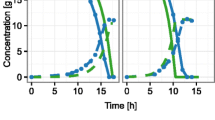
Abstract
Comparative profilings of cell growth and lipid production in the morphologically engineered strain (Δags1) and the wild type (WT) of Aspergillus oryzae BCC7051 were implemented. Using various nitrogen sources, a discrimination in cell morphology between the two strains was found, of which the Δags1 culture exhibited mycelial growth as small pellets in contrast to the WT. Of them, sodium nitrate and potassium nitrate were optimal for lipid production of the WT and Δags1 strains, respectively, which the highest lipid concentrations of 7.2 and 7.9 g L−1 were obtained in the respective cultures. The mathematical models of the growth kinetics and lipid phenotypes of both fungal strains were developed, enabling to distinguish three lipid-producing stages, including low lipid-producing, lipid accumulation, and lipid turnover stages. The model validation showed good performances in all nitrogen sources tested for the WT, but only NaNO3 and mixed yeast extract/NH4Cl were fitted well for the Δags1. The difference in the period of lipid-producing stages between the WT and Δags1 indicated the metabolic alterations of A. oryzae by the defect of a gene involved in the cell wall biosynthesis, which exhibited benefits for bioprocessing practices in addition to the high productivities of biomass and lipid. These findings would further permit the manipulation in the metabolic hub of the fungal production platform for other industrial purposes.
Graphic abstract








Similar content being viewed by others
Abbreviations
- C cal :
-
Calculated value derived from the model (g L−1)
- C exp :
-
Experimental value (g L−1)
- \(\overline{C} _{{{\text{exp}}}}\) :
-
Average value of experimental set (g L−1)
- C P :
-
Lipid concentration (g L−1)
- C S :
-
Glucose concentration (g L−1)
- C X :
-
Biomass concentration or dry cell weight (DCW) (g L−1)
- C Xf :
-
Lipid-free biomass concentration (g L−1)
- C Xm :
-
Maximum biomass concentration (g L−1)
- k d :
-
Specific rate of lipid turnover (g g−1 day−1)
- m S :
-
Biomass maintenance coefficient from glucose (g g−1 day−1)
- n :
-
Degree of lipid depletion
- q P :
-
Specific rate of lipid production (g g−1 day−1)
- q S :
-
Specific rate of glucose consumption (g g−1 day−1)
- Q L :
-
Volumetric rate of lipid reduction (g L−1 day−1)
- Q P :
-
Volumetric rate of lipid production (g L−1 day−1)
- Q S :
-
Volumetric rate of glucose consumption (g L−1 day−1)
- Q X :
-
Volumetric rate of biomass production (g L−1 day−1)
- Q Xf :
-
Volumetric rate of lipid-free biomass production (g L−1 day−1)
- R 2 :
-
Determination coefficient
- t :
-
Time (d)
- Y P / S :
-
Lipid yield on glucose (g g−1)
- Y X / S :
-
Biomass yield on glucose (g g−1)
- Y Xf / P :
-
Lipid-free biomass yield on lipid (g g−1)
- Y Xf / S :
-
Lipid-free biomass yield on glucose (g g−1)
- α:
-
Growth-associated product coefficient (g g−1)
- Β:
-
Non-growth-associated product coefficient (g g−1 day−1)
- μ :
-
Specific growth rate (day−1)
- μ Xf :
-
Specific rate of lipid-free biomass production (day−1)
- μ max :
-
Maximum specific growth rate (day−1)
References
Abd-Aziz S, Fernandez CC, Salleh MM, Illias RM, Hassan MA (2008) Effect of agitation and aeration rates on chitinase production using Trichoderma virens UKM1 in 2-l stirred tank reactor. Appl Biochem Biotechnol 150:193–204. https://doi.org/10.1007/s12010-008-8140-4
Abdala AF, Gallardo AP, Olvera LG, Silva EME (2017) Hydrolysis of carotenoid esters from Tagetes erecta by the action of lipases from Yarrowia lipolytica. Bioresour Bioprocess 4:5. https://doi.org/10.1186/s40643-016-0131-7
Abou-Zeid AM, Abd El-Zaher EHF, Saad-Allah KM, Ahmed RU (2019) Screening and optimization of some Egyptian soil-born fungi for lipids production as possible source for biofuel. Egypt J Exp Biol 15:243–250. https://doi.org/10.5455/egyjebb.20190709123823
