Abstract
Marek’s disease (MD) is the leading cause of losses in chicken production in the world. Over the past 40 years significant progress has been made in the control of MD through the use of vaccines which reduce or delay tumor formation in vaccinated flocks. However, these vaccines fail to induce an immune response that protects against infection and virus shedding. Little is known about the genetic changes that lead to attenuation and are necessary for the generation of vaccine strains. Previous research has demonstrated that serial passage of virulent strains in cell culture results in the generation of attenuated progeny. Obtaining detailed knowledge of the changes which are needed for attenuation will be important for advancing our understanding of MD biology and should facilitate the development of more potent vaccines. We have determined the complete nucleotide sequence of a bacterial artificial chromosome (BAC) construct representing the 80th passage of a very virulent plus (vv+) MD virus strain termed 584A. Pathotyping studies have indicated that this strain (584Ap80) is indeed attenuated. Bioinformatic analysis of the sequencing data has identified numerous gross genetic changes clustering in the inverted repeat regions of the genome, as well as subtle changes (single nucleotide polymorphisms or SNPs) scattered throughout the genome. Relative to the parental strain (584Ap9), insertional mutations were identified in the MD-specific genes encoding RLORF1, RLORF3, RLORF6, 23 kDa, RLORF7 (Meq), vIL8, vLip, RSORF1, and five uncharacterized novel genes. Deletions were found in four locations within the 584Ap80 genome. A large deletion (297nt) was found in the diploid genes 85.6/98.6 and a 321 nt deletion within the intergenic region between the UL3 and UL3.5 genes is predicted to create a fusion polypeptide. A single nucleotide deletion was identified within the origin of replication. Both insertions and deletions were found in the dipoid genes MDV3.4/78.3 encoding the virulence factor RLORF4. The sequencing of the attenuated strain 584Ap80 and comparison to that of the virulent parent 584A passage 9 (584Ap9) has provided a wealth of information regarding genetic changes which have occurred during the attenuation process.
Similar content being viewed by others
Introduction
Marek’s disease (MD) is an economically important neoplastic and neurodegenerative disease of poultry caused by the highly contagious, cell-associated alphaherpesvirus gallid herpesvirus 2 (GaHV-2 or MDV). The disease is largely controlled through the use of live modified vaccine formulations consisting of antigenically related herpesvirus of turkey (HVT or MeHV-1, meleagrid herpesvirus), non-oncogenic gallid herpesvirus 3 (GaHV-3) used singularly or in combination with HVT, and attenuated strains of GaHV-2; the most commonly used is a strain known as CVI988 isolated by B. H. Rispens in l972 [32]. These live vaccines have been very efficient in reducing MD-related losses since their introduction in the early 1970s, but newer more virulent pathotypes, the so-called very virulent plus (vv+) strains, have appeared in vaccinated flocks necessitating continued improvement in vaccine development [45]. It has recently been reported that these new virulent strains can cause greater than 50 percent mortality in flocks vaccinated with the latest MD vaccine formulations [24].
Despite nearly 40 years of vaccine development, only one non-CVI988 derivative (R2/23 or Md11/75) has been licensed for use as a MD vaccine in the United States [44]. In order to develop vaccines that will protect against vv+ MDV field strains, various approaches have been taken. These have involved the generation of defined mutants using modern molecular manipulation techniques with either a collection of cosmid constructs containing overlapping fragments of the GaHV-2 genome or bacterial artificial chromosome (BAC) containing the complete genome including marker-rescue experiments with genes encoding β-gal, GFP, or ecogpt [3, 7, 22, 27, 31], and classical approaches involving passage of virulent strains serially in cell culture. This latter approach has resulted in a collection of vaccine candidates that induce varying degrees of protective immunity against vv+ challenge relative to their attenuation status. Generally partially attenuated strains (lower passage number) have been reported to induce higher protection than their fully attenuated counterpart [46]. One vv+ GaHV-2 strain (584A) when passaged 100 times in chicken embryo cell culture was demonstrated to be fully attenuated in susceptible, maternal antibody-negative (ab-) chickens. However, in protection trials in ab+ chickens, protection was on average 98.5% higher with a partially attenuated 584A (passage 80) than that with the fully attenuated (passage 100) counterpart. This partially attenuated strain (584Ap80C) elicited protective immunity not only when administered as a live vaccine but also when give intramuscularly as a DNA vaccine (BAC construct containing the 584Ap80C genome) [34, 42]. The mutations responsible for its attenuation, like those in the genomes of other licensed GaHV-2 vaccines, are only recently being characterized [26, 37, 39].
The aim of this study was to determine the complete DNA sequence of BAC20 containing the genome of 584Ap80. Genes containing mutations were identified relative to sequencing data generated from the non-attenuated very virulent plus parental clone at passage 9 (584Ap9).
Material and methods
Viruses and cells
Duck embryo fibroblasts (DEFs) were prepared from 11-day-old embryos and maintained at 37°C under 5% CO2 atmosphere in Leibovitz’s L-15 medium plus McCoy 5A medium (1:1) supplemented with 2.0% new born calf serum and antibiotics. The GaHV-2 strain (584Ap9) used in this study was kindly provided by Dr Robert F. Silva, Avian Diseases and Oncology Laboratory (ADOL), East Lansing, MI. This strain was propagated in duck embryo fibroblasts and cells were harvested 4 days post infection for DNA isolation. The construction of the bacterial artificial chromosome (BAC20) harboring the genome of 584Ap80C was described previously [34].
Purification of DNA from GaHV-2 infected cells and BAC20 transformed E. coli DH10B cells
DNA was extracted from avian fibroblasts infected with GaHV-2 strain 584Ap9 using the DNeasy Tissue Kit (Qiagen, Valencia, CA) according to the manufacturer’s instruction. Briefly, 200 μl of infected cellular suspension was treated with proteolytic enzymes at 70°C for 10 min. An equal volume of 100% ethanol was then added to the lysate and the DNA was purified by mini column chromatography and centrifugation. Columns were washed with ethanol/salt solutions to remove cellular proteins. The DNA was eluted in 10 mM Tris, pH 8.0, and quantified at OD260. Large-scale preparation of BAC20 DNA from Escherichia coli DH10B cells harboring 584Ap80C BAC was achieved by silica-based affinity chromatography using commercially available kits (Qiagen, Valencia, CA).
