Abstract
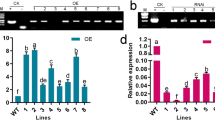
WRKY transcription factors play important roles in abiotic stress by directly regulating stress-related genes. However, the molecular mechanism of its involvement in salt stress in pak-choi is still poorly understood. In this study, we elucidated the function of BcWRKY1 from pak-choi (Brassica rapa ssp. chinensis) in salt stress. The expression level of BcWRKY1 showed the highest in rosette leaves among different tissues and was induced by salt and ABA treatment in pak-choi. Subcellular localization showed that BcWRKY1 was located in nucleus. The transgenic Arabidopsis overexpressing BcWRKY1 exhibited enhanced salt sensitivity and higher H2O2 contents, which were further confirmed by silencing BcWRKY1 in pak-choi. In addition, the expression of ZAT12 was negatively regulated with BcWRKY1 under salt stress both in pak-choi and Arabidopsis. Yeast one-hybrid and dual luciferase reporter assay showed that BcWRKY1 could bind to the promoter of BcZAT12, and BcsAPX expression was activated by BcZAT12. To sum up, we propose a BcWRKY1–BcZAT12–BcsAPX regulatory model that involves in pak-choi salt stress response.
Key message
BcWRKY1 repress the expression of BcZAT12 to inhibit reactive oxygen species scavenging, leading to enhanced salt tolerance in BcWRKY1 silencing pak-choi.










Similar content being viewed by others
Data availability
All the data supporting the findings of this study can be found in papers and supplementary materials published online.
References
Bao W, Wang X, Chen M, Chai T, Wang H (2018) A WRKY transcription factor, PcWRKY33, from Polygonum cuspidatum reduces salt tolerance in transgenic Arabidopsis thaliana. Plant Cell Rep 37:1033–1048. https://doi.org/10.1007/s00299-018-2289-2
Barrero JM, Rodríguez PL, Quesada V, Piqueras P, Ponce MR, Micol JL (2006) Both abscisic acid (ABA)-dependent and ABA-independent pathways govern the induction of NCED3, AAO3 and ABA1 in response to salt stress. Plant Cell Environ 29:2000–2008. https://doi.org/10.1111/j.1365-3040.2006.01576.x
Barrs H, Weatherley P (1962) A re-examination of the relative turgidity technique for estimating water deficits in leaves. Aust J Biol Sci 15:413. https://doi.org/10.1071/bi9620413
Besseau S, Li J, Palva ET (2012) WRKY54 and WRKY70 co-operate as negative regulators of leaf senescence in Arabidopsis thaliana. J Exp Bot 63:2667–2679. https://doi.org/10.1093/jxb/err450
Cai R, Dai W, Zhang C, Wang Y, Wu M, Zhao Y, Ma Q, Xiang Y, Cheng B (2017) The maize WRKY transcription factor ZmWRKY17 negatively regulates salt stress tolerance in transgenic Arabidopsis plants. Planta 246:1215–1231. https://doi.org/10.1007/s00425-017-2766-9
Campos PS, Quartin V, Ramalho JC, Nunes MA (2003) Electrolyte leakage and lipid degradation account for cold sensitivity in leaves of Coffea sp. plants. J Plant Physiol 160:283–292. https://doi.org/10.1078/0176-1617-00833
Chen H, Lai Z, Shi J, Xiao Y, Chen Z, Xu X (2010) Roles of Arabidopsis WRKY18, WRKY40 and WRKY60 transcription factors in plant responses to abscisic acid and abiotic stress. BMC Plant Biol 10:281. https://doi.org/10.1186/1471-2229-10-281
Chen F, Hu Y, Vannozzi A, Wu K, Cai H, Qin Y, Mullis A, Lin Z, Zhang L (2017) The WRKY transcription factor family in model plants and crops. CRC Crit Rev Plant Sci 36:311–335