Akpinar-Bayizit A (2014) Fungal lipids: the biochemistry of lipid accumulation. Int J Chem Eng Appl 5:409–414. https://doi.org/10.7763/IJCEA.2014.V5.419
American Public Health Association (2005) Standard methods for the examination of water and wastewater, twenty-first ed. American Public Health Association, American Water Works Association and Water Pollution Control Federation, Washington
Amirsadeghi M, Shields-Menard S, French WT, Hernandez R (2015) Lipid production by Rhodotorula glutinis from pulp and paper wastewater for biodiesel production. J Sustain Bioenergy Syst 5:114–125. https://doi.org/10.4236/jsbs.2015.53011
Bhargava S, Wenger KS, Marten MR (2003) Pulsed addition of limiting-carbon during Aspergillus oryzae fermentation leads to improved productivity of a recombinantenzyme. Biotechnol Bioeng 82:111–117. https://doi.org/10.1002/bit.10548
Čertík M, Adamechová Z, Laoteng K (2012) Microbial production of γ-linolenic acid: submerged versus solid-state fermentations. Food Sci Biotechnol 21:921–926. https://doi.org/10.1007/s10068-012-0121-2
Chen HC, Chang CC (1996) Production of γ-linolenic acid by the fungus Cunninghamella echinulata CCRC 31840. Biotechnol Prog 12:338–341. https://doi.org/10.1021/bp960009y
Chutrakul C, Jeennor S, Panchanawaporn S, Cheawchanlertfa P, Suttiwattanakul S, Mayura V, Laoteng K (2016) Metabolic engineering of long chain-polyunsaturatedfatty acid biosynthetic pathway in oleaginous fungus for dihomo-gamma linolenicacid production. J Biotechnol 218:85–93. https://doi.org/10.1016/j.jbiotec.2015.12.003
Driouch H, Hänsch R, Wucherpfennig T, Krull R, Wittmann C (2012) Improved enzyme production by bio-pellets of Aspergillus niger targeted morphology engineering using titanate microparticles. Biotechnol Bioeng 109:462–471. https://doi.org/10.1002/bit.23313
Fakas S, Galiotou-Panayotou M, Papanikolaou S, Komaitis M, Aggelis G (2007) Compositional shifts in lipid fractions during lipid turnover in Cunninghamella echinulata. Enzyme Microb Technol 40:1321–1327. https://doi.org/10.1016/j.enzmictec.2006.10.005
Fang X, Zhao G, Dai J, Liu H, Wang P, Wang L, Song J, Zheng Z (2018) Macro-morphological characterization and kinetics of Mortierella alpina colonies during batch cultivation. PLoS ONE 13:e0192803. https://doi.org/10.1371/journal.pone.0192803
Folch J, Lees M, Sloane-Stanley GH (1957) A simple method for the isolation and purification of total lipids from animal tissues. J Biol Chem 226:497–509
Gao D, Zeng J, Yu X, Dong T, Chen S (2014) Improved lipid accumulation by morphology engineering of oleaginous fungus Mortierella isabellina. Biotechnol Bioeng 111:1758–1766. https://doi.org/10.1002/bit.25242
He XM, Suzuki A (2003) Effect of nitrogen resources and pH on growth and fruit body formation of Coprinopsis phlyctidospora. Fungal Divers 12:35–44
Jeennor S, Laoteng K, Tanticharoen M, Cheevadhanarak S (2008) Evaluation of inoculum performance for enhancing gamma-linolenic acid production from Mucor rouxii. Lett Appl Microbiol 46:421–427. https://doi.org/10.1111/j.1472-765X.2007.02315.x
Jeennor S, Anantayanon J, Panchanawaporn S, Chutrakul C, Laoteng K (2019) Morphologically engineered strain of Aspergillus oryzae as a cell chassis for production development of functional lipids. Gene 718:144073. https://doi.org/10.1016/j.gene.2019.144073
Kumar KV (2007) Pseudo second-order models for the adsorption of safranin onto activated carbon: comparison oflinear and non-linear regression methods. J Hazard Mater 142:564–567. https://doi.org/10.1016/j.jhazmat.2006.08.018
Lang W, Dejma C, Sirisansaneeyakul S, Sakairi N (2009) Biosorption of nonylphenol on dead biomass of Rhizopus arrhizus encapsulated in chitosan beads. Bioresour Technol 100:5616–5623. https://doi.org/10.1016/j.biortech.2009.06.021