DNA sequencing
Sequencing of 5.0 μg BAC20 (200 ng/μl) was carried out commercially using a pyrosequencing platform, the Genome Sequencer 20 (GS20) System (454 Life Science Corporation). This involved the construction of a random library of the BAC20 DNA using the methodology described by Margulies et al. [21] with slight modifications as detailed by 454 Life Sciences. Briefly, BAC20 DNA was sheared by nebulization to an average size of 500 bp. The ends of the sheared DNA were repaired and phosphorylated using T4 DNA polymerase and T4 polynucleotide kinase. Adaptor oligonucleotides were added to the repaired ends using T4 DNA ligase. Purified DNA containing adaptors were hybridized to DNA capture beads to ensure only one DNA fragment per bead and clonally amplified using emulsion PCR. After disruption of the emulsion bubbles, DNA capture beads were sieved onto a 40 × 75 mm PicoTiterPlate equipped with an eight-lane gasket and flooded with reagent needed for pyrosequencing. Sequence reads, contigs and quality scores for sequences and contigs were obtained from 454 Life Sciences. Problematic regions containing mononucleotide reiterations or repetitive regions (i.e. 132-bp direct repeat region within the repeat long regions and a-like sequences [36, 39]) were sequenced from PCR products generated in reactions containing Platinum Taq DNA polymerase (Invitrogen, Carlsbad, CA) and numerous custom primers. PCR conditions were optimized using an Eppendorf Mastercycler Gradient to determine the optimal annealing temperatures and 5% DMSO was included in the reaction mixtures to increase product yields. Sanger-based DNA sequencing was performed at the South Atlantic Area sequencing facility (Athens, GA) and Polymorphic DNA Technologies, Inc. (Alameda, CA) using the BigDye terminator cycle sequencing protocol and analyzed on a model ABI-3730 XL DNA Analyzer (Applied Biosystems, Foster City, CA).
Copy number determination of a-like sequences and 132-bp direct repeats
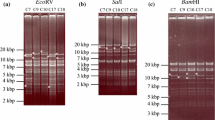
The sizing of two PCR products (1.5 and 1.7 kbp) generated in amplification reactions with primers that flank the a-like sequences and restriction endonuclease analysis using restriction endonucleases XhoI, AclI, AgeI, and BamHI of 584Ap80 DNA allowed for the determination of the number of a-like sequences within the IRL/IRS and TRS/TRL junctions. All amplification reactions contained 100 ng of BAC20 DNA, primers, Platinum Taq DNA polymerase, buffer, dNTP, and 5% DMSO. The placement of the repeat long regions relative to the termini of the unique long was based on BamHI restriction endonuclease digestion of 584Ap80.
PCR with primers that flanked the 132-bp direct repeats within the repeat long was also used to determine their copy number. A single ladder of 16 copies was resolvable by gel electrophoresis. Copy numbers greater that 16 were not resolved, indicating the PCR conditions were suboptimal or 16 copies were present in both repeat region. Subsequent restriction digestions using the enzymes mentioned above with 584Ap80 DNA indicated differences in the 132 bp copy number within the repeat long regions.
DNA sequence analysis
BAC20 DNA sequences were assembled from 60,782 reads (average length 150 nt) using the Sequencher Program (Gene Codes, Ann Arbor, MI). On average the final sequence represents a 32-fold coverage at each base pair. Ambiguities in the BAC20 sequencing data were resolved by resequencing using Sanger-based methodology. Sequence length and intragenic single nucleotide polymorphisms were identified by comparison with Sanger-based sequencing data generated using the parental 584Ap9 DNA. DNA sequences were maintained and analyzed using Lasergene (DNASTAR, Madison, WI), NCBI Entrez, and other web-based tools. Homology searches were conducted using the NCBI programs blastP and PsiBlast [33] with default settings. Published mRNA [20] and cDNA [28–30] data were compared to the 584Ap80 genome using PROT_MAP (SoftBerry, Mount Kisco, NY). Multiple alignments of proteins and nucleotide sequences were generated using MAFFT, MUSCLE, MultAlin, and MEGA 3.1 [5, 8, 13, 16]. The 132-bp direct repeat elements were investigated using the Tandyman program (Los Alamos National Laboratory, NM). The sequences of the GaHV-2 strains Md5, Md11, GA, CVI988, and RB-1B and the repeat long region of 584Ap9 used in comparisons were obtained from GenBank [4, 18, 26, 35, 40, 43]. Nucleotide sequence data reported in this paper has been submitted to GenBank nucleotide sequence database and has been assigned the accession number EU627065.
Results and discussion
Genome organization
The complete 584Ap80 genome, including the BAC cassette insertion within the US2 gene, is 187,156 bp. The lengths of the 584Ap80 subgenomic regions relative to other sequenced GaHV-2 strains are presented in Table 1. The UL region is 113,283 bp in length and extends from positions 18,035–131,317. The US region including the BAC cassette extends from positions 157,343 to 175,730. Without the BAC sequences and including the deleted US2 sequence the US region is 10,655 bp in length, a size similar to those found within the genomes of the attenuated strains CU-2 and CVI988. The terminal repeat long (TRL) and internal repeat long (IRL) extend from positions 288 to 18,034 and 131,318 to 145,897, respectively, and are 17,747 and 14,580 bp in length, respectively. The internal repeat short (IRS) is 11,166 bp in length and extends from positions 146,177 to 157,342. The terminal repeat short (TRS) is 11,167 bp in length and extends from positions 175,731 to 186,897. The first 288 and last 258 nucleotides at the termini of the 584Ap80 genomes contain the a-like sequences. The calculated size of the 548Ap80 genome (without the BAC sequences and containing the US2 sequences) is 179,434 bp and is the largest GaHV-2 genome sequenced to date. Its large size is caused by the high copy number of the 132-bp repeat elements within the repeat long regions. Each long repeat region contains a different number of the 132 bp element (16 and 40) and therefore is imperfect. Copy numbers were determined in PCR experiments with primers that flank the 132-bp repeat elements (data not shown). A ladder of 16 copies (the largest is approximately 2200 kbp) indicated that at least one long repeat contains 16 copies. Restriction endonuclease analysis with a battery of enzymes indicated that 40 copies were present in the other repeat region. Only two copies of the 132 bp repeats were found in the parental strain 584Ap9. In 584Ap9 two single nucleotide polymorphisms (C68 to T68 or C85 to A85) are present in the 132 bp repeats, (T68TTATTAAATGTGAGTTA85) and (C68TTATTAAATGTGAGTTC85). During in vitro propagation of 584Ap9, the latter repeat (e.g. T68 thru A85) is lost, while the C68 through C85 motif is amplified repeatedly. The C68 through C85 motif is only found within the 132-bp repeats of 584Ap80. Investigations into the prevalence of these SNPs within other GaHV-2 genomes reveal that the A85 SNP is unique to 584Ap9, which is biologically classified as vv MDV, while the T68 is also found in the vv MDV strains 549A and 595 and the vv+ strains 648A and 686.
