwood J, Schmutz J, Grauke LJ (2015) Towards a reference pecan genome sequence. Acta Hortic 1070:101–108. https://doi.org/10.17660/ActaHortic.2015.1070.11
Jia XD, Wang T, Zhai M, Li YR, Guo ZR (2011) Genetic diversity and identification of Chinese-grown pecan using ISSR and SSR markers. Molecules 16:10078–10092. https://doi.org/10.3390/molecules161210078
Kassambara A (2017) ggpubr:“ggplot2” based publication ready plots. R package version 01 6
Konar A, Choudhury O, Bullis R, Fiedler L, Kruser JM, Stephens MT, Gailing O, Schlarbaum S, Coggeshall MV, Staton ME, Carlson JE, Emrich S, Romero-Severson J (2017) High-quality genetic mapping with ddRADseq in the non-model tree Quercus rubra. BMC Genomics 18:417. https://doi.org/10.1186/s12864-017-3765-8
Ladejobi O, Mackay IJ, Poland J, Praud S, Hibberd J, Bentley AR (2019) Reference genome anchoring of high density markers for association mapping and genomic prediction in European winter wheat. Front Plant Sci 10:1278. https://doi.org/10.3389/fpls.2019.01278
Li J, Zeng YR, Shen DF, Xia GH, Huang YZ, Huang YJ, Chang J, Huang JQ, Wang ZJ (2014) Development of SSR markers in hickory (Carya cathayensis Sarg.) and their transferability to other species of Carya. Curr Genom 15:357–379. https://doi.org/10.2174/138920291505141106103734
Lin M, Lou X-Y, Chang M, Wu R (2003) A general statistical framework for mapping quantitative trait loci in nonmodel systems: issue for characterizing linkage phases. Genetics 165:901–913
Lu Q, Cui Y, Wu R (2004) A multilocus likelihood approach to joint modeling of linkage, parental diplotype and gene order in a full-sib family. BMC Genet 5:20. https://doi.org/10.1186/1471-2156-5-20
Lv J, Gao B, Liu P, Li J, Meng X (2017) Linkage mapping aided by de novo genome and transcriptome assembly in Portunus trituberculatus: applications in growth-related QTL and gene identification. Sci Rep 7:1–13. https://doi.org/10.1038/s41598-017-08256-8
Mahoney LL, Sargent DJ, Abebe-Akele F, Wood DJ, Ward JA, Bassil NV, Hancock JF, Folta KM, Davis TM (2016) A high-density linkage map of the ancestral diploid strawberry, Fragaria iinumae, constructed with single nucleotide polymorphism markers from the IStraw90 array and genotyping by sequencing. Plant Genome 9. https://doi.org/10.3835/plantgenome2015.08.0071
Maliepaard C, Jansen J, Van Ooijen J (1997) Linkage analysis in a full-sib family of an outbreeding plant species: overview and consequences for applications. Genet Res 70:237–250. https://doi.org/10.1017/S00166723970030058
Margarido GR, Souza AP, Garcia AA (2007) OneMap: software for genetic mapping in outcrossing species. Hereditas 144:78–79. https://doi.org/10.1111/j.2007.0018-0661.02000.x
Marrano A, Sideli GM, Leslie CA, Cheng H, Neale DB (2019) Deciphering of the genetic control of phenology, yield, and pellicle color in Persian walnut (Juglans regia L.). Front Plant Sci 10:1140. https://doi.org/10.3389/fpls.2019.01140
Marrano A, Britton M, Zaini PA, Zimin AV, Workman RE, Puiu D, Bianco L et al (2020) High-quality chromosome-scale assembly of the walnut (Juglans Regia L.) reference genome. GigaScience 9(5). https://doi.org/10.1093/gigascience/giaa050
Morishige DT, Klein PE, Hilley JL, Sahraeian SM, Sharma A, Mullet JE (2013) Digital genotyping of sorghum - a diverse plant species with a large repeat-rich genome. BMC Genomics 14:448. https://doi.org/10.1186/1471-2164-14-448
Muranty H, Jorge V, Bastien C, Lepoittevin C, Bouffier L, Sanchez L (2014) Potential for marker-assisted selection for forest tree breeding: lessons from 20 years of MAS in crops. Tree Genet Genomes 10:1491–1510. https://doi.org/10.1007/s11295-014-0790-5
NASS (2020) Pecan production. Agricultural Statistics Board
Noé L, Kucherov G (2005) YASS: enhancing the sensitivity of DNA similarity search. Nucleic Acids Res 33:W540–W543. https://doi.org/10.1093/nar/gki478
Parchman TL, Jahner JP, Uckele KA, Galland LM, Eckert AJ (2018) RADseq approaches and applications for forest tree genetics. Tree Genet Genomes 14:39. https://doi.org/10.1007/s11295-018-1251-3
Rasband WS (1997) ImageJ. Bethesda
Rastas P (2017) Lep-MAP3: robust linkage mapping even for low-coverage whole genome sequencing data. Bioinformatics 33:3726–3732. https://doi.org/10.1093/bioinformatics/btx494
Revelle W (2017) psych: procedures for personality and psychological research
Ritter E, Gebhardt C, Salamini F (1990) Estimation of recombination frequencies and construction of RFLP linkage maps in plants from crosses between heterozygous parents. Genetics 125:645–654