Choudhury FK, Rivero RM, Blumwald E, Mittler R (2017) Reactive oxygen species, abiotic stress and stress combination. Plant J 90:856–867. https://doi.org/10.1111/tpj.13299
Clough SJ, Bent AF (1998) Floral dip: A simplified method for Agrobacterium-mediated transformation of Arabidopsis thaliana. Plant J 16:735–743. https://doi.org/10.1046/j.1365-313X.1998.00343.x
Dai W, Wang M, Gong X, Liu JH (2018) The transcription factor FcWRKY40 of Fortunella crassifolia functions positively in salt tolerance through modulation of ion homeostasis and proline biosynthesis by directly regulating SOS2 and P5CS1 homologs. New Phytol 219:972–989. https://doi.org/10.1111/nph.15240
Davletova S, Schlauch K, Coutu J, Mittler R (2005) The zinc-finger protein Zat12 plays a central role in reactive oxygen and abiotic stress signaling in Arabidopsis. Plant Physiol 139:847–856. https://doi.org/10.1104/pp.105.068254
Del Río LA (2015) ROS and RNS in plant physiology: an overview. J Exp Bot 66:2827–2837
Dionisio-Sese ML, Tobita S (1998) Antioxidant responses of rice seedlings to salinity stress. Plant Sci 135:1–9. https://doi.org/10.1016/S0168-9452(98)00025-9
Du C, Zhao P, Zhang H, Li N, Zheng L, Wang Y (2017) The Reaumuria trigyna transcription factor RtWRKY1 confers tolerance to salt stress in transgenic Arabidopsis. J Plant Physiol 215:48–58. https://doi.org/10.1016/j.jplph.2017.05.002
Eulgem T, Rushton PJ, Robatzek S, Somssich IE (2000) The WRKY superfamily of plant transcription factors. Trends Plant Sci 5:199–206. https://doi.org/10.1016/S1360-1385(00)01600-9
Funck D, Baumgarten L, Stift M, von Wirén N, Schönemann L (2020) Differential contribution of P5CS isoforms to stress tolerance in Arabidopsis. Front Plant Sci 11:1–15. https://doi.org/10.3389/fpls.2020.565134
Han C, Yang P (2015) Studies on the molecular mechanisms of seed germination. Proteomics 15:1671–1679
Hayat S, Hayat Q, Alyemeni MN, Wani AS, Pichtel J, Ahmad A (2012) Role of proline under changing environments: a review. Plant Signal Behav 7:1456–1466
He F, Niu MX, Feng CH, Li HG, Su Y, Su WL, Pang H, Yang Y, Yu X, Wang HL, Wang J, Liu C, Yin W, Xia X (2020) PeSTZ1 confers salt stress tolerance by scavenging the accumulation of ROS through regulating the expression of PeZAT12 and PeAPX2 in Populus. Tree Physiol 40:1292–1311. https://doi.org/10.1093/treephys/tpaa050
Hua ZM, Yang X, Fromm ME (2006) Activation of the NaCl- and drought-induced RD29A and RD29B promoters by constitutively active Arabidopsis MAPKK or MAPK proteins. Plant Cell Environ 29:1761–1770. https://doi.org/10.1111/j.1365-3040.2006.01552.x
Huang F, Liu T, Tang J, Duan W, Hou X (2019) BcMAF2 activates BcTEM1 and represses flowering in Pak-choi (Brassica rapa ssp. chinensis). Plant Mol Biol 100:19–32. https://doi.org/10.1007/s11103-019-00867-1
Jiang J, Ma S, Ye N, Jiang M, Cao J, Zhang J (2017) WRKY transcription factors in plant responses to stresses. J Integr Plant Biol 59:86–101. https://doi.org/10.1111/jipb.12513
Jiang Y, Tong S, Chen N, Liu B, Bai Q, Chen Y, Bi H, Zhang Z, Lou S, Tang H, Liu J, Ma T, Liu H (2020) The PalWRKY77 transcription factor negatively regulates salt tolerance and ABA signaling in Populus. Plant J. https://doi.org/10.1111/tpj.15109