Laoteng K, Jitsue S, Dandusitapunth Y, Cheevadhanarak S (2008) Ethanol-induced changes in expression profiles of cell growth, fatty acid and desaturase genes of Mucor rouxii. Fungal Genet Biol 45:61–67. https://doi.org/10.1016/j.fgb.2007.04.006
Lepage G, Roy CC (1984) Improved recovery of fatty acid through direct transesterification without priorextraction or purification. J Lipid Res 25:1391–1396
Marudkla J, Lee WC, Wannawilai S, Chisti Y, Sirisansaneeyakul S (2018) Model of acetic acid-affected growth and poly(3-hydroxybutyrate) production by Cupriavidus necator DSM 545. J Biotechnol 268:12–20. https://doi.org/10.1016/j.jbiotec.2018.01.004
Mata-Alvarez J, Mitchell DA (2009) Mathematical modeling in biotechnology. In: Doelle HW, Rokem JS, Berovic M (eds) Biotechnology—fundamentals in biotechnology, vol 2. EOLSS Publishers, Paris, pp 102–139
Meeuwse P, Akbari P, Tramper J, Rinzema A (2012) Modeling growth, lipid accumulation and lipid turnover in submerged batch cultures of Umbelopsis isabellina. Bioproc Biosyst Eng 35:591–603. https://doi.org/10.1007/s00449-011-0632-x
Moubasher MH, Hamouda MS, Tammam AM, Tahoun IF (2016) Biodiesel productionfrom agricultural and agroindustrial wastes by Fusarium oxysporum. Egypt J Bot 56:733–751. https://doi.org/10.21608/EJBO.2016.2731
Olsvik ES, Kristiansen B (1994) Rheology of filamentous fermentations. Biotechnol Adv 12:1–39. https://doi.org/10.1016/0734-9750(94)90288-7
Papanikolaou S, Sarantou S, Komaitis M, Aggelis G (2004) Repression of reserve lipid turnover in Cunninghamella echinulata and Mortierella isabellina cultivated in multiple-limited media. J Appl Microbiol 97:867–875. https://doi.org/10.1111/j.1365-2672.2004.02376.x
Park WS, Murphy PA, Glatz BA (1990) Lipid metabolism and cell composition of the oleaginous yeast Apiotrichum curvatum grown at different carbon to nitrogen ratios. Can J Microbiol 36:318–326. https://doi.org/10.1139/m90-056
Patrovsky M, Sinovska K, Branska B, Patakova P (2019) Effect of initial pH, different nitrogen sources, and cultivation time on the production of yellow or orange Monascus purpureus pigments and the mycotoxin citrinin. Food Sci Nutr 7:3494–3500. https://doi.org/10.1002/fsn3.1197
Pirozzi D, Toscano G, Caputo G, Florio C, Rugari F, Travaglini G, Yousuf A, Zuccaro G (2015) Effect of nickel contamination on the growth of oleaginous yeasts in hydrolysates of Arundo donax. Environ Eng Manag J 14:1683–1690. https://doi.org/10.30638/eemj.2015.179
Ratledge C (1991) Microorganisms for lipids. Acta Biotechnol 11:429–438. https://doi.org/10.1002/abio.370110506
Ren HY, Liu BF, Ma C, Zhao L, Ren NQ (2013) A new lipid-rich microalga Scenedesmus sp. strain R-16 isolated using Nile red staining: effects of carbon and nitrogen sources and initial pH on the biomass and lipid production. Biotechnol Biofuels 6:143. https://doi.org/10.1186/1754-6834-6-143
Rossi M, Amaretti A, Raimondi S, Leonardi A (2011) Getting lipids for biodiesel production from oleaginous fungi. In: Stoytcheva M, Montero G (eds) Biodiesel-feedstocks and processing technologies. InTech, Croatia, pp 71–92
Shuler ML, Kargi F (2002) Bioprocess engineering. Prentice-Hall International Inc., New Jersey
Somashekar D, Venkateshwaran G, Sambaiah K, Lokesh BR (2002) Effect of culture conditions on lipid and gamma-linolenic acid production by mucoraceous fungi. Process Biochem 38:1719–1724. https://doi.org/10.1016/S0032-9592(02)00258-3
Surendhiran D, Vijay M, Sivaprakash B, Sirajunnisa A (2015) Kinetic modeling of microalgal growth and lipid synthesisfor biodiesel production. 3 Biotech 5:663–669. https://doi.org/10.1007/s13205-014-0264-3
Thammarongtham C, Nookaew I, Vorapreeda T, Srisuk T, Land ML, Jeennor S, Laoteng K (2018) Genome characterization of oleaginous Aspergillus oryzae BCC7051: a potential fungal-based platform for lipid production. Curr Microbiol 75:57–70. https://doi.org/10.1007/s00284-017-1350-7
