Abstract
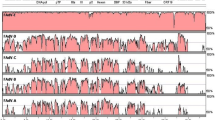
Fowl adenoviruses (FAdVs) are common in broiler operations, and the most frequently isolated FAdVs belong to serotypes 1, 8, and 11. Serotype 1 viruses are considered nonpathogenic. While some serotype 8 and 11 viruses cause inclusion body hepatitis (IBH), these virus serotypes can also be isolated from non-IBH cases. The fiber protein is one of the major constituents of the adenoviral capsid, involved in virus entry, and it has been implicated in the variation of virulence of FAdVs. The fiber gene sequences of four FAdV-8 and four FAdV-11 isolates from both IBH and non-IBH cases were determined and analyzed for a possible association of the fiber gene sequence in virulence. The fiber protein can be divided into tail, shaft, and head domains comprising some specific features. The conserved “RKRP” sequence motif (aa 17–aa 20) fit the consensus sequence predicted for the nuclear localization signal, while the “VYPF” motif (aa 53–aa 56), involved in the penton base interaction, was also found. Similar to mammalian adenoviruses, 17 pseudo-repeats with an average length of 16 aa were detected in the FAdV-8 fiber shaft region, while 20 pseudo-repeats with an average length of 18 aa were found in FAdV-11 fibers. There was a 144–147 nt difference between the fiber genes of the two FAdV serotypes. In the shaft region, the TLWT motif that marks the beginning of the fiber head domain of the mastadenovirus was not evident among examined FAdVs. The FAdV-11 isolates had 99.1 % aa sequence identity and 99.3 % similarity to each other, and there was no conserved aa substitution within the fibers. The FAdV-8 fiber proteins showed an overall lower, 89 % aa sequence identity and 93.4 % similarity, to each other and 22 nonsynonymous mutations were detected. Virulence markers were not detected in the analyzed fiber gene sequences of the different pathotypes of the two FAdV serotypes.


Similar content being viewed by others
Abbreviations
- FAdV-8:
-
Fowl adenovirus serotype 8
- FAdV-11:
-
Fowl adenovirus serotype 11
- IBH:
-
Inclusion body hepatitis
- NLS:
-
Nuclear localization signal
References
B. Harrach, M. Benkő, G.W. Both, M. Brown, A.J. Davison, M. Echavarría, M. Hess, M.S. Jones, A. Kajon et al., in Virus Taxonomy: Classification and Nomenclature of Viruses: Ninth Report of the International Committee on Taxonomy of Viruses, ed. by A.M.Q. King, M.J. Adams, E.B. Carstens, E.J. Lefkowitz (Elsevier, San Diego, 2011), pp. 125–141
B.M. Adair, S.D. Fitzgerald, in Diseases of Poultry, 12th edn., ed. by Y.M. Saif, A.M. Fadly, J.R. Glisson, L.R. McDougald, L.K. Nolan, D.E. Swayne (Blackwell Publishing, Ames, 2008), pp. 251–291
M. Saifuddin, C. Wilks, Arch. Virol. 116, 33 (1991)
S. Gomis, R. Goodhope, D. Ojkic, P. Willson, Avian Dis. 50, 550 (2006)
D. Ojkic, E. Martin, J. Swinton, J.P. Vaillancourt, M. Boulinanne, S. Gomis, Avian Pathol. 37, 95 (2008)
D. Ojkic, T. Tuboly, P.J. Krell, É. Nagy, Can. J. Vet. Res. 72, 236 (2008)
J. Pallister, P.J. Wright, M. Sheppard, J. Virol. 70, 5115 (1996)