Schiffthaler B, Bernhardsson C, Ingvarsson PK, Street NR (2017) BatchMap: a parallel implementation of the OneMap R package for fast computation of F1 linkage maps in outcrossing species. PLoS One 12:e0189256. https://doi.org/10.1371/journal.pone.0189256
Shi J, Ye S, Yu S, Wang Z (2013) SSR analysis of genetic diversity of the 37 new pecan cultivars. J Anhui Agric Univ 40:42–46
Sparks D (1993) Chilling and heating model for pecan budbreak. J Am Soc Hortic Sci 118:29–35. https://doi.org/10.21273/JASHS.118.1.29
Sparks D, Yates IE, Bertrand PF, Brenneman TB (2009) The relative impacts of elevation and rainy days on the incidence of scab damage of pecan nuts in the southeastern USA. J Hortic Sci Biotechnol 84:137–142. https://doi.org/10.1080/14620316.2009.11512494
Team RC (2016) R: a language and environment for statistical computing. R Foundation for Statistical Computing, Vienna
Temnykh S, DeClerck G, Lukashova A, Lipovich L, Cartinhour S, McCouch S (2001) Computational and experimental analysis of microsatellites in rice (Oryza sativa L.): frequency, length variation, transposon associations, and genetic marker potential. Genome Res 11:1441–1452. https://doi.org/10.1101/gr.184001
Thompson TE, Grauke LJ (1991) Pecans and other hickories (Carya). Acta Hortic 290:839–906. https://doi.org/10.17660/ActaHortic.1991.290.19
Thompson TE, Grauke LJ (1994) Genetic-resistance to scab disease in pecan. HortScience 29:1078–1080. https://doi.org/10.21273/HORTSCI.29.9.1078
Thompson TE, Romberg LD (1985) Inheritance of heterodichogamy in pecan. J Hered 76:456–458. https://doi.org/10.1093/oxfordjournals.jhered.a110144
Turechek WW, Stevenson KL (1998) Effects of host resistance, temperature, leaf wetness, and leaf age on infection and lesion development of pecan scab. Phytopathology 88:1294–1301. https://doi.org/10.1094/PHYTO.1998.88.12.1294
Van Ooijen J (2006) JoinMap 4. Software for the calculation of genetic linkage maps in experimental populations Kyazma BV, Wageningen, Netherlands 33
Vendrame WA, Kochert G, Wetzstein HY (1999) AFLP analysis of variation in pecan somatic embryos. Plant Cell Rep 18:853–857. https://doi.org/10.1007/s002990050673
Vendrame WA, Kochert GD, Sparks D, Wetzstein HY (2000) Field performance and molecular evaluations of pecan trees regenerated from somatic embryogenic cultures. J Am Soc Hortic Sci 125:542–546. https://doi.org/10.21273/JASHS.125.5.542
Wickham H (2016) ggplot2: elegant graphics for data analysis. Springer, New York
Wilke CO (2016) cowplot: streamlined plot theme and plot annotations for ‘ggplot2’. CRAN Repos
Wood BW, Smith MW, Worley RE, Anderson PC, Thompson TT, Grauke LJ (1997) Reproductive and vegetative characteristics of pecan cultivars. HortScience 32:1028–1033. https://doi.org/10.21273/HORTSCI.32.6.1028
Wood BW, Grauke LJ, Payne JA (1998) Provenance variation in pecan. J Am Soc Hortic Sci 123:1023–1028. https://doi.org/10.21273/JASHS.123.6.1023
Wu R, Ma C-X, Painter I, Zeng Z-B (2002) Simultaneous maximum likelihood estimation of linkage and linkage phases in outcrossing species. Theor Popul Biol 61:349–363. https://doi.org/10.1006/tpbi.2002.1577
Yadav N, De Vos S, Bock C, Wood B (2013) Development and validation of standard area diagrams to aid assessment of pecan scab symptoms on fruit. Plant Pathol 62:325–335. https://doi.org/10.1111/j.1365-3059.2012.02641.x
Yan M, Byrne DH, Klein PE, Yang J, Dong Q, Anderson N (2018) Genotyping-by-sequencing application on diploid rose and a resulting high-density SNP-based consensus map. Hortic Res 5:1–14. https://doi.org/10.1038/s41438-018-0021-6
Zhang C, Yao X, Ren H, Chang J, Wu J, Shao W, Fang Q (2020) Characterization and development of genomic SSRs in pecan (Carya illinoinensis). Forests 11:61. https://doi.org/10.3390/f11010061
Zhu T et al (2019) Sequencing a Juglans regia × J. microcarpa hybrid yields high-quality genome assemblies of parental species. Hortic Res 6:55. https://doi.org/10.1038/s41438-019-0139-1
Acknowledgments
The authors wish to thank Ms. Julie McCollum and Ms. Natalie Patterson for help in pecan DNA extraction and Illumina template preparation and Ms. Rory Tucker for her assistance evaluating SSRs. Illumina sequencing was provided by Texas A&M AgriLife Research Genomics and Bioinformatics Services. We also thank Linwood Nursery for providing access to their proprietary cultivar ‘VC1-68’ for the creation of this population. Questions regarding living inventories should be directed to the USDA-ARS Pecan Breeding and Genetics program.