Another region that was difficult to determine was the a-like sequence that is involved in cleavage of concatemers during virus replication and important for efficient packaging of viral unit-length genomes into newly synthesized nucleocapsids [23]. Analysis of the raw sequencing data indicated that DR1 sequences (ggccgcgagagg or its compliment cctctcgcggcc) are flanked by unique motifs (e.g. pac1 or pac2) which are characteristic of a-like sequences. The results showed that two copies of the a-like sequence are present in the terminal repeat long/terminal repeat short junction within the circular form of 584Ap80 (BAC). This organization, in which a-like sequences share a common DR1 site, was further validated via restriction endonuclease analysis (unpublished data). Since two a-like sequences are head-to-tail in 584Ap80, we were able to define the “unit” a-like sequence and map the junction between IRL/a-like sequence and a-like/TRS sequences [23, 41]. In the linearized DNA sequence submitted to GenBank for the 584Ap80 genome, the a-like sequence is 288 bp in length and is bracketed by DR1 sequences (ggccgcgagagg). It starts at position 145,898 and ends at position 146,185. The sequences of the termini are defined by the DR1 sequence (cctctcgcggcc) ending at position 287 and starting at position 186,898. Cleavage is postulated to occur 9 bp downstream from the DR1 sequence (Spatz, unpublished data).
Open reading frames
A list of 202 “core” ORFs, originally defined by annotating the GaHV-2 genomes GA, Md5, Md11, CVI988, RB-1B, and CU-2, were examined for mutations, which may account for the attenuated phenotype of 584Ap80. Table 2 lists relevant haploid and diploid 584Ap80 ORFs that differ between attenuated (CVI988) and virulent (CU-2, Md5, Md11, GA, and RB-1B) strains based on the predicted amino acid lengths and SNPs. Overall, 21 ORFs (counting diploid genes only once) contained mutations clustering in the repeat regions of the 584Ap80 genome (Fig. 1). These genes were first identified via comparison with ORFs of other sequenced GaHV-2 strains and then verified by resequencing selected regions of low passage virulent strain 584Ap9. In addition to the mutated ORFs, six ORFs (MDV15.5, MDV81.0/103.0, 85.3/98.9, and 86.5), which are generally present in other GaHV-2 strains, were absent in the 584Ap80 genome (Table 3). Out of this group of genes, the parental 584Ap9 genome contains MDV15.5. This is not surprising since it is unlikely that MDV81.0/103.0, MDV85.3/98.9, and MDV86.5 encode functional proteins. The number of amino acids of the MDV81.0/103.0 gene products is heterogeneous among the sequenced strains. Moreover, the predicted polypeptide lengths of the MDV85.3/98.9 ORFs differ between strains as well as between their diploid counterparts in the same strain (e.g. CVI988) (Table 3). Likewise, putative ORF MDV86.5 has only been found in virulent strains Md5 and Md11 and its length is highly variable. The only gene that is missing in the genome of 584Ap80 and is likely to encode a polypeptide is MDV15.5, predicted to encode a UL3.5 homologue.
In order to investigate the ORFs within the genome of 584Ap80 containing mutations, multiple amino acid and nucleotide alignments with homologous sequences from other GaHV-2 strains as well as the parental strain 584Ap9 were performed. An inspection of these alignments (Figs. 2–6) reveals that all of the mutations arose during serial passage of 584A. None of the mutations were present in the parental strain. These mutations are grouped into five general categories: insertions, homopolymer differences (mononucleotide reiterations), deletions, alternative start codon usage, and mixed (insertions and deletions or insertions and homopolymer differences).
Multiple nucleotide alignments of select regions of four genes containing nucleotide insertions. Sequences were aligned relative to the reference sequences 584Ap9, CVI988, and Md5. Polynucleotides and dinucleotides unique to 584Ap80 are in bold. Nucleotides shaded in gray denote regions of heterogeneity. Nucleotides in black boxes indicate a 3 nuc deletion in the CVI988 strain. The bracketing nucleotide numbers indicate the locations of the sequences within the genomes. (a) MDV3.2/78.4 encoding RLORF3, (b) MDV5.1/77.5 encoding RLORF6, (c) MDV4.0/77.0 encoding 23 KDa protein, and (d) novel open reading frame MDV5.2/76.8
ORFs containing insertions
Open reading frames containing insertions were further categorized on whether the nucleotide insertions occur in unique sequences or in homopolymer stretches. MDV3.2/78.4, MDV5.1/77.5, MDV4.0/77.0, and MDV5.2/76.8 encoding RLORF3, RLORF6, 23 Kda, and MGGG polypeptides, respectively, all contain insertions in unique sequences (Fig. 2). Relative to all other sequenced GaHV-2 genomes, a tetranucleotide insertion (GCTT) is present after position 36 in the MDV3.2/78.4 gene. This insertion causes a frameshift mutation and would result in the production of a truncated RLORF3 polypeptide of 58 amino acids. Similarly, a dinucleotide insertion (CA) in the MDV5.1/77.5 gene after nucleotide 71 is also predicted to cause a frame shift. This would result in the production of a 178 aa polypeptide with only 23 amino terminal residues in common with all RLORF6 homologues. Three other genes (MDV4.0/77.0, MDV5.0/76.0, and MDV5.2/76.8) are affected by this dinucleotide insertion due to overlapping reading frames.