Van’t Riet K, Tramper J (1991) Basic bioreactor design. Marcel Dekker INC, New York
Vorapreeda T, Thammarongtham C, Cheevadhanarak S, Laoteng K (2012) Alternative routes of acetyl-CoA synthesis identified by comparative genomic analysis: involvement in the lipid production of oleaginous yeast and fungi. Microbiology 158:217–228. https://doi.org/10.1099/mic.0.051946-0
Wang XL, Han WJ, Peng K, Zhang HY (2010) Screening of oleaginous microorganisms from filamentous fungi for microbial lipids production. In: 4th int conf bioinformatics biomed eng. https://doi.org/10.1109/ICBBE.2010.5517481
Wannawilai S, Chisti Y, Sirisansaneeyakul S (2017) A model of furfural-inhibited growth and xylitol production by Candida magnoliae TISTR 5663. Food Bioprod Process 105:129–140. https://doi.org/10.1016/j.fbp.2017.07.002
Wheeler KA, Hurdman BF, Pitt JI (1991) Influence of pH on the growth of some toxigenic species of Aspergillus, Penicillium and Fusarium. Int J Food Microbiol 12:141–149. https://doi.org/10.1016/0168-1605(91)90063-U
Wucherpfennig T, Hestler T, Krull R (2011) Morphology engineering-osmolality and its effect on Aspergillus niger morphology and productivity. Microb Cell Factories 10:58. https://doi.org/10.1186/1475-2859-10-58
Xu XH, Liu ZX, Shi XY, Miao C, Sheng S, Xu Y, Wu FA, Wang J (2018) Fed-batch fermentation of Yarrowia lipolytica using defatted silkworm pupae hydrolysate:a dynamic model-based approach for high yield of lipid production. Waste Biomass Valori 9:2399–2411. https://doi.org/10.1007/s12649-017-0180-y
Yang J, Rasa E, Tantayotai P, Scow KM, Yuan H, Hristova KR (2011) Mathematical model of Chlorella minutissima UTEX2341 growth and lipid production under photoheterotrophic fermentation conditions. Bioresour Technol 102:3077–3082. https://doi.org/10.1016/j.biortech.2010.10.049
Yodsuwan N, Sawayama S, Sirisansaneeyakul S (2017) Effect of nitrogen concentration on growth, lipid production and fatty acid profiles of the marine diatom Phaeodactylum tricornutum. Agric Nat Resour 51:190–197. https://doi.org/10.1016/j.anres.2017.02.004
Zhan J, Hong Y, Hu H (2016) Effects of nitrogen sources and C/N ratios on the lipid-producing potential ofChlorella sp. HQ J Microbiol Biotechnol 26:1290–1302. https://doi.org/10.4014/jmb.1512.12074
Zhang M, Wu W, Guo X, Weichen Y, Qi F, Jiang X, Huang J (2018) Mathematical modeling of fed-batch fermentation of Schizochytrium sp. FJU-512 growth and DHA production using a shift control strategy. 3 Biotech 8:162. https://doi.org/10.1007/s13205-018-1187-1
Acknowledgements
We gratefully thank the Targeted Research Grant of Functional Ingredients and Food Innovation, National Science and Technology Development Agency, for supporting this research work (Project No. P19-52263). We would like to thank Mr. Podsawee Limsuwannarot for language proofreading.
Author information
Authors and Affiliations
Contributions
SW designed and carried out experiments, analyzed data, developed the mathematical models and wrote the manuscript. BK and SJ assisted the analysis of metabolites and cell morphology. KL conceived and designed the research project, interpreted the results, revised and finalized the manuscript. All authors read and approved the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Research involving human and animal rights
This article does not contain any studies with human participants or animals performed by any of the authors.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Wannawilai, S., Jeennor, S., Khongto, B. et al. Exploring differential traits of lipid-producing stages of the wild type and morphologically engineered strain of Aspergillus oryzae by comparative kinetic modeling. World J Microbiol Biotechnol 36, 183 (2020). https://doi.org/10.1007/s11274-020-02959-3
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11274-020-02959-3