Funding
This work was supported by the United States Department of Agriculture Agricultural Research Service (USDA-ARS) CRIS project 6202-21000-036-00D (Management and Characterization of Pecan Genetics Resources and Related Wild Populations), USDA-ARS CRIS project 6202-21000-035-00D (Pecan Improvement Through Breeding and Genetics), Specific Cooperative Agreement 58-6202-1-201 (Developing Molecular Markers for Carya), Specific Cooperative Agreement 58-3091-5-031 (Genomic Markers for Carya), USDA Hatch funds, and USDA-SCRI 58-6042-6009 (Coordinated Development of Genetic Tools for Pecan).
Author information
Authors and Affiliations
Contributions
LJ Grauke conceived the population’s creation, long-term care, and contributed expertise in monitoring phenotypes. Keith Kubenka contributed to the maintenance of the population as well as phenotypic evaluation. Erin Ruhlman and Nolan Bentley collected, extracted, and genotyped samples with SSRs including novel SSRs they developed. Robert Klein contributed access to the ABI3130xl and assistance with genotyping the population with SSRs. Robert Klein and Nolan Bentley contributed to GBS template DNA preparation. Xinwang Wang contributed the budbreak phenotype observations in 2018 and 2019. Patricia Klein analyzed the sequencing data and generated the SNP calls. Nolan Bentley contributed multiple phenotype evaluations, wrote the R scripts, and performed downstream analyses of the SNP and phenotypic data. Nolan Bentley wrote the manuscript with revisions and contributions to the interpretation from Patricia Klein, Robert Klein, and LJ Grauke.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no competing interests.
Data archiving statement
The sequences generated in this study have been submitted to the NCBI Sequence Read Archive (SRA) (http://www.ncbi.nlm.nih.gov/sra) under the BioProject number PRJNA602509. The reference sequence assembly used in this study (Oaxaca-v1.0, HudsonAlpha data unpublished) can be accessed with permission at https://pecantoolbox.nmsu.edu/.
Additional information
Communicated by E. Dirlewanger
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Nolan Bentley and L. J. Grauke are co-first authors
Supplementary Information
Figure S1
Scatter and density plots visualizing the per marker maximum percentage of observations explained (%Exp) by an intercross phase (%Expinter) on the x-axis and the %Exp by a testcross phase while imputing homozygous minor allele calls as heterozygous (%Exptest) on the y-axis. The scatterplot shows all 15,429 SNPs meeting minimum MAF and missing call frequency filters colored by the frequency of heterozygous calls (Het-Freq). Loci with Het-Freq closer to 0.5 (the expected frequency for intercross and testcross markers) were plotted first to reduce overplotting. The density plots show the %Exp distributions for the 11,876 markers predicted based on the observed parental genotypes to be testcrosses (blue, median = 95.7%, median absolute deviation = 3.1%), intercrosses (red, median = 89.6%, median absolute deviation = 4.3%) and the not predicted (green). Red lines show the cutoff values used to accept or reject marker phasing; A) the minimum %Exptest at testcross loci (89.6%), B) the maximum %Exptest at intercross loci (84.7%), C) the minimum %Expinter at intercross loci and the maximum %Expinter at testcross loci (81.1%). Arrows D and E indicate distributions with low Het-Freq primarily explained by cross-types where the non-heterozygous parent contains two (D) or one (E) null alleles. This is indicated by their high homozygous minor-allele frequency and testcross-like segregation patterns. Arrow F points at unmappable AAxBB loci with high Het-Freq. Plotted with ggpubr (Kassambara 2017), colored with viridis (Garnier et al. 2018), and arranged with cowplot (Wilke 2016). (PNG 850 kb)
Figure S2
Line plots showing the relationship between marker physical position and linkage position across each chromosome. ‘Elliott’ (red), ‘VC1-68’ (blue), and consensus map (grey) positions plotted separately. Rug plots show the density of markers in each map in the corresponding color. The black rug plot indicates the location of all markers predicted to be mappable based on the observed parent genotypes. (PNG 1398 kb)