A larger percentage (64%) of the insertional mutations was identified in stretches of mononucleotide reiterations (Fig. 3). These occur in two genes that are only found in the genomes of members of the Mardivirus genus (MDV3.0/78.0 and MDV10.0) and encode the virulence factors vIL-8 and vLIP, respectively. A polypeptide of 135 amino acids is predicted for the gene encoding vIL-8 due to two additional adenosine residues in the third exon causing a reading frame shift. A frameshift mutation (one extra adenosine in exon two) is also present in the gene encoding 584Ap80 vLIP. In this case, premature termination is predicted to yield a 496 aa polypeptide.
Multiple nucleotide alignments of select regions of seven genes containing differences in nucleotide homopolymer stretches. Individual 584Ap80 sequences were aligned relative to the parental strain 584Ap9 and two reference strains: the attenuated strain CVI988 and virulent strain Md5. Nucleotides within the homopolymer stretches that are unique to 584Ap80 are in bold. Nucleotides in black boxes represent regions of sequence heterogeneity where a mutation causes a premature termination of the reading frame within one strain. The bracketing nucleotide numbers indicate the locations of the sequences within the genomes. Well characterized ORFs (a) MDV3.0/78.0 encoding vIL8 and (b) MDV10.0 encoding vLIP are shown. Also shown are ORFs encoding novel polypeptides (c) MDV0.5/80.5, (d) MDV2.6/78.5, (e) MDV5.3/76.4, (f) MDV86.0/98.0, and (g) MDV86.2/97.6. Two 584Ap80 sequences were aligned in (f) and (g) due to interstrain heterogeneity of the diploid genes
ORFs with homopolymer differences
Differences in homopolymer stretches were also identified in five novel genes (Fig. 3). Relative to other GaHV-2 strains, extra guanosine, cytidine, thymidine, and adenosine residues are present in the genes encoding MDV0.5/80.5, MDV2.6/78.5, MDV86.0/98.0, and MDV86.2/97.6, respectively. The two additional guanosine residues after nucleotide 86 in MDV0.5/80.5 of 584Ap80 result in a frame shift, which is then predicted to encode a 50 amino acid polypeptide. An additional cytidine after nucleotide 135 in MDV2.6/78.5 of 584Ap80 would yield a polypeptide of 51 aa. Interestingly, the diploid pairs (MDV86.0/98.0 and MDV86.2/97.6) present in the repeat regions of 584Ap80 are different (Fig. 3f, g). Frameshift mutations would yield polypeptides containing 94 and 86 amino acids from MDV86.0 and MDV98.0, respectively. Polypeptides containing 59 and 37 aa residues are predicted for MDV86.2 and MDV97.6, respectively.
ORFs containing deletions
Open reading frames containing deletions were identified in two diploid genes (MDV85.6/98.6 and MDV7.0/74.0). A larger deletion of 297 nucleotides was identified in the diploid genes MDV85.6 and MDV98.6 of 584Ap80 and absent in the 584Ap9 genome, as well as all other GaHV-2 strains sequenced to date. This deletion causes a frame shift and a large polypeptide of 213 amino acids is predicted relative to the 69 aa polypeptide predicted from all other sequenced MDV85.6/98.6 genes.
A single nucleotide deletion (T residue) was identified in the diploid 7.0/74.0 genes encoding RLORF12. This deletion occurs in the 5′ terminus of the genes, so it is possible that alternative start codon usage could occur and consequently result in a 34 aa polypeptide (RLORF12a) or a 76 aa polypeptide (RLORF12b) that shares homology with either the amino or carboxyl terminus of full length RLORF12, respectively. The use of the second “in-frame” downstream ATG to generate the 76 aa polypeptide is by far favored by Kozak’s rules (GGATATGG vs CGTGATGG) [15]. Coincidentally, this single nucleotide deletion occurs in the origin of replication and is likely to have an effect on the replicative fitness of 584Ap80. Deletions and single nucleotide substitutions (Fig. 4) have been identified between the two ATG codons in other attenuated strains (R2/23 and CVI988 BP-5) and strains of lower virulence (JM/102 W and RM-1) [38]. Because of this and functional data demonstrating binding between RLORF12 and growth-related translationally controlled tumor protein [19], it is postulated that translation starts at the second ATG and RLORF12 is 76 amino acids in length.
Multiple nucleotide alignment of the 5′ termini of the diploid genes MDV 7.0/74.0 encoding RLORF12. Sequences from 5 attenuated (584Ap80, CVI988, CVI988-BP5, RM-1, and R2/23) and 5 virulent strains (584Ap9, Md5, JM/102 W, Md11, and 648A) are presented. These strains were chosen to illustrate the mutations within the origen of replication (Ori L). Nucleotides in black boxes represent regions of sequence heterogeneity within the (Ori L). The bracketing nucleotide numbers indicate the locations of the sequences within the genomes or subfragments. The GenBank accession numbers for these genomes and subfragments are: CVI988-DQ530348, CVI988-BP5-DQ534536, RM-1-DQ534542, R2/23-DQ534540, Md5-AF243438, JM/102 W-DQ534539, Md11-AY510475, and 648A-DQ534534
Another region of the584Ap80 genome that differs in comparison to 584Ap9 is the intergenic region between ORFs 15 and ORF15.5. As mentioned earlier 584Ap80 does not contain a MDV15.5 gene. A 321 bp deletion in the 584Ap80 genomes encompasses the MDV15.5 start codon. The result of this is a new ORF containing the 5′ terminus of MDV15 fused to the 3′ terminus of MDV15.5. The new polypeptide MDV15/15.5 (or UL3/UL3.5) would be 187 aa in length. This is of interest since a MDV recombinant containing a BAC cassette between UL3 and UL4 is attenuated in chickens (Dr. Hans Cheng, ADOL, personal communication). Homologues of the MDV15.5 gene (UL3.5) found in other animal herpesviruses pseudorabies (PRV) and bovine herpesvirus 1 (BHV-1) have been shown to be involved in virus egress [9, 10]. PRV UL3.5 null mutants are reportedly impaired in plaque formation with drastic reductions in the release of infectious virus and concurrent accumulation of naked nucleocapsids in the cytoplasm [14]. Therefore it is likely the UL3.5 gene of GaHV-2 does encode a virulence factor.