Le CTT, Brumbarova T, Ivanov R, Stoof C, Weber E, Mohrbacher J, Fink-Straube C, Bauer P (2016) Zinc finger of Arabidopsis thaliana12 (zat12) interacts with fer-like iron deficiency-induced transcription factor (FIT) linking iron deficiency and oxidative stress responses1[OPEN]. Plant Physiol 170:540–557. https://doi.org/10.1104/pp.15.01589
Li ZQ, Li JT, Bing J, Zhang GF (2019) The role analysis of APX gene family in the growth and developmental processes and in response to abiotic stresses in Arabidopsis thaliana. Yi Chuan 41:534–547
Ma Z, Li W, Wang H, Yu D (2020) WRKY transcription factors WRKY12 and WRKY13 interact with SPL10 to modulate age-mediated flowering. J Integr Plant Biol 00:1–15. https://doi.org/10.1111/jipb.12946
Mahi HE, Hormaeche JP, De LA, Villalta I, Espartero J, Arjona FG, Fernández JL, Bundó M, Mendoza I, Mieulet D, Lalanne E, Lee SY, Yun DJ, Guiderdoni E, Aguilar M, Leidi EO, Pardo JM, Quintero FJ (2019) A critical role of sodium flux via the plasma membrane na+/h+ exchanger sos1 in the salt tolerance of rice. Plant Physiol 180:1046–1065. https://doi.org/10.1104/pp.19.00324
Matsuo M, Johnson JM, Hieno A, Tokizawa M, Nomoto M, Tada Y, Godfrey R, Obokata J, Sherameti I, Yamamoto YY, Böhmer FD, Oelmüller R (2015) High REDOX RESPONSIVE TRANSCRIPTION FACTOR1 levels result in accumulation of reactive oxygen species in Arabidopsis thaliana shoots and roots. Mol Plant 8:1253–1273. https://doi.org/10.1016/j.molp.2015.03.011
Miller G, Suzuki N, Ciftci-Yilmaz S, Mittler R (2010) Reactive oxygen species homeostasis and signalling during drought and salinity stresses. Plant Cell Environ 33:453–467. https://doi.org/10.1111/j.1365-3040.2009.02041.x
Noctor G, Foyer CH (1998) Ascorbate and glutathione: keeping active oxygen under control. Annu Rev Plant Biol 49:249–279. https://doi.org/10.1146/annurev.arplant.49.1.249
Pandey SP, Somssich IE (2009) The role of WRKY transcription factors in plant immunity. Plant Physiol 150:1648–1655. https://doi.org/10.1104/pp.109.138990
Pandey SP, Roccaro M, Schön M, Logemann E, Somssich IE (2010) Transcriptional reprogramming regulated by WRKY18 and WRKY40 facilitates powdery mildew infection of Arabidopsis. Plant J 64:912–923. https://doi.org/10.1111/j.1365-313X.2010.04387.x
Pavlović I, Pěnčík A, Novák O, Vujčić V, Radić Brkanac S, Lepeduš H, Strnad M, Salopek-Sondi B (2018) Short-term salt stress in Brassica rapa seedlings causes alterations in auxin metabolism. Plant Physiol Biochem 125:74–84. https://doi.org/10.1016/j.plaphy.2018.01.026
Potschin M, Schlienger S, Bieker S, Zentgraf U (2014) Senescence networking: WRKY18 is an upstream regulator, a downstream target gene, and a protein interaction partner of WRKY53. J Plant Growth Regul 33:106–118. https://doi.org/10.1007/s00344-013-9380-2
Qiu N, Liu Q, Li J, Zhang Y, Wang F, Gao J (2017) Physiological and transcriptomic responses of Chinese cabbage (Brassica rapa L ssp. pekinensis) to salt stress. Int J Mol Sci. https://doi.org/10.3390/ijms18091953
Rai AN, Penna S (2013) Molecular evolution of plant P5CS gene involved in proline biosynthesis. Mol Biol Rep 40:6429–6435. https://doi.org/10.1007/s11033-013-2757-2
Rizhsky L, Davletova S, Liang H, Mittler R (2004) The zinc finger protein Zat12 is required for cytosolic ascorbate peroxidase 1 expression during oxidative stress in Arabidopsis. J Biol Chem 279:11736–11743. https://doi.org/10.1074/jbc.M313350200