Figure S3
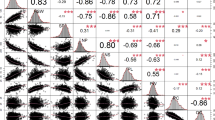
Pearson correlation between tree architecture phenotypes and associated measurements. Marker color based on position in the orchard (see row and column scatterplot). Red lines indicate line of best fit. Black line over histogram indicates density. p values of correlations (numbers) indicated as <0.05 (*), <0.01 (**), or < 0.001 (***). Correlations calculated and plotted using the psych (Revelle 2017) R package. (PNG 332 kb)
Figure S4
Pearson correlation between budbreak, pecan scab susceptibility, and associated measurements. Marker color based on position in the orchard (see row and column scatterplot). Red lines indicate line of best fit. Black line over histogram indicates density. p values of correlations (numbers) indicated as <0.05 (*), <0.01 (**), or < 0.001 (***). Correlations calculated and plotted using the psych (Revelle 2017) R package. (PNG 493 kb)
Figure S5
Scatterplot visualizing SNP marker positions comparing the results of the JoinMap v5 regression algorithm calculated linkage map (Table S5) to the R/QTL calculated consensus linkage map across all 16 chromosomes. All linkage groups calculated by JoinMap v5 consisted of markers from a single chromosome and parent. The JoinMap linkage maps were inverted when necessary to give the ‘Elliott’ informative markers a positive slope and the ‘VC1-68’ markers a negative slope to simplify visualization. Regressions calculated using lm function (Team RC 2016) and indicated by dark grey lines underneath the scatterplot. The r2 of this regression and the number of markers included (n) indicated per parental map. The overlying black lines show the reference sequence-based order of the markers and the color indicates the reference sequence position. (PNG 1058 kb)
Figure S6
Plots of LOD values associated with pecan scab incidence calculated in R/QTL (Broman et al. 2003) via the scanone Haley-Knott method with a step size of 0.1 cM. The line color and pattern indicate the origin of the plotted phenotype. Horizontal lines indicate the 98.75% cutoff for that trait determined by 10,000 permutations of the phenotypes. Rug plots show the relative LOD values (darker colors indicate greater LOD) in the ‘Elliott’ map (top rug plot), in the ‘VC1-68’ map (bottom rug plot), and at intercross loci in the consensus map (middle rug plot) plotted as a function of their position within the consensus map. (PNG 1196 kb)
Figure S7
Plots of LOD values associated with budbreak rating calculated in R/QTL (Broman et al. 2003) via the scanone Haley-Knott method with a step size of 0.1 cM. The line color and pattern indicate the origin of the plotted phenotype. Horizontal lines indicate the 98.75% cutoff for that trait determined by 10,000 permutations of the phenotypes. Rug plots show the relative LOD values (darker colors indicate greater LOD) in the ‘Elliott’ map (top rug plot), in the ‘VC1-68’ map (bottom rug plot), and at intercross loci in the consensus map (middle rug plot) plotted as a function of their position within the consensus map. (PNG 1166 kb)
Table S1
SSR primers, genotypes, and SNP marker associations (XLSX 56 kb)
Table S2
Marker statistics and phenotype associations (XLSX 8120 kb)
Table S3
Full calls and counts file for SNPs passing MAF and missing call frequency filters (XLSX 38210 kb)
Table S4
Results of tests comparing recombination frequencies between cross-direction sample groupings (XLSX 26632 kb)
Table S5
Re-analysis of marker phase and linkage position in JoinMap v5 performed across markers utilized in QTL mapping (XLSX 16 kb)
Table S6
Sample statistics and phenotypes (XLSX 47 kb)
Table S7
Summary of the scab incidence linear models and transformed marker analyses (XLSX 3368 kb)
Table S8
Summary of the budbreak linear models (XLSX 229 kb)
Table S9
Summary of the mqmscan results (XLSX 2405 kb)
Rights and permissions
About this article
Cite this article
Bentley, N., Grauke, L.J., Ruhlman, E. et al. Linkage mapping and QTL analysis of pecan (Carya illinoinensis) full-siblings using genotyping-by-sequencing. Tree Genetics & Genomes 16, 83 (2020). https://doi.org/10.1007/s11295-020-01476-6
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11295-020-01476-6