ORFs with alternative start codons
Another category of mutations involves alternative start codons. Like that postulated for RLORF12 encoding genes (MDV7.0/74.0), alternative initiation codon usage is likely to occur for two other 584Ap80 genes, namely MDV82/102 and MDV6.6/75.1. As shown in Fig. 5, MDV 82/102 encoding RSORF1 contains two extra thymidine residues. Because of this insertion, a 76 aa polypeptide (RSORFb) could result if the first start codon was used. However, it is possible that a 94 aa (RSORFa) polypeptide could result if a downstream “out-of-frame” ATG (favored by Kozak’s rules) was used as the start codon.
Multiple alignments of the 5′ termini of two diploid genes (a) 82.0/102.0 and (b) 6.6/75.1 encoding RSORF1 and B68 that classify as genes that may use alternative start codons. Nucleotides in bold within 584Ap80 sequences denote an insertion relative to reference sequences 584Ap9, CV988, and Md5. Nucleotides in black boxes represent regions of sequence heterogeneity. The bracketing nucleotide numbers indicate the locations of the sequences within the genomes. Arrows indicate the positions of the two possible start codons and the number of amino acids within each open reading frame
The diploid MDV6.6/75.1 gene of 584Ap80 may also use alternative start codons. This gene is predicted to encode a 73 aa polypeptide in contrast to the 63 aa polypeptide predicted from all other sequenced strains. Both forms share common COOH terminal sequences, but the 584Ap80 version contains 10 additional amino acids due to a point mutation upstream of the consensus start codon. This point mutation creates a new in-frame start codon.
ORFs with mixed-type mutations
Three diploid genes, MDV3.4/78.3, MDV2.0/79.0 and MDV5.0/76.0, within the 584Ap80 genome contain mutations that were categorized as mixed (Fig. 6). Relative to 584Ap9, CVI988, and Md5, the MDV3.4/78.3 gene present in BAC20 encoding RLORF4 contained both insertions and deletions, while insertions and homopolymer differences were identified in the MDV2.0/79.0 and MDV5.0/76.0 genes encoding RLORF1 and RLORF7 (Meq). An in-frame deletion of 69 nucleotides at position 185 and an extra adenosine residue at position 362 within the RLORF4 genes are predicted to yield a polypeptide of 114 aa. This is in agreement with sequencing data previously reported by Jarosinski et al., indicating similar mutations were present in 4 out of 6 attenuated MDV strains they examined including 584Ap80 [11]. This group also reported that MDV recombinants containing deletions in RLORF4 are attenuated [12]. Also shown in Fig. 6 is an alignment of MDV2.0/79.0 sequences (RLORF1) indicating a tri-nucleotide insertion and guanosine reiteration differences. This alignment also shows both intra- and interstrain heterogeneity within RLORF1 encoding genes (MDV2.0 vs MDV79.0). Two differing copies are present within the 584Ap80 genome; with both copies differing from those found in the genomes of other sequenced GaHV-2 strains. Polypeptides of 166 and 236 aa lengths are predicted for MDV2.0 and MDV79.0, respectively. Most other strains including Md5, CVI988, RB-1B, Md11, and CU-2 contain RLORF1 encoding genes that yield 198 and 239 (GA) aa polypeptides (Table 2). Four versions of the MDV2.0/79.0 genes are predicted in the GaHV-2 genomes and are currently available in GenBank.
Multiple alignments of select regions of three diploid genes containing mixed-type mutations (a) MDV3.4/078.3, (b) MDV2.0/79.0, and (c) MDV5.0/76.0 encoding RLORF4, RLORF1, and RLORF7, respectively. The bracketing nucleotide numbers indicate the locations of the sequences within the genomes or subfragments. The alignment (nuc. 105–415 of Md5 RLORF4) depicted in (a) shows the deleted region (185–254) that is in-frame and an insertion of an adenosine residue (in bold) at position 362 which causes a premature reading frame termination at position 414. Nucleotides 213–313 of RLORF1 (Md5) are shown in (b). A unique insertion (CTT) at position 254 is shown in bold. Guanosine residues after position 295 that cause reading frame shifts are also shown in bold. The RLORF7 (Meq) alignment in (c) shows two stretches of sequences relative to RLORF 7 of Md5 (1–104 and 537–575). In the 1–104 alignment an Adenosine residue after the polyadenosine stretch is in bold within the 584Ap80 sequence. In the 537–575 alignment a unique sequence (CAGAGC) is present in 584Ap80. Also included in this alignment are nucleotide ACC insertions (in bold) within RLORF7 sequences of attenuated strains R2/23 and RM-1
Mixed-type mutations within the Meq loci of 584Ap80
The most intriguing result of the comparative sequencing effort preformed in this study, however, was the identification of mutations in the Meq gene encoded by 584Ap80. This is in agreement with the hypothesis that attenuated GaHV-2 strains contain mutations in this (diploid) gene that encodes the MDV oncoprotein. The mutations, classified as both insertions and mononucleotide reiterations, are not found in the parental strain (Fig. 6). An extra adenosine residue occurs after position 100 and a unique sequence CAGAGC is present at position 547 in the Meq alignment (Fig. 6c). Two polypeptides are predicted to result from this doubly mutated gene: RLORF7a and RLORF7b. RLORF7a starts at the wild type ATG and terminates shortly after to yield a polypeptide of 49 aa. RLORF7b (238 aa) starts at an in-frame ATG at position 376 (Md5 RLORF7) and shares 57 amino acids in common with the wild type Meq protein (full length is 339 aa). The two in-frame translation products are likely devoid of functions significant to Meq, due to the fact that they both lack critical functional domains. RLORF7a is severely truncated and only contains the amino terminal proline-rich and polylysine stretches implicated in transrepression. The larger RLORF7b is missing the critical Arg/Lys rich (basic) domain important in binding to AP-1 sites on DNA [17] and missing the leucine zipper domains important for protein-protein interactions, both homodimerization and heterodimerization, with the cellular transcription factors Jun, Fos, and AFT [25]. Taken together, the 584Ap80 Meq gene is grossly mutated and probably not functional. Future transcription studies are needed to determine whether the two newly predicted proteins resulting from the mutations in this gene are likely to be expressed. However, further inspection of the multiple alignment indicates that other attenuated strains (Fig. 6c, nucleotides in bold for R2/23 and RM-1) contain Meq mutations in the same general region (530–583) as those found in 584Ap80. This suggests that Meq may play a role in attenuation. Primers, which flank this region, may therefore have future relevance in designing PCR-based diagnostic assays that could replace the time-consuming bioassay currently in use for pathotyping [47].