Rushton PJ, Somssich IE, Ringler P, Shen QJ (2010) WRKY transcription factors. Trends Plant Sci 15:247–258. https://doi.org/10.1016/J.TPLANTS.2010.02.006
Schön M, Töller A, Diezel C, Roth C, Westphal L, Wiermer M, Somssich IE (2013) Analyses of wrky18 wrky40 plants reveal critical roles of SA/EDS1 signaling and indole-glucosinolate biosynthesis for Golovinomyces orontii resistance and a loss-of resistance towards Pseudomonas syringae pv. tomato AvrRPS4. Mol Plant-Microbe Interact 26:758–767. https://doi.org/10.1094/mpmi-11-12-0265-r
Shang Y, Yan L, Liu ZQ, Cao Z, Mei C, Xin Q, Wu FQ, Wang XF, Du SY, Jiang T, Zhang XF, Zhao R, Sun HL, Liu R, Yu YT, Zhang DP (2010) The Mg-chelatase H subunit of Arabidopsis antagonizes a group of WRKY transcription repressors to relieve ABA-responsive genes of inhibition. Plant Cell 22:1909–1935. https://doi.org/10.1105/tpc.110.073874
Sharma P, Jha AB, Dubey RS, Pessarakli M (2012) Reactive oxygen species, oxidative damage, and antioxidative defense mechanism in plants under stressful conditions. J Bot 2012:1–26. https://doi.org/10.1155/2012/217037
Song XS, Hu WH, Mao WH, Ogweno JO, Zhou YH, Yu JQ (2005) Response of ascorbate peroxidase isoenzymes and ascorbate regeneration system to abiotic stresses in Cucumis sativus L. Plant Physiol Biochem 43:1082–1088. https://doi.org/10.1016/j.plaphy.2005.11.003
Song Y, Li J, Sui Y, Han G, Zhang Y, Guo S, Sui N (2020) The sweet sorghum SbWRKY50 is negatively involved in salt response by regulating ion homeostasis. Plant Mol Biol 102:603–614. https://doi.org/10.1007/s11103-020-00966-4
Su J, Wu S, Xu Z, Qiu S, Luo T, Yang Y, Chen Q, Xia Y, Zou S, Huang B-L, Huang B (2013) Comparison of salt tolerance in Brassicas and some related species. Am J Plant Sci 04:1911–1917. https://doi.org/10.4236/ajps.2013.410234
Téllez-Robledo B, Manzano C, Saez A, Navarro-Neila S, Silva-Navas J, de Lorenzo L, González-García MP, Toribio R, Hunt AG, Baigorri R, Casimiro I, Brady SM, Castellano MM, del Pozo JC (2019) The polyadenylation factor FIP1 is important for plant development and root responses to abiotic stresses. Plant J 99:1203–1219. https://doi.org/10.1111/tpj.14416
Ullah A, Dutta D, Fliegel L (2016) Expression and characterization of the SOS1 Arabidopsis salt tolerance protein. Mol Cell Biochem 415:133–143. https://doi.org/10.1007/s11010-016-2685-2
Ullah A, Sun H, Hakim YX, Zhang X (2018) A novel cotton WRKY gene, GhWRKY6-like, improves salt tolerance by activating the ABA signaling pathway and scavenging of reactive oxygen species. Physiol Plant 162:439–454. https://doi.org/10.1111/ppl.12651
Van Zelm E, Zhang Y, Testerink C (2020) Salt tolerance mechanisms of plants. Annu Rev Plant Biol 71:403–433. https://doi.org/10.1146/annurev-arplant-050718-100005
Vogel JT, Zarka DG, Van Buskirk HA, Fowler SG, Thomashow MF (2005) Roles of the CBF2 and ZAT12 transcription factors in configuring the low temperature transcriptome of Arabidopsis. Plant J 41:195–211. https://doi.org/10.1111/j.1365-313X.2004.02288.x
Wang Y, Hou X, Shen S (2007) Cloning and characterization of a full-length cDNA of WRKY transcription factor gene in Brassica campestris ssp chinensis. Acta Horticult Sin 33(5):985–988