A. Marek, V. Nolte, A. Schachner, E. Berger, C. Schlötterer, M. Hess, Vet. Microbiol. 156, 411 (2012)
U.B. Rasmussen, Y. Schlesinger, A. Pavirani, M. Mehtali, Gene 159, 279 (1995)
H.S. Alexander, P. Huber, J.X. Cao, P.J. Krell, É. Nagy, J. Virol. Methods 74, 9 (1998)
D. Ojkic, É. Nagy, Virology 283, 197 (2001)
H. Grgić, D.-H. Yang, É. Nagy, Virus Res. 156, 91 (2011)
J.C. Corredor, A. Garceac, P.J. Krell, É. Nagy, Virus Genes 36, 331 (2008)
S. Chiocca, R. Kurzbauer, G. Schaffner, A. Baker, V. Mautner, M. Cotten, J. Virol. 70, 2939 (1996)
B.D. Griffin, É. Nagy, J. Gen. Virol. 92, 1260 (2011)
D. Ojkic, É. Nagy, J. Gen. Virol. 81, 1833 (2000)
A. Marek, C. Kosiol, B. Harrach, G.L. Kaján, C. Schlötterer, M. Hess, Vet. Microbiol. 166, 250 (2013)
N.M. Green, N.G. Wrigley, W.C. Russell, S.R. Martin, A.D. McLachlan, EMBO J. 2, 1357 (1983)
J. Chroboczek, B. Jacrot, Virology 161, 549 (1987)
J.S. Hong, K.G. Mullis, J.A. Engler, Virology 167(2), 545 (1988)
C. Signas, G. Akusjarvi, U. Petterson, J. Virol. 53, 672 (1985)
G.L. Kaján, A.J. Davison, V. Palya, B. Harrach, M. Benko, J. Gen. Virol. 93(Pt 11), 2457 (2012)
G.L. Kaján, R. Stefancsik, K. Ursu, V. Palya, M. Benko, Virus Res. 153(2), 226 (2010)
M. Hess, A. Cuzange, R.W.H. Ruigrok, J. Chroboczek, B. Jacrot, J. Mol. Biol. 252, 379 (1995)
M.L. Caillet-Boudin, J. Mol. Biol. 208, 195 (1989)
S.S. Hong, P. Boulanger, EMBO J. 14, 4714 (1995)
C. Zubieta, G. Schoehn, J. Chroboczek, S. Cusack, Mol. Cell 17, 121 (2005)
S. Vrati, D. Boyle, R. Kocherhans, G.W. Both, Virology 209, 400 (1995)
M. Hess, H. Blocker, P. Brandt, Virology 238, 145 (1997)
X. Li, S.K. Tikoo, Virus Genes 25, 59 (2002)
M. Sheppard, W. Werner, M.A. Johnson, DNA Seq. 8, 391 (1998)
M.J. van Raaij, A. Mitraki, G. Lavigne, S. Cusack, Nature 401, 935 (1999)
D. Xia, L.J. Henry, R.D. Gerard, J. Deisenhofer, Structure 2, 1259 (1994)
M. Mase, N. Kikuyasu, T. Imada, J. Vet. Diagn. Invest. 22, 218 (2010)
J.X. Cao, P.J. Krell, É. Nagy, J. Gen. Virol. 79, 2507 (1998)
A. Marek, E. Schulz, C. Hess, M. Hess, J. Vet. Diagn. Invest. 6, 937 (2010)
Acknowledgments
This study was supported by the Natural Sciences and Engineering Research Council of Canada (NSERC), the Canadian Poultry Research Council (CPRC), and the Ontario Ministry of Agriculture and Food (OMAF). The authors thank Dr. Davor Ojkic for providing the FAdV isolates.
Author information
Authors and Affiliations
Corresponding author
Additional information
The GenBank accession numbers of the sequences reported in this paper are: Fiber JQ034214; protein AFD32281, Fiber JQ034215; protein AFD32282, Fiber JQ034216; protein AFD32283, Fiber JQ034217; protein AFD32284, Fiber JQ034218; protein AFD32285, Fiber JQ034219; protein AFD32286, Fiber JQ034220; protein AFD32287, Fiber JQ034221; protein AFD32288.
Rights and permissions
About this article
Cite this article
Grgić, H., Krell, P.J. & Nagy, É. Comparison of fiber gene sequences of inclusion body hepatitis (IBH) and non-IBH strains of serotype 8 and 11 fowl adenoviruses. Virus Genes 48, 74–80 (2014). https://doi.org/10.1007/s11262-013-0995-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11262-013-0995-y