Single nucleotide polymorphisms
In addition to the gross mutations mentioned above, we also mapped amino acid polymorphisms (Table 4) shared between 584Ap80 and the attenuated strain CVI988 that differ from virulent strains (Md11, RB-1B and Md5) as well as the mildly virulent strain CU-2. SNPs were found in the diploid genes (MDV3.0/78) encoding vIL-8 and the haploid gene (MDV92.0) encoding US3 kinase. The strains listed in Table 4 either contained the AAA or AAC SNP at position 356 within the MDV3.0/78 genes. Attenuated strains contain the Lys119 (AAA) polymorphism while virulent strains Md5, RB-1B, Md11 and CU-2 all contain the Asn119 (AAC) polymorphism. Two SNPs (ACT/AGT and CCG/TCG) encoding Thr4/Ser4 and Pro109/Ser109 polymorphisms, respectively, were found in the gene (MDV92.0) encoding US3 kinase. Attenuated strains contain the Thr4 (ACT) polymorphism while all virulent strains contain the Ser4 (AGT) polymorphism. The Pro109 (CCG) polymorphism is predominantly found within US3 kinases of attenuated strains, but it is also found in the mildly virulent strain CU-2, perhaps contributing to its reduced virulence phenotype. It is postulated that the mutations in the genes encoding vIL-8 and the US3 kinase may contribute to the attenuated nature of 584Ap80. However these mutations are probably of lesser importance to the attenuation of 548A then the gross mutations. Further research creating mutants containing these polymorphisms in a virulent background is needed to prove their involvement in attenuation.
Conclusions
Comparison of the complete sequence of the attenuated strain 584Ap80 to the parental, virulent strain 584Ap9 allowed for the identification of gross mutations and single nucleotide polymorphisms that, together with others sequences, will undoubtedly give insight into the mechanics of the attenuation process and other more general aspects of the pathobiology of MDV. In total, 40 ORFs contained genetic aberrations. Of these 15 ORFs (counting diploid ORFs once) contained mutations that changed the reading frame while two genes contained mutations that would allow alternative start codon usage. The remaining 23 genes contain SNPs relative to virulent strains Md5, Md11, and RB-1B, mildly attenuated strain CU-2, and attenuated strain CVI988, which were identified using BLAST searches.
Since the construction of 584Ap80 as a BAC (known as BAC20) is widely available to researchers in the field, it is strongly advised that care be taken when interpreting experimental results. Recently this BAC construct was further modified to contain a VP22/GPF fusion gene and used in morphogenesis studies [6]. Generalized statements regarding the morphogenesis of MD virus particles (e.g. lack of intracellular enveloped virions) in cell culture based on experiments that use this grossly mutated strain 584Ap80 should be made cautiously. Such observation may simply be explained as the result of the deletions found in the genomes of this highly attenuated strain, such as the MDV15.5 gene encoding the egress protein UL3.5 of 584Ap80.
References
S.F. Altschul, W. Gish, W. Miller, E.W. Myers, D.J. Lipman, Basic local alignment search tool. J. Mol. Biol. 215, 403–410 (1990)
S.F. Altschul, T.L. Madden, A.A. Schaffer, J. Zhang, Z. Zhang, W. Miller, D.J. Lipman, Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res. 25, 3389–3402 (1997). doi:https://doi.org/10.1093/nar/25.17.3389
A.S. Anderson, M.S. Parcells, R.W. Morgan, The glycoprotein D (US6) homolog is not essential for oncogenicity or horizontal transmission of Marek’s disease virus. J. Virol. 72, 2548–2553 (1998)
P. Brunovskis, L.F. Velicer, The Marek’s disease virus (MDV) unique short region: alphaherpesvirus-homologous, fowlpox virus-homologous, and MDV-specific genes. Virology 206, 324–338 (1995). doi:https://doi.org/10.1016/S0042-6822(95)80048-4
F. Corpet, Multiple sequence alignment with hierarchical clustering. Nucleic Acids Res. 16, 10881–10890 (1988). doi:https://doi.org/10.1093/nar/16.22.10881
C. Denesvre, C. Blondeau, M. Lemesle, Y. Le Vern, D. Vautherot, P. Roingeard, J.F. Vautherot, Morphogenesis of a highly replicative EGFPVP22 recombinant Marek’s disease virus in cell culture. J. Virol. 81, 12348–12359 (2007). doi:https://doi.org/10.1128/JVI.01177-07
R.L. Dienglewicz, M.S. Parcells, Establishment of a lymphoblastoid cell line using a mutant MDV containing a green fluorescent protein expression cassette. Acta Virol. 43, 106–112 (1999)
R.C. Edgar, MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res. 32, 1792–1797 (2004). doi:https://doi.org/10.1093/nar/gkh340
W. Fuchs, H. Granzow, T.C. Mettenleiter, Functional complementation of UL3.5-negative pseudorabies virus by the bovine herpesvirus 1 UL3.5 homolog. J. Virol. 71, 8886–8892 (1997)
W. Fuchs, B.G. Klupp, H. Granzow, H.J. Rziha, T.C. Mettenleiter, Identification and characterization of the pseudorabies virus UL3.5 protein, which is involved in virus egress. J. Virol. 70, 3517–3527 (1996)