Wang M, Yuan J, Qin L, Shi W, Xia G, Liu S (2020) TaCYP81D5, one member in a wheat cytochrome P450 gene cluster, confers salinity tolerance via reactive oxygen species scavenging. Plant Biotechnol J 18:791–804. https://doi.org/10.1111/pbi.13247
Wei W, Liang DW, Bian XH, Shen M, Xiao JH, Zhang WK, Ma B, Lin Q, Lv J, Chen X, Chen SY, Zhang JS (2019) GmWRKY54 improves drought tolerance through activating genes in abscisic acid and Ca2+ signaling pathways in transgenic soybean. Plant J 100:384–398. https://doi.org/10.1111/tpj.14449
Xing C, Liu Y, Zhao L, Zhang S, Huang X (2019) A novel MYB transcription factor regulates ascorbic acid synthesis and affects cold tolerance. Plant Cell Environ 42:832–845. https://doi.org/10.1111/pce.13387
Xiong L, Zhu JK (2002) Molecular and genetic aspects of plant responses to osmotic stress. Plant Cell Environ 25:131–139. https://doi.org/10.1046/j.1365-3040.2002.00782.x
Xiong L, Schumaker KS, Zhu JK (2002) Cell signaling during cold, drought, and salt stress. Plant Cell 14:165–183. https://doi.org/10.1105/tpc.000596
Yang Y, Guo Y (2018a) Elucidating the molecular mechanisms mediating plant salt-stress responses. New Phytol 217:523–539. https://doi.org/10.1111/nph.14920
Yang Y, Guo Y (2018b) Unraveling salt stress signaling in plants. J Integr Plant Biol 60:796–804. https://doi.org/10.1111/jipb.12689
Yang Y, Chi Y, Wang Z, Zhou Y, Fan B, Chen Z (2016) Functional analysis of structurally related soybean GmWRKY58 and GmWRKY76 in plant growth and development. J Exp Bot 67:4727–4742. https://doi.org/10.1093/jxb/erw252
Zhang X, Lu G, Long W, Zou X, Li F, Nishio T (2014) Recent progress in drought and salt tolerance studies in Brassica crops. Breed Sci 64:60–73. https://doi.org/10.1270/jsbbs.64.60
Zhang P, Wang R, Yang X, Ju Q, Li W, Shiyou L, Tran LSP, Xu J (2020) The R2R3-MYB transcription factor AtMYB49 modulates salt tolerance in Arabidopsis by modulating the cuticle formation and antioxidant defence. Plant Cell Environ 43:1925–1943. https://doi.org/10.1111/pce.13784
Acknowledgements
We are grateful to Dr Isabelle Jupin for providing the plasmid pTY-S. This work was supported by National Vegetable Industry Technology System (CARS-23-A-16), Jiangsu Agricultural Science and Technology Innovation Fund (CX (20)2017), the National Natural Science Foundation of China (31272173) and a project funded by the Priority Academic Program Development of Jiangsu Higher Education Institutions.
Funding
Funding was provided by National Vegetable Industry Technology System (Grant No.: CARS-23-A-16), Jiangsu Agricultural Science and Technology Innovation Fund (Grant No.: CX (20)2017), the Priority Academic Program Development of Jiangsu Higher Education Institutions, and the National Natural Science Foundation of China (Grant No.: 31272173).
Author information
Authors and Affiliations
Contributions
SY and YW performed the experiments. DH and YL designed the experiments. CS analyzed data. SY write the manuscript. DH, FC, XH, CZ, TL and YL revised the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that there is no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Yuan, S., Hu, D., Wang, Y. et al. BcWRKY1 confers salt sensitivity via inhibiting Reactive oxygen species scavenging. Plant Mol Biol 109, 741–759 (2022). https://doi.org/10.1007/s11103-022-01272-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11103-022-01272-x