K.W. Jarosinski, P.H. O’Connell, K.A. Schat, Impact of deletions within the Bam HI-L fragment of attenuated Marek’s disease virus on vIL-8 expression and the newly identified transcript of open reading frame LORF4. Virus Genes 26, 255–269 (2003). doi:https://doi.org/10.1023/A:1024447230464
K.W. Jarosinski, N. Osterrieder, V.K. Nair, K.A. Schat, Attenuation of Marek’s disease virus by deletion of open reading frame RLORF4 but not RLORF5a. J. Virol. 79, 11647–11659 (2005). doi:https://doi.org/10.1128/JVI.79.18.11647-11659.2005
K. Katoh, K. Kuma, H. Toh, T. Miyata, MAFFT version 5: improvement in accuracy of multiple sequence alignment. Nucleic Acids Res. 33, 511–518 (2005). doi:https://doi.org/10.1093/nar/gki198
B.G. Klupp, H. Granzow, W. Fuchs, E. Mundt, T.C. Mettenleiter, Pseudorabies virus UL3 gene codes for a nuclear protein which is dispensable for viral replication. J. Virol. 78, 464–472 (2004). doi:https://doi.org/10.1128/JVI.78.1.464-472.2004
M. Kozak, An analysis of 5′-noncoding sequences from 699 vertebrate messenger RNAs. Nucleic Acids Res. 15, 8125–8148 (1987). doi:https://doi.org/10.1093/nar/15.20.8125
S. Kumar, K. Tamura, M. Nei, MEGA3: integrated software for molecular evolutionary genetics analysis and sequence alignment. Brief. Bioinform. 5, 150–163 (2004). doi:https://doi.org/10.1093/bib/5.2.150
H.J. Kung, L. Xia, P. Brunovskis, D. Li, J.-L. Liu, L.F. Lee, Meq: an MDV-specific bZIP transactivator with transforming properties, in Marek’s Disease, ed. by K. Hirai (Springer, Berlin, 2000)
L.F. Lee, P. Wu, D. Sui, D. Ren, J. Kamil, H.J. Kung, R.L. Witter, The complete unique long sequence and the overall genomic organization of the GA strain of Marek’s disease virus. Proc. Natl Acad. Sci. U.S.A. 97, 6091–6096 (2000). doi:https://doi.org/10.1073/pnas.97.11.6091
H.C. Liu, M. Niikura, J.E. Fulton, H.H. Cheng, Identification of chicken lymphocyte antigen 6 complex, locus E (LY6E, alias SCA2) as a putative Marek’s disease resistance gene via a virus-host protein interaction screen. Cytogenet. Genome Res. 102, 304–308 (2003). doi:https://doi.org/10.1159/000075767
T. Maray, M. Malkinson, Y. Becker, RNA transcripts of Marek’s disease virus (MDV) serotype-1 in infected and transformed cells. Virus Genes 2, 49–68 (1988). doi:https://doi.org/10.1007/BF00569736
M. Margulies, M. Egholm, W.E. Altman, S. Attiya, J.S. Bader, L.A. Bemben, J. Berka, M.S. Braverman, Y.J. Chen, Z. Chen, S.B. Dewell, L. Du, J.M. Fierro, X.V. Gomes, B.C. Godwin, W. He, S. Helgesen, C.H. Ho, G.P. Irzyk, S.C. Jando, M.L. Alenquer, T.P. Jarvie, K.B. Jirage, J.B. Kim, J.R. Knight, J.R. Lanza, J.H. Leamon, S.M. Lefkowitz, M. Lei, J. Li, K.L. Lohman, H. Lu, V.B. Makhijani, K.E. McDade, M.P. McKenna, E.W. Myers, E. Nickerson, J.R. Nobile, R. Plant, B.P. Puc, M.T. Ronan, G.T. Roth, G.J. Sarkis, J.F. Simons, J.W. Simpson, M. Srinivasan, K.R. Tartaro, A. Tomasz, K.A. Vogt, G.A. Volkmer, S.H. Wang, Y. Wang, M.P. Weiner, P. Yu, R.F. Begley, J.M. Rothberg, Genome sequencing in microfabricated high-density picolitre reactors. Nature 437, 376–380 (2005)
D.R. Marshall, J.D. Reilly, X. Liu, R.F. Silva, Selection of Marek’s disease virus recombinants expressing the Escherichia coli gpt gene. Virology 195, 638–648 (1993). doi:https://doi.org/10.1006/viro.1993.1415
M.A. McVoy, D.E. Nixon, S.P. Adler, E.S. Mocarski, Sequences within the herpesvirus-conserved pac1 and pac2 motifs are required for cleavage and packaging of the murine cytomegalovirus genome. J. Virol. 72, 48–56 (1998)
V. Nair, Evolution of Marek’s disease—a paradigm for incessant race between the pathogen and the host. Vet. J. 170, 175–183 (2005)
V. Nair, H.-J. Kung, Marek’s disease virus oncogenicity: molecular mechanisms, in Marek’s Disease: An Emerging Problem, ed. by F. Davidson, V. Nair (Elsevier, Oxford, UK, 2004), pp. 32–48
M. Niikura, J. Dodgson, H. Cheng, Direct evidence of host genome acquisition by the alphaherpesvirus Marek’s disease virus. Arch. Virol. 151, 537–549 (2005). doi:https://doi.org/10.1007/s00705-005-0633-7
M.S. Parcells, S.F. Lin, R.L. Dienglewicz, V. Majerciak, D.R. Robinson, H.C. Chen, Z. Wu, G.R. Dubyak, P. Brunovskis, H.D. Hunt, L.F. Lee, H.J. Kung, Marek’s disease virus (MDV) encodes an interleukin-8 homolog (vIL-8): characterization of the vIL-8 protein and a vIL-8 deletion mutant MDV. J. Virol. 75, 5159–5173 (2001). doi:https://doi.org/10.1128/JVI.75.11.5159-5173.2001
F. Peng, G. Bradley, A. Tanaka, G. Lancz, M. Nonoyama, Isolation and characterization of cDNAs from BamHI-H gene family RNAs associated with the tumorigenicity of Marek’s disease virus. J. Virol. 66, 7389–7396 (1992)
Q. Peng, Y. Shirazi, Isolation and characterization of Marek’s disease virus (MDV) cDNAs from a MDV-transformed lymphoblastoid cell line: identification of an open reading frame antisense to the MDV Eco-Q protein (Meq). Virology 221, 368–374 (1996). doi:https://doi.org/10.1006/viro.1996.0388
Q. Peng, M. Zeng, Z.A. Bhuiyan, E. Ubukata, A. Tanaka, M. Nonoyama, Y. Shirazi, Isolation and characterization of Marek’s disease virus (MDV) cDNAs mapping to the BamHI-I2, BamHI-Q2, and BamHI-L fragments of the MDV genome from lymphoblastoid cells transformed and persistently infected with MDV. Virology 213, 590–599 (1995). doi:https://doi.org/10.1006/viro.1995.0031
J.T. Prigge, V. Majerciak, H.D. Hunt, R.L. Dienglewicz, M.S. Parcells, Construction and characterization of Marek’s disease viruses having green fluorescent protein expression tied directly or indirectly to phosphoprotein 38 expression. Avian Dis. 48, 471–487 (2004). doi:https://doi.org/10.1637/7110
B.H. Rispens, H. van Vloten, N. Mastenbroek, H.J. Maas, K.A. Schat, Control of Marek’s disease in the Netherlands. I. Isolation of an avirulent Marek’s disease virus (strain CVI 988) and its use in laboratory vaccination trials. Avian Dis. 16, 108–125 (1972). doi:https://doi.org/10.2307/1588905
A.A. Schaffer, L. Aravind, T.L. Madden, S. Shavirin, J.L. Spouge, Y.I. Wolf, E.V. Koonin, S.F. Altschul, Improving the accuracy of PSI-BLAST protein database searches with composition-based statistics and other refinements. Nucleic Acids Res. 29, 2994–3005 (2001). doi:https://doi.org/10.1093/nar/29.14.2994
D. Schumacher, B.K. Tischer, W. Fuchs, N. Osterrieder, Reconstitution of Marek’s disease virus serotype 1 (MDV-1) from DNA cloned as a bacterial artificial chromosome and characterization of a glycoprotein B-negative MDV-1 mutant. J. Virol. 74, 11088–11098 (2000). doi:https://doi.org/10.1128/JVI.74.23.11088-11098.2000
B. Silva, L.F. Lee, M. Heidari, I. Gimeno, S. Spatz, J. Zelepsky, The role of UL13 in generating cell-free MDV, in 4th International Workshop on the Molecular Pathogenesis of Marek’s Disease Virus, ed. by M.S. Parcells, R.W. Morgan, J. Burnside, C.J. Schmidt, E.L. Bernberg, C. Kenny, E. Eppler (University of Delaware, Newark, Delaware, 2006), p. 28
R.F. Silva, R.L. Witter, Genomic expansion of Marek’s disease virus DNA is associated with serial in vitro passage. J. Virol. 54, 690–696 (1985)
S.J. Spatz, L. Petherbridge, Y. Zhao, V. Nair, Comparative full-length sequence analysis of oncogenic and vaccine (Rispens) strains of Marek’s disease virus. J. Gen. Virol. 88, 1080–1096 (2007). doi:https://doi.org/10.1099/vir.0.82600-0
S.J. Spatz, R.F. Silva, Polymorphisms in the repeat long regions of oncogenic and attenuated pathotypes of Marek’s disease virus 1. Virus Genes 35, 41–53 (2007). doi:https://doi.org/10.1007/s11262-006-0024-5
S.J. Spatz, R.F. Silva, Sequence determination of variable regions within the genomes of gallid herpesvirus-2 pathotypes. Arch. Virol. 152, 1665–1678 (2007). doi:https://doi.org/10.1007/s00705-007-0992-3
S.J. Spatz, Y. Zhao, L. Petherbridge, L.P. Smith, S.J. Baigent, V. Nair, Comparative sequence analysis of a highly oncogenic but horizontal spread-defective clone of Marek’s disease virus. Virus Genes 35, 753–766 (2007)
B.L. Strang, N.D. Stow, Circularization of the herpes simplex virus type 1 genome upon lytic infection. J. Virol. 79, 12487–12494 (2005). doi:https://doi.org/10.1128/JVI.79.19.12487-12494.2005
B.K. Tischer, D. Schumacher, M. Beer, J. Beyer, J.P. Teifke, K. Osterrieder, K. Wink, V. Zelnik, F. Fehler, N. Osterrieder, A DNA vaccine containing an infectious Marek’s disease virus genome can confer protection against tumorigenic Marek’s disease in chickens. J. Gen. Virol. 83, 2367–2376 (2002)
E.R. Tulman, C.L. Afonso, Z. Lu, L. Zsak, D.L. Rock, G.F. Kutish, The genome of a very virulent Marek’s disease virus. J. Virol. 74, 7980–7988 (2000). doi:https://doi.org/10.1128/JVI.74.17.7980-7988.2000
K. Venugopal, Marek’s disease: an update on oncogenic mechanisms and control. Res. Vet. Sci. 69, 17–23 (2000). doi:https://doi.org/10.1053/rvsc.2000.0396
R.L. Witter, Increased virulence of Marek’s disease virus field isolates. Avian Dis. 41, 149–163 (1997). doi:https://doi.org/10.2307/1592455
R.L. Witter, Induction of strong protection by vaccination with partially attenuated serotype 1 Marek’s disease viruses. Avian Dis. 46, 925–937 (2002). doi:https://doi.org/10.1637/0005-2086(2002)046[0925:IOSPBV]2.0.CO;2
R.L. Witter, B.W. Calnek, C. Buscaglia, I.M. Gimeno, K.A. Schat, Classification of Marek’s disease viruses according to pathotype: philosophy and methodology. Avian Pathol. 34, 75–90 (2005). doi:https://doi.org/10.1080/03079450500059255
Acknowledgments
The authors would like to thank Jeremy Volkening for his excellent laboratory, and bioinformatic and graphic computer skills. This research was funded by the United States Department of Agriculture CRIS program (project number 6612-32000-043).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Spatz, S.J., Rue, C., Schumacher, D. et al. Clustering of mutations within the inverted repeat regions of a serially passaged attenuated gallid herpesvirus type 2 strain. Virus Genes 37, 69–80 (2008). https://doi.org/10.1007/s11262-008-0242-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11262-008-0242-0